Abstract
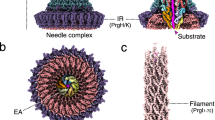
Type 3 secretion systems use 3.5-megadalton syringe-like, membrane-embedded 'injectisomes', each containing an ~800-Å-long needle complex to connect intracellular compartments of infectious bacteria and hosts. Here we identify requirements for substrate association with, transport through and exit from the injectisome of Salmonella enterica serovar Typhimurium. This guided the design of substrates that become trapped within the secretion path and enabled visualization of injectisomes in action in situ. We used cryo-EM to define the secretion path, providing a structural explanation as to why effector proteins must be unfolded during transport. Furthermore, trapping of a heterologous substrate in the needle prevents secretion of natural bacterial effectors. Together, the data reveal the path of protein secretion across multiple membranes and show that mechanisms rejecting unacceptable substrates can be undermined, and transport of bacterial effectors across an already assembled type 3 secretion system can be inhibited.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hueck, C.J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433 (1998).
Coburn, B., Sekirov, I. & Finlay, B.B. Type III secretion systems and disease. Clin. Microbiol. Rev. 20, 535–549 (2007).
Kubori, T. et al. Supramolecular structure of the Salmonella typhimurium type III protein secretion system. Science 280, 602–605 (1998).
Sekiya, K. et al. Supermolecular structure of the enteropathogenic Escherichia coli type III secretion system and its direct interaction with the EspA-sheath-like structure. Proc. Natl. Acad. Sci. USA 98, 11638–11643 (2001).
Galán, J.E. & Wolf-Watz, H. Protein delivery into eukaryotic cells by type III secretion machines. Nature 444, 567–573 (2006).
Edgren, T., Forsberg, A., Rosqvist, R. & Wolf-Watz, H. Type III secretion in Yersinia: injectisome or not? PLoS Pathog. 8, e1002669 (2012).
Marlovits, T.C. et al. Structural insights into the assembly of the type III secretion needle complex. Science 306, 1040–1042 (2004).
Schraidt, O. & Marlovits, T.C. Three-dimensional model of Salmonella's needle complex at subnanometer resolution. Science 331, 1192–1195 (2011).
Hodgkinson, J.L. et al. Three-dimensional reconstruction of the Shigella T3SS transmembrane regions reveals 12-fold symmetry and novel features throughout. Nat. Struct. Mol. Biol. 16, 477–485 (2009).
Winnen, B. et al. Hierarchical effector protein transport by the Salmonella Typhimurium SPI-1 type III secretion system. PLoS ONE 3, e2178 (2008).
Stebbins, C.E. & Galan, J.E. Modulation of host signaling by a bacterial mimic: structure of the Salmonella effector SptP bound to Rac1. Mol. Cell 6, 1449–1460 (2000).
Loquet, A. et al. Atomic model of the type III secretion system needle. Nature 486, 276–279 (2012).
Fujii, T. et al. Structure of a type III secretion needle at 7-Å resolution provides insights into its assembly and signaling mechanisms. Proc. Natl. Acad. Sci. USA 109, 4461–4466 (2012).
Galkin, V.E., Schmied, W.H., Schraidt, O., Marlovits, T.C. & Egelman, E.H. The structure of the Salmonella typhimurium type III secretion system needle shows divergence from the flagellar system. J. Mol. Biol. 396, 1392–1397 (2010).
Lilic, M., Vujanac, M. & Stebbins, C.E. A common structural motif in the binding of virulence factors to bacterial secretion chaperones. Mol. Cell 21, 653–664 (2006).
Stebbins, C.E. & Galan, J.E. Structural mimicry in bacterial virulence. Nature 412, 701–705 (2001).
Stebbins, C.E. & Galán, J.E. Maintenance of an unfolded polypeptide by a cognate chaperone in bacterial type III secretion. Nature 414, 77–81 (2001).
Akeda, Y. & Galan, J.E. Chaperone release and unfolding of substrates in type III secretion. Nature 437, 911–915 (2005).
Wulff-Strobel, C.R., Williams, A.W. & Straley, S.C. LcrQ and SycH function together at the Ysc type III secretion system in Yersinia pestis to impose a hierarchy of secretion. Mol. Microbiol. 43, 411–423 (2002).
Lara-Tejero, M., Kato, J., Wagner, S., Liu, X. & Galan, J.E. A sorting platform determines the order of protein secretion in bacterial type III systems. Science 331, 1188–1191 (2011).
Pettersson, J. et al. Modulation of virulence factor expression by pathogen target cell contact. Science 273, 1231–1233 (1996).
Kosarewicz, A., Konigsmaier, L. & Marlovits, T.C. The blueprint of the type-3 injectisome. Phil. Trans. R. Soc. Lond. B 367, 1140–1154 (2012).
Cornelis, G.R. & Van Gijsegem, F. Assembly and function of type III secretory systems. Annu. Rev. Microbiol. 54, 735–774 (2000).
Geibel, S., Procko, E., Hultgren, S.J., Baker, D. & Waksman, G. Structural and energetic basis of folded-protein transport by the FimD usher. Nature 496, 243–246 (2013).
Lee, V.T. & Schneewind, O. Yop fusions to tightly folded protein domains and their effects on Yersinia enterocolitica type III secretion. J. Bacteriol. 184, 3740–3745 (2002).
Johnsson, N. & Varshavsky, A. Ubiquitin-assisted dissection of protein transport across membranes. EMBO J. 13, 2686–2698 (1994).
Collazo, C.M. & Galan, J.E. Requirement for exported proteins in secretion through the invasion-associated type III system of Salmonella typhimurium. Infect. Immun. 64, 3524–3531 (1996).
Sukhan, A., Kubori, T., Wilson, J. & Galan, J.E. Genetic analysis of assembly of the Salmonella enterica serovar Typhimurium type III secretion-associated needle complex. J. Bacteriol. 183, 1159–1167 (2001).
Wagner, S. et al. Organization and coordinated assembly of the type III secretion export apparatus. Proc. Natl. Acad. Sci. USA 107, 17745–17750 (2010).
Diepold, A., Wiesand, U. & Cornelis, G.R. The assembly of the export apparatus (YscR,S,T,U,V) of the Yersinia type III secretion apparatus occurs independently of other structural components and involves the formation of an YscV oligomer. Mol. Microbiol. 82, 502–514 (2011).
Jung, T. & Grune, T. Structure of the proteasome. Prog. Mol. Biol. Transl. Sci. 109, 1–39 (2012).
Rüßmann, F. et al. Folding of large multidomain proteins by partial encapsulation in the chaperonin TRiC/CCT. Proc. Natl. Acad. Sci. USA 109, 21208–21215 (2012).
Kubori, T., Sukhan, A., Aizawa, S.I. & Galan, J.E. Molecular characterization and assembly of the needle complex of the Salmonella typhimurium type III protein secretion system. Proc. Natl. Acad. Sci. USA 97, 10225–10230 (2000).
Müller, S.A. et al. Double hexameric ring assembly of the type III protein translocase ATPase HrcN. Mol. Microbiol. 61, 119–125 (2006).
Abrusci, P. et al. Architecture of the major component of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 20, 99–104 (2013).
Kaniga, K., Bossio, J.C. & Galan, J.E. The Salmonella typhimurium invasion genes invF and invG encode homologues of the AraC and PulD family of proteins. Mol. Microbiol. 13, 555–568 (1994).
Schraidt, O. et al. Topology and organization of the Salmonella typhimurium type III secretion needle complex components. PLoS Pathog. 6, e1000824 (2010).
Marlovits, T.C. et al. Assembly of the inner rod determines needle length in the type III secretion injectisome. Nature 441, 637–640 (2006).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Fernández, J.J., Luque, D., Caston, J.R. & Carrascosa, J.L. Sharpening high resolution information in single particle electron cryomicroscopy. J. Struct. Biol. 164, 170–175 (2008).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Resch, G.P., Brandstetter, M., Konigsmaier, L., Urban, E. & Pickl-Herk, A.M. Immersion freezing of suspended particles and cells for cryo-electron microscopy. Cold Spring Harb. Protoc. 2011, 803–814 (2011).
Mastronarde, D.N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J.R., Mastronarde, D.N. & McIntosh, J.R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Acknowledgements
We would like to thank M. Brunner for the establishment of automated image data acquisition; O. Schraidt for discussions at early phases of the project; the EM, MS, bioinformatics and sequencing facilities at the Vienna Biocenter Campus for technical support; and members of the Marlovits laboratory for critical reading of the manuscript. We further would like to thank J.E. Galan (Yale University) and members of his laboratory for continuous discussion, generous support and materials. This work was supported by a PhD fellowship from Boehringer Ingelheim Fonds (to L.K.). Research in the Marlovits laboratory is generously supported by the Research Institute of Molecular Biotechnology (Austrian Academy of Sciences) and the Research Institute of Molecular Pathology and through a research grant from the Zentrum für Innovation and Technologie (Center for Molecular and Cellular Nanostructure (CMCN Vienna)) and the Oberösterreichische Landesbank Raika (no. 5000).
Author information
Authors and Affiliations
Contributions
J.R. and L.K. cloned constructs and performed biochemical assays and purifications. J.R. collected cryo-EM data for single particles, and L.K. collected and processed cryo-electron tomography data. T.C.M. analyzed and refined EM data and wrote the manuscript. All authors discussed the results and commented on the manuscript. T.C.M. directed and supervised the project.
Corresponding author
Ethics declarations
Competing interests
J.R., L.K. and T.C.M. are inventors on a patent application providing a cell that fully or partially secretes a protein of interest via a T3SS.
Integrated supplementary information
Supplementary Figure 1 Rationale of the design of substrates for the substrate-trapping experiment.
Newly designed substrates are based on the natural effector protein SptP. (a) Scheme of the concatemeric design of SptP3-GFP with three effector domains (GFP green fluorescent protein). (b) Model of substrates entering the needle complex with the N-terminal region first (left panel) at the cytoplasmic site and exiting at the extracellular tip of the needle filament. Complete transport is prohibited by the C-terminally fused GFP. As a consequence, designed substrates may then become trapped within injectisomes with the majority of the substrate being located within, and the N- and the C-terminal ends being located outside needle complexes. (c) Table of designed substrates used in this study (cbd = chaperone binding domain; Ub = ubiquitin).
Supplementary Figure 2 In situ visualization of active type 3 secretion systems by cryo-electron tomography.
(a) Reconstituted tomogram of Salmonella typhimurium cells containing T3SSs: SB905 cells containing wild type T3SSs (“Substrate-free”, WT) (left panel), and SB905 cells expressing SptP3-GFP (“Substrate-trapped”, +pSptP3-GFP) (right panel). Nine slices of the tomogram covering 179 Å were averaged (n = 9) (cy. = cytoplasm, ex. = extracellular space; bar = 100 nm). Dimensions of bases of injectisomes in situ or after purification are similar between each other and during substrate transport. (b) Upper panel: individual slices through a zoomed-in view of the tomogram containing membrane embedded, substrate-free T3SSs. T3SSs are displayed from left to right (bar = 50 nm), traversing through z-axis from top to bottom (voxel size 19.88 Å) Lower panel: individual slices through a zoomed-in view of the tomogram containing membrane embedded, substrate-trapped T3SSs. T3SSs are displayed from left to right (bar = 50 nm), traversing through z-axis from top to bottom (voxel size 19.88 Å).
Supplementary Figure 3 Designed substrates are stably bound to needle complexes.
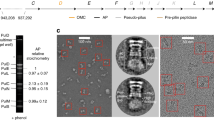
(a) Needle complexes (NCs) were purified from Salmonella strain SB905 as well as from strains expressing SptP1-GFP or SptP3-GFP. Substrates co-purified with NCs in CsCl gradient fractions. Top panel: Coomassie stained gels (4-20% SDS-polyacrylamide) of individual CsCl fractions. Lower panels: Western blotting of corresponding samples (rabbit anti-NC; rabbit anti-SptP, and mouse anti-FLAG). (b) Pooled CsCl fractions containing NCs (WT) or NCs and substrates (SptP1-GFP, SptP3-GFP) were subsequently separated on sucrose gradients. Western blotting results showed that designed substrates (SptP1-GFP, SptP3-GFP) are stably associated into NCs. Isolation of WT NCs showed no co-migration of natural substrates. (c) Peak fractions of sucrose gradients of w.t., and substrate-bound complexes (+pSptP1-GFP and +pSptP3-GFP) are negatively stained (2% PTA) for analysis by transmission electron microscopy: In total 27% of NCs containing the substrate SptP3-GFP displayed a clearly visible needle-tip density (ntotal= 298; ntip-density = 83). In contrast, all (100%) WT (substrate-free) NCs (ntotal= 287) were free of a visible density at the needle tip.
Supplementary Figure 4 Visibility of substrate domains from SptP3-GFP bound to needle complexes.
(a) Flexibility of emerging substrate at the tip of needle complexes. Collection of class averages of immuno-purified, substrate-trapped (SptP3-GFP) needle complexes. Class averages were calculated from negatively stained (2% (w/v) PTA) needle complexes. The different position of the needle-tip density comprising the emerging substrate (SptP3-GFP) and its diffuse character indicated a high flexibility and/or folding state of this domain. (b) Localization of the C-terminal substrate domain in substrate-trapped injectisomes Immuno-labeling (anti-GFP-gold) of substrate-free and substrate-trapped (SptP3-GFP) needle complexes on the electron microscopy grid. Highly specific labels close to the basal site of needle complexes could only be found in the substrate-trapped sample (2% (w/v/) PTA). (c) Sketch of a selection of various possible substrate-trapped injectisomes. The substrate might be trapped at various vertical positions and the C-terminally fused GFP in SptP3-GFP can be present at various horizontal positions relative to the needle complex (cbd = chaperone binding domain; N-term = N-terminal region; GFP = green fluorescent protein) (d) GFP (encircled with orange dots) is visible in the difference volume by lowering the threshold for the volume display (34.3 to 26.3).
Supplementary Figure 5 Single-particle analysis of substrate-free and substrate-trapped needle complexes.
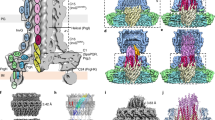
(a) Difference volume from substrate-trapped and substrate free needle complexes reconstructed without applying symmetry (C1). Surface views of half-sectioned and 15 degrees tilted three-dimensional volume of substrate-free (gray) and difference volume of substrate-trapped (blue) needle complexes reconstructed without applying symmetry (C1). (Difference corresponding to presence of substrate (Δsub); difference observed at the larger, inner ring-1, (ΔIR1)). Note, that the length of the extracellular needle filament is approximately 500 Å, however, due to a smaller boxsize and additional masking used during image data processing and reconstruction, the displayed needle filament is truncated. For comparison, entire isolated needle complexes are shown in Fig. 2b. (b) Resolution assessment by Fourier Shell Correlation. Upper panel: The resolution of substrate-free and substrate-trapped needle complexes reconstituted without symmetry (C1) corresponds to approximately 12 Angstroem (FSC=0.5). Lower panel: The resolution of substrate-free and substrate-trapped needle complexes reconstituted without symmetry (C1) and subsequently three-fold symmetrized (C3) corresponded to approximately 10 Angstroem measured at 0.5 FSC.
Supplementary Figure 6 Only minor structural changes are evident in substrate-free and substrate-trapped needle complexes.
Surface views of half-sectioned three-dimensional volume of substrate-free (gray) and substrate-trapped (yellow) needle complexes show only minor differences at the cup and socket region of the export apparatus (gray circle). The inner rod is substantially better defined in reconstructions obtained from substrate-free (gray) than in substrate-trapped (yellow) needle complexes, which could indicate a higher flexibility of the inner rod during substrate transport.
Supplementary Figure 7 Comparison of needle lengths of substrate-free (WT) and substrate-trapped (+pSptP3-GFP) needle complexes.
(a,b) Substrate-trapped sucrose-gradient purified needle complexes displayed a broader distribution of needle lengths than substrate-free (WT) needle complexes. (The most frequent needle length for substrate-trapped NCs was 41.4 nm, and 29.9 nm for substrate-free NCs, respectively.) This suggested that the substrate-trapped sample was composed of a mixture of empty and substrate-trapped needle complexes and indicated that substrate-containing complexes indeed had longer needles. However, the molecular mechanism leading to this observation is unknown, but may be explained by an increased stability of the needle filament in the presence of substrate during the purification. (Box plot: the left and right border of the box represented the first and third quartiles, and the band inside the box the second quartile (median).
Supplementary Figure 9 Validation of polyclonal antibodies.
(a) anti-SptP (b) anti-SipA
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Table 1 (PDF 16142 kb)
Cryo-electron tomography of wild-type Salmonella cells (substrate free).
The movie file shows a cryo electron tomogram of wild type Salmonella typhimurium after osmotic shock. (AVI 3438 kb)
Cryo-electron tomography of substrate-trapped Salmonella cells.
The movie file shows a cryo electron tomogram of wild type Salmonella typhimurium expressing SptP3-GFP after osmotic shock. (AVI 1530 kb)
Rights and permissions
About this article
Cite this article
Radics, J., Königsmaier, L. & Marlovits, T. Structure of a pathogenic type 3 secretion system in action. Nat Struct Mol Biol 21, 82–87 (2014). https://doi.org/10.1038/nsmb.2722
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2722
This article is cited by
-
Construction of a constitutively active type III secretion system for heterologous protein secretion
Applied Microbiology and Biotechnology (2023)
-
Substrate-engaged type III secretion system structures reveal gating mechanism for unfolded protein translocation
Nature Communications (2021)
-
Salmonella-based platform for efficient delivery of functional binding proteins to the cytosol
Communications Biology (2020)
-
The Rich Tapestry of Bacterial Protein Translocation Systems
The Protein Journal (2019)
-
Flagella-mediated secretion of a novel Vibrio cholerae cytotoxin affecting both vertebrate and invertebrate hosts
Communications Biology (2018)