Abstract
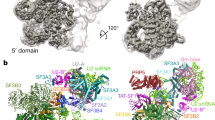
The spliceosome is a dynamic assembly of five small nuclear ribonucleoproteins (snRNPs) that removes introns from eukaryotic pre-mRNA. U6, the most conserved of the spliceosomal small nuclear RNAs (snRNAs), participates directly in catalysis. Here, we report the crystal structure of the Saccharomyces cerevisiae U6 snRNP core containing most of the U6 snRNA and all four RRM domains of the Prp24 protein. It reveals a unique interlocked RNP architecture that sequesters the 5′ splice site–binding bases of U6 snRNA. RRMs 1, 2 and 4 of Prp24 form an electropositive groove that binds double-stranded RNA and may nucleate annealing of U4 and U6 snRNAs. Substitutions in Prp24 that suppress a mutation in U6 localize to direct RNA-protein contacts. Our results provide the most comprehensive view to date of a multi-RRM protein bound to RNA and reveal striking coevolution of protein and RNA structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Brow, D.A. Allosteric cascade of spliceosome activation. Annu. Rev. Genet. 36, 333–360 (2002).
Shukla, G.C. & Padgett, R.A. A catalytically active group II intron domain 5 can function in the U12-dependent spliceosome. Mol. Cell 9, 1145–1150 (2002).
Fica, S.M. et al. RNA catalyses nuclear pre-mRNA splicing. Nature 503, 229–234 (2013).
Marcia, M. & Pyle, A.M. Visualizing group II intron catalysis through the stages of splicing. Cell 151, 497–507 (2012).
Brow, D.A. & Guthrie, C. Spliceosomal RNA U6 is remarkably conserved from yeast to mammals. Nature 334, 213–218 (1988).
Shannon, K.W. & Guthrie, C. Suppressors of a U4 snRNA mutation define a novel U6 snRNP protein with RNA-binding motifs. Genes Dev. 5, 773–785 (1991).
Mayes, A.E., Verdone, L., Legrain, P. & Beggs, J.D. Characterization of Sm-like proteins in yeast and their association with U6 snRNA. EMBO J. 18, 4321–4331 (1999).
Stevens, S.W. et al. Biochemical and genetic analyses of the U5, U6, and U4/U6 x U5 small nuclear ribonucleoproteins from Saccharomyces cerevisiae. RNA 7, 1543–1553 (2001).
Karaduman, R. et al. Structure of yeast U6 snRNPs: arrangement of Prp24p and the LSm complex as revealed by electron microscopy. RNA 14, 2528–2537 (2008).
Ghetti, A., Company, M. & Abelson, J. Specificity of Prp24 binding to RNA: a role for Prp24 in the dynamic interaction of U4 and U6 snRNAs. RNA 1, 132–145 (1995).
Raghunathan, P.L. & Guthrie, C. A spliceosomal recycling factor that reanneals U4 and U6 small nuclear ribonucleoprotein particles. Science 279, 857–860 (1998).
Vidaver, R.M., Fortner, D.M., Loos-Austin, L.S. & Brow, D.A. Multiple functions of Saccharomyces cerevisiae splicing protein Prp24 in U6 RNA structural rearrangements. Genetics 153, 1205–1218 (1999).
Rader, S.D. & Guthrie, C. A conserved Lsm-interaction motif in Prp24 required for efficient U4/U6 di-snRNP formation. RNA 8, 1378–1392 (2002).
Bae, E. et al. Structure and interactions of the first three RNA recognition motifs of splicing factor Prp24. J. Mol. Biol. 367, 1447–1458 (2007).
Martin-Tumasz, S., Richie, A.C., Clos, L.J. II, Brow, D.A. & Butcher, S.E. A novel occluded RNA recognition motif in Prp24 unwinds the U6 RNA internal stem loop. Nucleic Acids Res. 39, 7837–7847 (2011).
Jandrositz, A. & Guthrie, C. Evidence for a Prp24 binding site in U6 snRNA and in a putative intermediate in the annealing of U6 and U4 snRNAs. EMBO J. 14, 820–832 (1995).
Achsel, T. et al. A doughnut-shaped heteromer of human Sm-like proteins binds to the 3′-end of U6 snRNA, thereby facilitating U4/U6 duplex formation in vitro. EMBO J. 18, 5789–5802 (1999).
Verdone, L., Galardi, S., Page, D. & Beggs, J.D. Lsm proteins promote regeneration of pre-mRNA splicing activity. Curr. Biol. 14, 1487–1491 (2004).
Ryan, D.E., Stevens, S.W. & Abelson, J. The 5′ and 3′ domains of yeast U6 snRNA: Lsm proteins facilitate binding of Prp24 protein to the U6 telestem region. RNA 8, 1011–1033 (2002).
Dreyfuss, G., Swanson, M.S. & Pinol-Roma, S. Heterogeneous nuclear ribonucleoprotein particles and the pathway of mRNA formation. Trends Biochem. Sci. 13, 86–91 (1988).
Kenan, D.J., Query, C.C. & Keene, J.D. RNA recognition: towards identifying determinants of specificity. Trends Biochem. Sci. 16, 214–220 (1991).
Daubner, G.M., Clery, A. & Allain, F.H. RRM-RNA recognition: NMR or crystallography.and new findings. Curr. Opin. Struct. Biol. 23, 100–108 (2013).
Lunde, B.M., Moore, C. & Varani, G. RNA-binding proteins: modular design for efficient function. Nat. Rev. Mol. Cell Biol. 8, 479–490 (2007).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–D301 (2012).
Kwan, S.S. & Brow, D.A. The N- and C-terminal RNA recognition motifs of splicing factor Prp24 have distinct functions in U6 RNA binding. RNA 11, 808–820 (2005).
Martin-Tumasz, S., Reiter, N.J., Brow, D.A. & Butcher, S.E. Structure and functional implications of a complex containing a segment of U6 RNA bound by a domain of Prp24. RNA 16, 792–804 (2010).
Fortner, D.M., Troy, R.G. & Brow, D.A. A stem/loop in U6 RNA defines a conformational switch required for pre-mRNA splicing. Genes Dev. 8, 221–233 (1994).
Karaduman, R., Fabrizio, P., Hartmuth, K., Urlaub, H. & Luhrmann, R. RNA structure and RNA-protein interactions in purified yeast U6 snRNPs. J. Mol. Biol. 356, 1248–1262 (2006).
Huppler, A., Nikstad, L.J., Allmann, A.M., Brow, D.A. & Butcher, S.E. Metal binding and base ionization in the U6 RNA intramolecular stem-loop structure. Nat. Struct. Biol. 9, 431–435 (2002).
Sashital, D.G., Cornilescu, G., McManus, C.J., Brow, D.A. & Butcher, S.E. U2–U6 RNA folding reveals a group II intron-like domain and a four-helix junction. Nat. Struct. Mol. Biol. 11, 1237–1242 (2004).
Duarte, C.M., Wadley, L.M. & Pyle, A.M. RNA structure comparison, motif search and discovery using a reduced representation of RNA conformational space. Nucleic Acids Res. 31, 4755–4761 (2003).
Richardson, J.S. et al. RNA backbone: consensus all-angle conformers and modular string nomenclature (an RNA Ontology Consortium contribution). RNA 14, 465–481 (2008).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Toor, N., Keating, K.S., Taylor, S.D. & Pyle, A.M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008).
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013).
Cléry, A., Blatter, M. & Allain, F.H. RNA recognition motifs: boring? Not quite. Curr. Opin. Struct. Biol. 18, 290–298 (2008).
Madhani, H.D., Bordonne, R. & Guthrie, C. Multiple roles for U6 snRNA in the splicing pathway. Genes Dev. 4, 2264–2277 (1990).
Burke, J.E., Sashital, D.G., Zuo, X., Wang, Y.X. & Butcher, S.E. Structure of the yeast U2/U6 snRNA complex. RNA 18, 673–683 (2012).
Xu, D., Nouraini, S., Field, D., Tang, S.J. & Friesen, J.D. An RNA-dependent ATPase associated with U2/U6 snRNAs in pre-mRNA splicing. Nature 381, 709–713 (1996).
Small, E.C., Leggett, S.R., Winans, A.A. & Staley, J.P. The EF-G-like GTPase Snu114p regulates spliceosome dynamics mediated by Brr2p, a DExD/H box ATPase. Mol. Cell 23, 389–399 (2006).
Tseng, C.K. & Cheng, S.C. Both catalytic steps of nuclear pre-mRNA splicing are reversible. Science 320, 1782–1784 (2008).
Wells, S.E. et al. CUS1, a suppressor of cold-sensitive U2 snRNA mutations, is a novel yeast splicing factor homologous to human SAP 145. Genes Dev. 10, 220–232 (1996).
Kuhn, A.N., Li, Z. & Brow, D.A. Splicing factor Prp8 governs U4/U6 RNA unwinding during activation of the spliceosome. Mol. Cell 3, 65–75 (1999).
Kuhn, A.N. & Brow, D.A. Suppressors of a cold-sensitive mutation in yeast U4 RNA define five domains in the splicing factor Prp8 that influence spliceosome activation. Genetics 155, 1667–1682 (2000).
Galej, W.P., Oubridge, C., Newman, A.J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013).
Zhou, L. et al. Crystal structures of the Lsm complex bound to the 3′ end sequence of U6 small nuclear RNA. Nature 506, 116–120 (2014).
Staley, J.P. & Guthrie, C. Mechanical devices of the spliceosome: motors, clocks, springs, and things. Cell 92, 315–326 (1998).
Ares, M. Jr. & Weiser, B. Rearrangement of snRNA structure during assembly and function of the spliceosome. Prog. Nucleic Acid Res. Mol. Biol. 50, 131–159 (1995).
Wilkins, M.R. et al. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 112, 531–552 (1999).
Kibbe, W.A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Petrov, A.S., Bowman, J.C., Harvey, S.C. & Williams, L.D. Bidentate RNA-magnesium clamps: on the origin of the special role of magnesium in RNA folding. RNA 17, 291–297 (2011).
Klein, D.J., Moore, P.B. & Steitz, T.A. The contribution of metal ions to the structural stability of the large ribosomal subunit. RNA 10, 1366–1379 (2004).
Winn, M.D., Isupov, M.N. & Murshudov, G.N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 57, 122–133 (2001).
Ramachandran, G.N., Ramakrishnan, C. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 7, 95–99 (1963).
Krissinel, E. & Henrick, K. Detection of protein assemblies in crystals. Lect. Notes Comput. Sci. 3695, 163–174 (2005).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Sikorski, R.S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Acknowledgements
We are grateful to J. Doudna, C. Guthrie, A. Hoskins, H. Noller, A.M. Pyle and members of the Brow and Butcher laboratories for helpful discussions and critical reading of the manuscript, C. Bingman for advice and technical assistance in performing crystallization screening and B. Bhattacharyya and J. Keck for acquisition of diffraction data. Use of the Advanced Photon Source, an Office of Science User Facility operated for the US Department of Energy (DOE) Office of Science by Argonne National Laboratory, was supported by the US DOE under contract no. DE-AC02-06CH11357. Use of the LS-CAT Sector 21 was supported by grant 085P1000817. This work was supported by a grant from the US National Institutes of Health (no. GM065166 to S.E.B. and D.A.B.).
Author information
Authors and Affiliations
Contributions
E.J.M. and H.L. prepared crystallization samples; E.J.M. determined the structure; E.C.C., K.L.A., C.N.T. and D.A.B. performed the yeast genetic analysis; S.E.B. and D.A.B. supervised the work and wrote the manuscript with assistance from E.J.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Wall-eyed stereo image of the U6–Prp24 crystal structure.
(a) Cartoon representation of the interlocked RNP topology, colored as in Figure 2. (b) Electron density from the final 2mFo-DFc map contoured at 2.0 rmsd. The RNA (nucleotides 48-53 of the U6 asymmetric bulge) is colored yellow to enhance clarity, while the protein is colored as in Figure 2. Red spheres depict associated water molecules.
Supplementary Figure 2 RNA structural features of interest.
(a) U80, which plays an important role in pre-mRNA splicing63,64, is base-paired with C67 in the U6-A62G-Prp24 crystal structure. The red sphere depicts associated solvent. (b) The observed structure of the ISL is within experimental error (RMSD < 1 Å) of the solution structure of the isolated ISL65. (c) Similar to an unusual backbone conformation observed previously in duplex RNA66, there are two inverted ribose moieties in U6; however, this ribose inversion is dictated through extensive contacts between RNA and protein. The directionality of the backbone is depicted by highlighted O4' atoms (green). Inverted nucleotide A40 makes a non-Watson-Crick base pair with A91 at the end of the U6 telestem. The inversion in A42 is part of the aspartate bridge motif described in Figures 4a and 6a. (RRM3, which forms extensive contacts with RNA in this region of the structure, has been omitted for clarity.) RNA and protein are colored as in Figure 2.
Supplementary Figure 3 Mutations in PRP24 isolated as genomic suppressors of U4-G14C6 also suppress U6-A62G and U6-UA cold sensitivity.
Strains containing the indicated U6 and PRP24 alleles were serially diluted and plated at 16 °C and 37 °C. The Prp24-L217P substitution confers heat-sensitivity at 37 °C in addition to suppression at 16 °C6.
Supplementary Figure 4 Agreement between chemical-probing data and the U6–Prp24 crystal structure.
Observed contacts between Prp24 and U6 along the single-stranded asymmetric bulge help to explain previous chemical probing studies28, where blue nucleotides were protected from chemical modification, pink nucleotides were moderately reactive, and red nucleotides were strongly reactive. (Gray indicates no information.) Reactive base-paired regions in the ISL and telestem are consistent with helical breathing during the long incubation times used for chemical modification (40-80 minutes at 4°C)28. The RRM domains are colored as in Figure 2 and are shown with a partially transparent solvent accessible surface (calculated using a 1.4 Å probe radius).
Supplementary Figure 5 Proposed assembly mechanism of the interlocked U6–Prp24 structure.
The telestem, which is relatively unstable due to three non-canonical A-A or A-G pairs (Fig. 2a), likely exists only transiently in free U6 RNA. Binding of RRM2 to the ACAGA-box region induces binding of the RRM2 loop spanning residues 149-160 to oRRM4 (middle panel). Binding of RRM3 to U6 nucleotides 39-44 and 91-92 then stabilizes the telestem.
Supplementary Figure 6 Proposed architecture of the U6 snRNP from yeast.
(a) The Lsm ring46 (gray) is placed near the 3' tail of U6 RNA to allow association of the 3' nucleotide with the torus of the ring. (b) The register of the ring is made to be consistent with observed cross-linking9 between G30 (red surface) and Lsm2 (yellow) and (c) yeast two-hybrid13 data suggesting the C-terminal tail of Prp24 interacts with Lsm5, Lsm7 and Lsm8 (light green). The positioning of the SNFFL box in close proximity to the U6 telestem could explain previously reported cross-links between U6 nucleotides 28 and 29 with unidentified residues in Prp24 (ref. 28).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1 and 2, and Supplementary Note (PDF 8289 kb)
Architecture of the U6–Prp24 crystal structure
Structure of the U6 RNA–Prp24 complex highlighting the interlocked RNA-protein topology, aspartate bridged base pair, and extensive interactions of RRM2 with the conserved U6 ACAGA sequence. (MP4 19362 kb)
Rights and permissions
About this article
Cite this article
Montemayor, E., Curran, E., Liao, H. et al. Core structure of the U6 small nuclear ribonucleoprotein at 1.7-Å resolution. Nat Struct Mol Biol 21, 544–551 (2014). https://doi.org/10.1038/nsmb.2832
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2832
This article is cited by
-
Architecture of the U6 snRNP reveals specific recognition of 3′-end processed U6 snRNA
Nature Communications (2018)
-
Mechanistic insights into precursor messenger RNA splicing by the spliceosome
Nature Reviews Molecular Cell Biology (2017)
-
Usb1 controls U6 snRNP assembly through evolutionarily divergent cyclic phosphodiesterase activities
Nature Communications (2017)
-
Stereomicroscopic 3D-pattern profiling of murine and human intestinal inflammation reveals unique structural phenotypes
Nature Communications (2015)
-
Specificity and nonspecificity in RNA–protein interactions
Nature Reviews Molecular Cell Biology (2015)