Key Points
-
The gastrointestinal tract (GIT) and respiratory tract, although separate organs, are part of a shared mucosal immune system termed the gut–lung axis.
-
The microbiota of the GIT and the respiratory tract are involved in the gut–lung axis, influencing immune responses both locally and at distant sites.
-
Current research has identified specific bacterial taxa, their components and metabolites that can influence host immunity.
-
With greater knowledge of the gut–lung axis and microbial influences of immunity, advances have been made in understanding the role of the microbiota in respiratory diseases, such as asthma, chronic obstructive pulmonary disease (COPD) and respiratory infection.
-
This newfound understanding has created several possible therapeutic strategies for the treatment or prevention of acute and chronic respiratory diseases. However, several technical challenges and unanswered questions remain.
Abstract
The microbiota is vital for the development of the immune system and homeostasis. Changes in microbial composition and function, termed dysbiosis, in the respiratory tract and the gut have recently been linked to alterations in immune responses and to disease development in the lungs. In this Opinion article, we review the microbial species that are usually found in healthy gastrointestinal and respiratory tracts, their dysbiosis in disease and interactions with the gut–lung axis. Although the gut–lung axis is only beginning to be understood, emerging evidence indicates that there is potential for manipulation of the gut microbiota in the treatment of lung diseases.
Similar content being viewed by others
Main
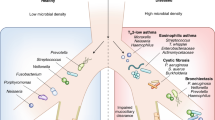
Chronic lung diseases, such as asthma and chronic obstructive pulmonary disease (COPD), are common and often occur together with chronic gastrointestinal tract (GIT) diseases, such as inflammatory bowel disease (IBD) or irritable bowel syndrome (IBS)1,2. Up to 50% of adults with IBD and 33% of patients with IBS have pulmonary involvement, such as inflammation or impaired lung function, although many patients have no history of acute or chronic respiratory disease3,4. Furthermore, patients with COPD are 2–3 times more likely to be diagnosed with IBD4. Individuals with asthma have functional and structural alterations in their intestinal mucosa, and patients with COPD typically have increased intestinal permeability2,5. Although the mature GIT and respiratory tract have different environments and functions, they have the same embryonic origin and, consequently, have structural similarities. Thus, it is not surprising that the two sites might interact in health and disease (Fig. 1); however, the underlying mechanisms are not well understood.
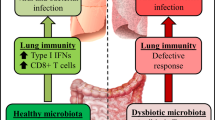
A healthy intestinal microbiota maintains homeostatic local immune responses through the exposure of structural ligands (for example, lipopolysaccharide (LPS) and/or peptidoglycan) and secreted metabolites (for example, short-chain fatty acids (SCFAs)). Invading microorganisms and absorbed metabolites influence circulating lymphocytes and contribute to the regulation of systemic immune responses. When the gut microbiota is disturbed, for example, during infection or antibiotic exposure, the normal microbiota-derived signals are altered, which leads to a changed immune response. In early life, when the immune system is still developing, this disturbance can substantially alter the way in which the immune system perceives its surroundings in later life, which can lead to the development of chronic inflammatory disorders in the gut and lung. In adulthood, dysbiosis of the gut microbiota, for example, through exposure to cigarette smoke, can cause systemic inflammation and an outgrowth of opportunistic pathogens, which can lead to chronic inflammation at distal sites. Although the specific taxa, ligands, metabolites and/or host responses may differ in specific disease situations, these broad principles outline the role of the microbiota in gut–lung crosstalk.
An emerging area of intense interest is the influence of the host-associated microbiota on local and systemic immunity. This is exemplified in germ-free mice, which lack an appropriately developed immune system and show mucosal alterations, both of which can be restored through colonization with gut microbiota6,7. The microbiota changes over time from birth, to adulthood and into old age, and in response to environmental factors, such as diet, and drug and environmental exposures8.
In this ever-expanding field, researchers are now investigating how the local microbiota influences immunity at distal sites, in particular how the gut microbiota influences other organs, such as the brain, liver or lungs. This has led to the coining of terms such as the 'gut–brain axis' and 'gut–lung axis'. For example, antibiotic-induced alterations in the gut microbiota in early life increase the risk of developing allergic airway disease9,10,11,12; such findings add to our understanding of the links between exposure to microorganisms and allergy and autoimmunity (Box 1). The mechanisms by which the gut microbiota affects the immune responses in the lungs, and vice versa, are being uncovered, but many questions remain. In this Opinion article, we summarize the emerging role of the microbiota in the gut–lung axis, highlighting gaps in our knowledge and the potential for therapeutic intervention.
Microbiota of the healthy gut and lungs
The GIT remains, by far, the best-studied host-associated microbial ecosystem, partly owing to its abundance of microorganisms and partly because the microbiota can be profiled through faeces, which is easily obtainable. Both the abundance and diversity of the commensal microbiota generally increase along the GIT, and there are site-specific variations in the mucosa and the lumen13,14. These differences are governed by the prevailing environment, including pH, the concentration of bile acids, digesta retention time, mucin properties and host defence factors15. Despite these variations, the GIT is dominated by members of only four bacterial phyla, Firmicutes, Bacteroidetes, Proteobacteria and Actinobacteria; with lesser and sporadic representation of other phyla, including Fusobacteria, Verrucomicrobia and Spirochaetes. This 'core' gut microbial community comprises up to 14 bacterial genera and 150 bacterial 'species', many of which have not yet been cultured16,17,18.
We are beginning to understand the lung microbiota through programmes such as the Lung HIV Microbiome Project, which is a multi-centre study that examines both individuals who are infected with HIV and uninfected individuals who have varying histories of lung and/or respiratory disease19. The lungs have a large surface area with high environmental exposure and are equipped with effective antimicrobial defences. Healthy lungs were long considered to be sterile; however, the advent of culture-independent approaches for microbial community profiling has resulted in the detection of microbial DNA in the lungs of healthy individuals19,20. These microorganisms probably reached the lungs from the oral cavity through microaspiration, as the taxonomic profiles of the two sites were similar19,20. Compared with surrounding sites, the lungs had a decreased abundance of Prevotella-affiliated taxa and an enrichment of Proteobacteria, specifically Enterobacteriaceae, Ralstonia spp., and Haemophilus spp.19, which may have resulted from host immunity and environment, such as redox state and oxygen availability. The lung microbiota might not be resident in healthy individuals, but rather transiently recolonized through microaspiration and breathing. The lungs have a comparatively low microbial biomass and remarkably similar composition to adjacent sites, even though the lungs are continuously exposed to entering microorganisms and their environmental conditions differ vastly from other body sites. These observations support the hypothesis that entry and selective elimination of a transient microbiota is the major determinant of microbial composition in the lungs, rather than resident and viable microorganisms. This does not negate the importance of host–microorganism interactions in the lungs, as evidenced by correlations between the composition of microbial communities and pulmonary inflammation and disease21. Rather, it highlights the delicate balance between microbial exposure and elimination; the possibility of dysbiosis at oral sites preceding and/or causing dysbiosis in the lungs and contributing to disease pathogenesis19; and the importance in distinguishing whether microbial DNA that is detected by culture-independent techniques is truly representative of viable bacteria in the lungs22. Technical challenges, such as low microbial biomass and bronchoscope contamination, constant seeding from oral and GIT sites, and mucociliary and immune clearance, have hindered the identification of a viable and resident, or a transiently recolonizing, microbiota in the lungs, as well as further research into host–microorganism interactions. Novel methods of sampling tissue with minimal contamination23, longitudinal studies to identify temporal changes in the microbiota, and the increasing use of metagenomic analyses to facilitate the cultivation of fastidious bacterial species24, will provide a clearer picture of the role of the respiratory microbiota and enable the better design of interventional studies to develop a more complete understanding of host–microorganism interactions in the lungs.
Interactions between the gut and lungs
Interactions of microorganisms between the sites. The epithelial surfaces of the GIT and respiratory tract are exposed to a wide variety of microorganisms; ingested microorganisms can access both sites and the microbiota from the GIT can enter the lungs through aspiration. Both the gut and respiratory mucosa provide a physical barrier against microbial penetration, and colonization with the normal microbiota generates resistance to pathogens; for example, through the production of bacteriocins15. Furthermore, a rapidly expanding collection of commensal gut bacteria, including segmented filamentous bacteria (SFB), Bifidobacterium spp. and members of the colonic Bacteroides genus, induce the production of antimicrobial peptides, secretory immunoglobulin A (sIgA) and pro-inflammatory cytokines. Non-pathogenic Salmonella strains downregulate inflammatory responses in GIT epithelial cells by inhibiting the ubiquitylation of nuclear factor-κB (NF-κB) inhibitor-α (IκBα)25, whereas some Clostridium spp. promote anti-inflammatory regulatory T cell (Treg cell) responses26. In the respiratory tract, Streptococcus pneumoniae and Haemophilus influenzae synergistically activate host p38 mitogen-activated protein kinase (MAPK) in a Toll-like receptor (TLR)-independent manner to amplify pro-inflammatory responses27. Conversely, non-pathogenic S. pneumoniae and other bacteria and their components can suppress allergic airway disease by inducing Treg cells28,29,30,31. In recipients of lung transplants, the microbiota of the respiratory tract alters immunity in the lungs. Firmicutes-dominated and Proteobacteria-dominated dysbiosis were associated with the expression of inflammatory genes in pulmonary leukocytes, whereas Bacteroidetes-dominated dysbiosis was linked to a gene expression profile that is characteristic of tissue remodelling32. In both cell culture32 and animal models33, the inflammatory response that is induced by pathogenic species is larger than the response induced by commensal microorganisms, which indicates that a diverse lung microbiota protects against pathology by 'diluting' the more pro-inflammatory stimuli of pathogens. Although the transfer of microorganisms from faecal suspensions has been used to determine the role of the gut microbiota, such techniques have not yet been used to transfer the respiratory microbiota between animals, which limits our understanding of their roles.
Several studies have shown the effects of GIT colonization with orally administered bacteria on lung function. Oral gavage of faecal suspensions in mice that were first treated with antibiotics provided short-term improvements in some, but not all, measures in an S. pneumoniae infection model34.Although the nature of this 'gut–lung axis' has been challenged owing to the potential confounding effects of faecal administration by oral gavage and antibiotic use35, the concept warrants systematic and controlled evaluation. In infants, the composition of the gut microbiota and caesarean section have been linked to atopic manifestations, and colonization by Clostridium difficile at one month of age was associated with wheeze and eczema throughout early life, and with asthma after 6–7 years36. Positive associations between the presence of 'beneficial' bacteria, such as Bifidobacterium longum, in the gut and a lower incidence of asthma have also been identified37, although larger and longer studies are required to evaluate these associations.
Considerable evidence suggests that host epithelia and other structural and immune cells assimilate information directly from microorganisms and from the concomitant local cytokine response to adjust inflammatory responses, and that this shapes immune responses at distal sites, such as the lungs38,39 (Fig. 2). There is less evidence of the direct transfer of microorganisms between sites, although the translocation of bacteria from the GIT to the lungs has been observed in sepsis and acute respiratory distress syndrome, in which barrier integrity is compromised40. In addition, some environmental factors, such as dietary fibre, can produce similar changes in the microbiota of the GIT and the lungs39. Whether these result from diet-driven changes in microbial metabolites, changes in innate immune responses or a combination of both remains to be determined.
The gut and respiratory tract epithelia have substantial differences in functional purpose and exist in different environments; however, they retain some anatomical similarities. Both are derived from the endoderm and consist of columnar epithelial cells with projections of microvilli (gut) or cilia (respiratory tract) that function as a physical barrier and as sentinels for the immune system in conjunction with associated lymphoid tissue. Both secrete mucus through goblet cells as well as secretory immunoglobulin A (sIgA; although less in the lung). The alveoli that are found in terminal airways in the lungs differ substantially, consisting of squamous epithelial cells that either secrete surfactant (type 2 alveolar cells) or function in gas exchange (type 1 alveolar cells). However, the similarities end here: the intestinal lumen is an oxygen-poor environment and functions to digest food and absorb nutrients. The movement of matter is unidirectional (mouth to anus), with the exception of reflux or vomiting. Furthermore, the pH, enzyme presence and structure vary along the gastrointestinal tract (GIT). By contrast, the respiratory tract and alveoli are oxygen-rich and movement is bidirectional (inhalation and exhalation). The temperature in the gut is relatively uniform at 37 °C, whereas the temperature in the respiratory tract differs depending on the proximity to the pharynx. Thus, it is unsurprising that the microbial life in each environment is distinct. Changes in diet and exposure to therapeutics and environmental particulates can directly affect the composition of the microbiota. Both the gut and lungs are able to influence each other's immune responses. Dendritic cells in the intestines and respiratory tract, and macrophages in the lungs, sample antigens in the lumen. Lymphocytes in the associated lymphoid tissues circulate through the lymphatic system to affect systemic immunity. Bacteria from the gut can travel to the lungs through aspiration of vomit or oesophageal reflux. During times of dysbiosis, disturbed epithelial integrity may enable bacteria and their components and metabolites to enter the circulatory system, which can cause systemic inflammation.
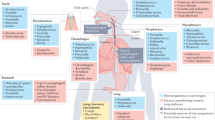
Microbial species-specific effects on host immunity. The crucial role of the microbiota in lung homeostasis and immunity is demonstrated by the poor outcomes of germ-free mice that were exposed to acute infections41 and their susceptibility to allergic airway disease42. Current research is assessing the effects of selected members of the commensal gut microbiota on systemic immunity, including in the lungs, as well as the use of probiotics and prebiotics to prevent and treat acute and chronic pulmonary disease (Fig. 3). For example, SFB in the gut, when present naturally or introduced by probiotic dosing or co-housing of mice, stimulated pulmonary T helper 17 (TH17) responses and protection from S. pneumoniae infection and mortality43. Intriguingly, a respiratory microbiota enriched with oral-related taxa, such as Prevotella spp., Rothia spp. and Veillonella spp., was associated with TH17-mediated immunity in the lungs of healthy human hosts21. Whether these links are correlative or causative remain unclear. Exposure of mice to dog-associated house dust altered the caecal microbiota, and, in particular, increased the abundance of Lactobacillus johnsonii and other Firmicutes-related lineages, such as species in the Peptococcaceae and Lachnospiraceae families44. Mice that were either exposed to dog dust or inoculated with L. johnsonii showed decreases in respiratory tract TH2 cytokine responses, and L. johnsonii treatment protected against exposure to respiratory syncytial virus and allergens such as ovalbumin. Other examples of microbial influences on host immunity include the ability of various Bacteroides spp. to expand Treg cell populations or bias the TH1/TH2 phenotype in either direction in a strain-specific manner, or the suppression of host inflammatory responses by the common bacterial metabolites short-chain fatty acids (SCFAs), which act through free fatty acid (FFA) receptors and/or epigenetic regulation of immune cells45.
Secreted and structural components of microorganisms can influence the host immune response both locally and at distal sites. Microbial metabolites, such as short chain fatty acids (SCFAs), bind to free fatty acid receptors or promote epigenetic changes in host leukocytes, which induce anti-inflammatory responses and decrease inflammation. Virulence factors from pathogenic bacteria, such as Helicobacter pylori or Bacteroides fragilis, can downregulate host immune responses, whereas structural components from commensal bacteria can influence inflammatory responses through the activation of pattern recognition receptors. GGT, γ-glutamyl transpeptidase; IL-10, interleukin-10; LPS, lipopolysaccharide; LTA, lipoteichoic acid; NAP, neutrophil-activating protein; PSA, polysaccharide A; Treg cell, regulatory T cell; UreB, urease subunit-β; VacA, vacuolating cytotoxin A.
In related human studies, seropositivity to the gut-specific pathogen Helicobacter pylori, in particular, cytotoxin-associated gene A positive (cagA+) strains, has long been linked with decreased incidence of asthma and allergy46,47,48. Conversely, two recent meta-analyses suggest that infection with H. pylori is positively associated with increased incidence of COPD and other chronic bronchial diseases49. Although these differences might be partly attributable to genetic, environmental and lifestyle factors, these findings raise the possibility that systemic immune responses that are triggered by H. pylori might have different roles in the aetiology of different lung disorders. Strain variations, in addition to the expression of cagA, might also affect Treg cell responses50.
Clearly, the incredible diversity and abundance of species in the gut microbiota results in many immunomodulatory signals, which have considerable combined effects on host health. Although much has been uncovered about the activity of specific bacterial species, current research has only just begun to assess the structure–function relationships of the gut and lung microbiota with host immunity.
Components and metabolites of the gut microbiota that influence the lung. Early studies showed that germ-free mice have reduced responsiveness to lipopolysaccharide (LPS)-induced pathology and that this oral tolerance to microbial components was due to interleukin-10 (IL-10)-mediated hypo-responsiveness; however, subsequent exposure to LPS was no longer tolerated and the immune response became similar to that seen in conventional mice51,52. Furthermore, a robust response to LPS by colonic macrophages could be restored by commensal microorganisms53.
Bacterial components can also have anti-inflammatory effects, which attenuate GIT pathology. Polysaccharide A (PSA) from Bacteroides fragilis induces the production of IL-10 by T cells and protects against intestinal inflammatory disease caused either chemically or by infection with Helicobacter hepaticus54. Sphingolipids, which are naturally occurring cell membrane components of many anaerobic gut genera including Bacteroides, decrease the number of invariant natural killer T cells in the colon — cells that have been implicated in the development of colitis55. The best-studied metabolites, SCFAs, are by-products of the microbial fermentation of dietary fibre, have anti-inflammatory properties, are a source of energy for colonocytes, and regulate fatty acid and lipid synthesis in the host56.
Much less is known about the influence of microbial components and metabolites at other sites, including the lungs. Decreases in Faecalibacterium spp., Lachnospira spp., Veillonella spp. and Rothia spp. in the gut, and the urine levels of some microbial bile acid metabolites correlate with the development of atopic wheeze in children, although whether they are a cause or a consequence of wheeze is not known12. Oral administration of SCFAs has been shown experimentally to alleviate allergic airway disease39,57. Microbial components and metabolites have been implicated in other disorders, such as tryptophan in brain health, PSA in disorders of the central nervous system and trimethylamine N-oxide in atherosclerosis, which further highlights their importance in extra-intestinal environments55. In studies of other diseases, Bacteroides spp. were associated with early-onset autoimmune diseases, which may be a consequence of potent activation of immunity by LPS produced by these bacteria58.
Gut microbiota and lung diseases
Asthma. An increased risk of asthma has been connected to the disruption of the gut microbiota in early life (Box 1), and several studies have sought to characterize the precise microbial constituents that are associated with the development of the disease in infants.
The overall composition of the gut microbial community is not altered in infants at risk of the development of asthma, but subtle transient changes in select taxa can be detected in the first few months of life12,59. Increased risk of asthma has been associated with an increase in the abundance of B. fragilis and total anaerobes in early life60, as well as reduced microbial diversity59 and decreases in Escherichia coli61 and the relative abundances of Faecalibacterium spp., Lachnospira spp., Rothia spp. and Veillonella spp.12, although these findings were not consistent across all studies. In addition, although models of allergic airway disease support the existence of a critical developmental window early in life42,62, only one study has provided direct evidence that restoring the altered gut microbiota through probiotic treatment can decrease susceptibility to asthma12.
Similarly, in adults, the overall composition of the faecal microbiota in individuals with allergic asthma does not differ from healthy individuals63,64. There are taxa-specific differences, such as the enrichment of Bifidobacterium adolescentis, which negatively correlated with the time since asthma diagnosis63. Interestingly, heat-inactivated Bifidobacterium spp. that were isolated from infants with allergic asthma induced larger pro-inflammatory responses than those isolated from healthy individuals65.
There are several proposed mechanisms through which the gut microbiota can attenuate the risk of developing asthma. Infants who were at risk of developing asthma had decreased levels of LPS in their faeces12, whereas PSA from B. fragilis protected against the development of allergic airways disease in mice by inducing IL-10 responses in T cells66. H. pylori alleviated murine allergic airway disease in several ways, namely through the direct activation of Treg cells by neutrophil-activating protein67, or indirectly through urease subunit-β, which promotes tolerogenic reprogramming of dendritic cells68. In addition, γ-glutamyl transpeptidase and vacuolating cytotoxin from H. pylori altered dendritic cell function, but did not require Treg cells to alleviate symptoms69. Commensal bacteria can also influence the development of asthma through the production and secretion of metabolites, specifically SCFAs. The risk of asthma in infants was associated with a decrease in the concentration of acetate in faeces12 and inversely correlated with serum acetate concentrations in their mothers when they were pregnant57. A high-fibre diet, which increased levels of SCFAs in serum and faeces, protected mice against the development of asthma symptoms, a phenomenon that could be replicated through the direct administration of acetate or propionate before disease onset to promote tolerogenic immune responses in dendritic cells and Treg cells39,57. The benefits of a high-fibre diet were associated with a decreased ratio of Firmicutes/Bacteroidetes and an enrichment of Bacteroidaceae in both the faeces and lungs, which highlights the necessity of investigating microbial communities at several body sites for a complete understanding of the influence of microorganisms on host health. These studies did not directly explore the relationship between the composition of the microbial communities at the two sites, or the relative importance of the gut or lung microbiota in protection against disease. Such studies would be valuable in determining which body site to target with therapeutic interventions. An important but understudied area is the role that interactions between microorganisms have in the development of asthma. For example, the loss of intestinal bacteria and the subsequent outgrowth of commensal fungi triggered prostaglandin E2-induced changes in alveolar macrophages and increased allergic airway inflammation70. Furthermore, gut helminth infection protected mice against allergic airway disease, which was associated with an increase in the abundance of Lachnospiraceae and other Clostridiales members, the production of SCFAs and subsequent robust Treg cell responses in the lungs71. Although the Treg cell-promoting capability of Clostridium spp. has previously been demonstrated in the colon26,72, it is increasingly being explored for the treatment of diseases at other body sites, including asthma73,74.
COPD. Respiratory microbiota research in COPD has assessed changes in the disease state and with smoke exposure, which is a major risk factor for the development of this disease. Interestingly, although the lung microbiota is similar in healthy smokers and non-smokers, the oral microbiota differs substantially between the two groups19. As enrichment of the lung microbiota with taxa from the oral cavity is associated with increased inflammation in smokers75, it is plausible that changes in the oral microbiota and a failure to effectively clear aspirated microorganisms contribute to disease development, and may help explain why only a subset of smokers develop COPD. Moreover, there are substantial differences between the lung microbiota of patients with COPD compared with 'healthy' smokers76,77, which led to the proposal that the respiratory microbiota may be useful in the early diagnosis of COPD. By contrast, no study to date has investigated changes in the gut microbiota of patients with COPD. Nevertheless, in 'healthy' smokers, the faecal microbiota is characterized by an increase in the abundance of Bacteroides–Prevotella78, and a decreased Firmicutes/Bacteroidetes ratio79 compared with non-smokers. These changes in the composition of the faecal microbiota have been associated with intestinal inflammation and IBD80,81. Smokers also have a decreased abundance of Bifidobacterium spp.79,82, and hence may lose the anti-inflammatory effects that are often associated with this genus.
The causes of smoking-associated changes in the composition of the gut microbiota are probably a combination of environmental, host and microbial changes, such as intestinal and immune disruption, impaired clearance of pathogens83,84, acidification of gastric contents85 and ingestion of bacteria that are present in cigarettes86. Furthermore, cigarette smoke can directly affect the virulence of bacteria87 and fungi88, as well as alter the growth and exopolysaccharide structure of known gut bacteria, such as Bifidobacterium animalis89, which may contribute to dysbiosis. Even after the cessation of smoking, many of the changes that cause dysbiosis persist for prolonged periods of time, and thus any therapeutic intervention to restore the gut microbiota may potentially require repeated administration to prevent relapse.
In the absence of longitudinal or interventional studies, it is difficult to ascertain whether changes in the gut or respiratory microbiota are a cause or a consequence of COPD. Most likely, both are true and operate simultaneously or at different stages of disease. Exposure to environmental stimuli and the onset of disease cause dysbiosis, which, in turn, probably contributes to disease progression. Moreover, defined probiotic use may benefit patients with COPD, particularly if used as an early preventive intervention. Oral administration of Lactobacillus casei improved the previously defective function of peripheral natural killer cells in adult male smokers90, whereas Bifidobacterium breve and Lactobacillus rhamnosus reduced lung pathology in a mouse model of COPD91 and reduced inflammatory responses in macrophages that were exposed to cigarette smoke extract in vitro92. Similarly, a diet that increased the production of SCFAs protected against elastase-induced inflammation and emphysema93. Although a causal relationship between SCFAs and protection in this study was not confirmed, both cigarette smoke94 and environmental particulate matter95 decreased SCFA concentrations in rodents, and cigarette smoke condensate decreased the production of SCFAs in vitro89. Furthermore, increased intestinal translocation of bacteria and their products occurred after exposure to particulate matter or the development of COPD2,95,96. Bacterial toxins, such as enterotoxin97 or LPS98, can contribute to the pathogenesis of COPD and microbiota-associated intestinal inflammation may become systemic and also contribute. The potential of SCFAs to improve intestinal barrier function may account for their benefits in animal models of COPD99,100, although this is yet to be explored in clinical studies.
Respiratory infections. The gut microbiota is broadly protective against respiratory infection, as its depletion or absence in mice leads to impaired immune responses and worsens outcomes following bacterial or viral respiratory infection34,41,101,102,103. Administration of SFB improved resistance to Staphylococcus aureus pneumonia43 and Bifidobacterium spp. protected against both bacterial104 and viral pulmonary infection in mice103,105. Lactobacillus spp. and Bifidobacterium spp.-based probiotics also improved the incidence and outcomes of respiratory infections in humans106,107,108,109.
Several aspects of experimental design influence the results of infection studies, including the route of administration of bacterial ligands102,110, the facility from which research animals are sourced43, the type of antibiotic used for the depletion of the microbiota62,102 and the infecting pathogen. For example, herpes simplex virus type 2 or Legionella pneumophila do not seem to be influenced by antibiotic-mediated depletion of the microbiota102.
Nevertheless, several important mechanisms by which the gut microbiota promotes the clearance of pathogens have been identified. Innate immune responses to bacteria in the lungs are greatly enhanced by exposure to nucleotide-binding oligomerization domain (NOD)-like receptor and TLR agonists in the GIT, which include peptidoglycan, LPS, lipoteichoic acid and CpG DNA41,101,110. Similarly, stimulation of TLRs by cell wall components and flagellin of gut bacteria is necessary for effective adaptive immune responses to influenza102,111, whereas the anti-inflammatory effects of oral SCFA administration are linked to reduced pulmonary pathology following both bacterial104,112 and viral113 infection in mice. However, the microbiota can also drive gut pathology in pulmonary infection. Influenza virus infection in mice increased the number of lung-derived CC-chemokine receptor 9 positive (CCR9+)CD4+ T cells, which preferentially migrate to the GIT under the guidance of C-C motif ligand 25 (CCL25) expressed on intestinal epithelial cells114. This resulted in the outgrowth of E. coli and the induction of aberrant TH17 responses and intestinal damage.
Conclusions and perspectives
Many studies have identified the presence of a lung microbiota in health and disease. However, we believe that the healthy lung microbiota may be transient and best described as a progression of taxa that are influenced by adjacent body sites and the external environment, rather than an actively reproducing core resident community. This is not down-playing the importance of a transient microbiota in healthy lungs, which could still have important roles in inflammatory responses whether viable or not. By contrast, the microbiota is more likely to be persistent and resident in the respiratory tract and lungs of individuals with respiratory disease, although whether it is a cause or a consequence of disease remains to be elucidated. Furthermore, the lung microbiota could affect, or be affected by, microorganisms or immune responses at distal sites.
The crosstalk between microorganisms and the host is complex and our current understanding of these interactions is in its infancy. It is unlikely that any one of these interactions is solely responsible for the functions of the microbiota, and alterations in any part of these relationships may be enough to affect health and disease. It is unclear whether changes in the microbiota at one site affect many distal sites equally, or whether these systemic effects might be specific to certain tissues. To date, no such broad study investigating these systemic widespread effects has been carried out.
Thus far, gut–lung microbiota studies have had two major limitations: the first is discerning causative over correlative effects and the second is timing. Most studies have been associative. Furthermore, culture-independent identification of microorganisms has not yet replaced the need to isolate and culture suspected opportunistic pathogens or probiotics to study their effects, and many members of the microbiota cannot be easily cultured. Thus, it is typically unclear whether changes that are observed in the microbiota are the cause or effect of disease. In the case of timing, most experimental data have described the effect of the gut microbiota on the development of lung disease and not on established lung disease. Longitudinal studies in humans and animals that associate changes in the microbiota with the severity of established chronic lung disease are required. Research into manipulations of the microbiota during lung disease is necessary to improve our understanding and inform the development of novel therapies (Box 2).
Increasingly, microbiota research is moving towards defining functional guilds. As taxonomic variation between sites and individuals is so large, and the microbiota consists of thousands of species, it is highly likely that there is redundancy between species in terms of their interactions with other microorganisms and in the metabolites that they produce. Thus, next-generation 'omics' approaches are required to identify functional guilds to aid in defining how the microbiota of the gut and the microbiota of the lungs interact with each other and influence health and disease.
In summary, the lung microbiota in a healthy individual may be transient and constantly re-seeded from the environment and cleared by the immune system, but may still influence health and disease. In respiratory diseases the lung microbiota probably becomes persistent and may be both a cause and a consequence of the disease, forming a pathogenic feedback loop. It is clear that bacterial components and metabolites in the gut and the lungs have the capacity to modulate systemic and local immunity, with specific taxa able to influence the pathogenesis of respiratory diseases, such as asthma, COPD and respiratory infections. Such relationships have been identified in other respiratory diseases, such as cystic fibrosis115, which, as a genetic disease, is a special case. Respiratory challenges with environmental factors such as pollution, cigarette smoke, antibiotics and diet, influence disease risk and probably drive pathogenesis through their ability to modulate the composition of the microbiota, although the mechanisms of these effects remain unknown. Further longitudinal studies and improved interventional experiments will help to elucidate the role of the microbiota and gut–lung crosstalk in respiratory disease, and will potentially lead to the identification of new and effective avenues for treatment.
References
Roussos, A., Koursarakos, P., Patsopoulos, D., Gerogianni, I. & Philippou, N. Increased prevalence of irritable bowel syndrome in patients with bronchial asthma. Respir. Med. 97, 75–79 (2003).
Rutten, E. P., Lenaerts, K., Buurman, W. A. & Wouters, E. F. Disturbed intestinal integrity in patients with COPD: effects of activities of daily living. Chest 145, 245–252 (2014).
Yazar, A. et al. Respiratory symptoms and pulmonary functional changes in patients with irritable bowel syndrome. Am. J. Gastroenterol. 96, 1511–1516 (2001).
Keely, S., Talley, N. J. & Hansbro, P. M. Pulmonary–intestinal cross-talk in mucosal inflammatory disease. Mucosal Immunol. 5, 7–18 (2012).
Vieira, W. A. & Pretorius, E. The impact of asthma on the gastrointestinal tract (GIT). J. Asthma Allergy 3, 123–130 (2010).
Wymore Brand, M. et al. The altered Schaedler flora: continued applications of a defined murine microbial community. ILAR J. 56, 169–178 (2015).
Al-Asmakh, M. & Zadjali, F. Use of germ-free animal models in microbiota-related research. J. Microbiol. Biotechnol. 25, 1583–1588 (2015).
Quercia, S. et al. From lifetime to evolution: timescales of human gut microbiota adaptation. Front. Microbiol. 5, 587 (2014).
Noverr, M. C., Falkowski, N. R., McDonald, R. A., McKenzie, A. N. & Huffnagle, G. B. Development of allergic airway disease in mice following antibiotic therapy and fungal microbiota increase: role of host genetics, antigen, and interleukin-13. Infect. Immun. 73, 30–38 (2005).
Russell, S. L. et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 13, 440–447 (2012).
Russell, S. L. et al. Perinatal antibiotic-induced shifts in gut microbiota have differential effects on inflammatory lung diseases. J. Allergy Clin. Immunol. 135, 100–109 (2015).
Arrieta, M. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Aguirre de Carcer, D. et al. Numerical ecology validates a biogeographical distribution and gender-based effect on mucosa-associated bacteria along the human colon. ISME J. 5, 801–809 (2011).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Buffie, C. G. & Pamer, E. G. Microbiota-mediated colonization resistance against intestinal pathogens. Nat. Rev. Immunol. 13, 790–801 (2013).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Ormerod, K. L. et al. Genomic characterization of the uncultured Bacteroidales family S24-7 inhabiting the guts of homeothermic animals. Microbiome 4, 36 (2016).
Morris, A. et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am. J. Respir. Crit. Care Med. 187, 1067–1075 (2013).
Bassis, C. M. et al. Analysis of the upper respiratory tract microbiotas as the source of the lung and gastric microbiotas in healthy individuals. mBio 6, e00037 (2015).
Segal, L. N. et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a TH17 phenotype. Nat. Microbiol. 1, 16031 (2016).
Rogers, G. B. et al. Assessing the diagnostic importance of nonviable bacterial cells in respiratory infections. Diagn. Microbiol. Infect. Dis. 62, 133–141 (2008).
Shanahan, E. R., Zhong, L., Talley, N. J., Morrison, M. & Holtmann, G. Characterisation of the gastrointestinal mucosa-associated microbiota: a novel technique to prevent cross-contamination during endoscopic procedures. Aliment. Pharmacol. Ther. 43, 1186–1196 (2016).
Pope, P. B. et al. Isolation of Succinivibrionaceae implicated in low methane emissions from Tammar wallabies. Science 333, 646–648 (2011).
Neish, A. S. et al. Prokaryotic regulation of epithelial responses by inhibition of IκB-α ubiquitination. Science 289, 1560–1563 (2000).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Ratner, A. J., Lysenko, E. S., Paul, M. N. & Weiser, J. N. Synergistic proinflammatory responses induced by polymicrobial colonization of epithelial surfaces. Proc. Natl Acad. Sci. USA 102, 3429–3434 (2005).
Preston, J. A. et al. Inhibition of allergic airways disease by immunomodulatory therapy with whole killed Streptococcus pneumoniae. Vaccine 25, 8154–8162 (2007).
Thorburn, A. N., Foster, P. S., Gibson, P. G. & Hansbro, P. M. Components of Streptococcus pneumoniae suppress allergic airways disease and NKT cells by inducing regulatory T cells. J. Immunol. 188, 4611–4620 (2012).
Thorburn, A. N. & Hansbro, P. M. Harnessing regulatory T cells to suppress asthma: from potential to therapy. Am. J. Respir. Cell Mol. Biol. 43, 511–519 (2010).
Preston, J. A. et al. Streptococcus pneumoniae infection suppresses allergic airways disease by inducing regulatory T-cells. Eur. Respir. J. 37, 53–64 (2011).
Bernasconi, E. et al. Airway microbiota determines innate cell inflammatory or tissue remodeling profiles in lung transplantation. Am. J. Respir. Crit. Care Med. http://dx.doi.org/10.1164/rccm.201512-2424OC (2016).
Larsen, J. M. et al. Chronic obstructive pulmonary disease and asthma-associated Proteobacteria, but not commensal Prevotella spp., promote Toll-like receptor 2-independent lung inflammation and pathology. Immunology 144, 333–342 (2015).
Schuijt, T. J. et al. The gut microbiota plays a protective role in the host defence against pneumococcal pneumonia. Gut 65, 575–583 (2016).
Dickson, R. P. & Cox, M. J. The premature invocation of a 'gut–lung axis' may obscure the direct effects of respiratory microbiota on pneumonia susceptibility. Gut http://dx.doi.org/10.1136/gutjnl-2016-311823 (2016).
van Nimwegen, F. A. et al. Mode and place of delivery, gastrointestinal microbiota, and their influence on asthma and atopy. J. Allergy Clin. Immunol. 128, 948–955 (2011).
Akay, H. K. et al. The relationship between bifidobacteria and allergic asthma and/or allergic dermatitis: a prospective study of 0–3 years-old children in Turkey. Anaerobe 28, 98–103 (2014).
Marsland, B. J., Trompette, A. & Gollwitzer, E. S. The gut–lung axis in respiratory disease. Ann. Am. Thorac. Soc. 12 (Suppl. 2), S150–S156 (2015).
Trompette, A. et al. Gut microbiota metabolism of dietary fibre influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Dickson, R. P. et al. Enrichment of the lung microbiome with gut bacteria in sepsis and the acute respiratory distress syndrome. Nat. Microbiol. 1, 16113 (2016).
Fagundes, C. T. et al. Transient TLR activation restores inflammatory response and ability to control pulmonary bacterial infection in germfree mice. J. Immunol. 188, 1411–1420 (2012).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
Gauguet, S. et al. Intestinal microbiota of mice influences resistance to Staphylococcus aureus pneumonia. Infect. Immun. 83, 4003–4014 (2015).
Fujimura, K. E. et al. House dust exposure mediates gut microbiome Lactobacillus enrichment and airway immune defense against allergens and virus infection. Proc. Natl Acad. Sci. USA 111, 805–810 (2014).
Samuelson, D. R., Welsh, D. A. & Shellito, J. E. Regulation of lung immunity and host defense by the intestinal microbiota. Front. Microbiol. 6, 1085 (2015).
Chen, Y. & Blaser, M. J. Inverse associations of Helicobacter pylori with asthma and allergy. Arch. Intern. Med. 167, 821–827 (2007).
Reibman, J. et al. Asthma is inversely associated with Helicobacter pylori status in an urban population. PLoS ONE 3, e4060 (2008).
Chen, Y. & Blaser, M. J. Helicobacter pylori colonization is inversely associated with childhood asthma. J. Infect. Dis. 198, 553–560 (2008).
Wang, F., Liu, J., Zhang, Y. & Lei, P. Association of Helicobacter pylori infection with chronic obstructive pulmonary disease and chronic bronchitis: a meta-analysis of 16 studies. Infect. Dis. (Lond.) 47, 597–603 (2015).
Hussain, K. et al. Helicobacter pylori-mediated protection from allergy is associated with IL-10-secreting peripheral blood regulatory T cells. Front. Immunol. 7, 71 (2016).
McGhee, J. R. et al. Lipopolysaccharide (LPS) regulation of the immune response: T lymphocytes from normal mice suppress mitogenic and immunogenic responses to LPS. J. Immunol. 124, 1603–1611 (1980).
Michalek, S. M., Kiyono, H., Wannemuehler, M. J., Mosteller, L. M. & McGhee, J. R. Lipopolysaccharide (LPS) regulation of the immune response: LPS influence on oral tolerance induction. J. Immunol. 128, 1992–1998 (1982).
Ueda, Y. et al. Commensal microbiota induce LPS hyporesponsiveness in colonic macrophages via the production of IL-10. Int. Immunol. 22, 953–962 (2010).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Sharon, G. et al. Specialized metabolites from the microbiome in health and disease. Cell Metab. 20, 719–730 (2014).
den Besten, G. et al. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 54, 2325–2340 (2013).
Thorburn, A. N. et al. Evidence that asthma is a developmental origin disease influenced by maternal diet and bacterial metabolites. Nat. Commun. 6, 7320 (2015).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
Abrahamsson, T. R. et al. Low gut microbiota diversity in early infancy precedes asthma at school age. Clin. Exp. Allergy 44, 842–850 (2014).
Vael, C., Nelen, V., Verhulst, S. L., Goossens, H. & Desager, K. N. Early intestinal Bacteroides fragilis colonisation and development of asthma. BMC Pulm. Med. 8, 19 (2008).
Orivuori, L. et al. High level of faecal calprotectin at age 2 months as a marker of intestinal inflammation predicts atopic dermatitis and asthma by age 6. Clin. Exp. Allergy 45, 928–939 (2015).
Russell, S. L. et al. Perinatal antibiotic treatment affects murine microbiota, immune responses and allergic asthma. Gut Microbes 4, 158–164 (2013).
Hevia, A. et al. Allergic patients with long-term asthma display low levels of Bifidobacterium adolescentis. PLoS ONE 11, e0147809 (2016).
Hua, X., Goedert, J. J., Pu, A., Yu, G. & Shi, J. Allergy associations with the adult fecal microbiota: analysis of the American Gut Project. EBioMedicine 3, 172–179 (2016).
He, F. et al. Stimulation of the secretion of pro-inflammatory cytokines by Bifidobacterium strains. Microbiol. Immunol. 46, 781–785 (2002).
Johnson, J. L., Jones, M. B. & Cobb, B. A. Bacterial capsular polysaccharide prevents the onset of asthma through T cell activation. Glycobiology 25, 368–375 (2015).
Sehrawat, A., Sinha, S. & Saxena, A. Helicobacter pylori neutrophil-activating protein: a potential Treg modulator suppressing allergic asthma. Front. Microbiol. 6, 493 (2015).
Koch, K. N. et al. Helicobacter urease-induced activation of the TLR2/NLRP3/IL-18 axis protects against asthma. J. Clin. Invest. 125, 3297–3302 (2015).
Engler, D. B. et al. Effective treatment of allergic airway inflammation with Helicobacter pylori immunomodulators requires BATF3-dependent dendritic cells and IL-10. Proc. Natl Acad. Sci. USA 111, 11810–11815 (2014).
Kim, Y. G. et al. Gut dysbiosis promotes M2 macrophage polarization and allergic airway inflammation via fungi-induced PGE(2). Cell Host Microbe 15, 95–102 (2014).
Zaiss, M. M. et al. The intestinal microbiota contributes to the ability of helminths to modulate allergic inflammation. Immunity 45, 998–1010 (2015).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Huang, F. et al. Early-life exposure to Clostridium leptum causes pulmonary immunosuppression. PLoS ONE 10, e0141717 (2015).
Li, Y. N. et al. Effect of oral feeding with Clostridium leptum on regulatory T-cell responses and allergic airway inflammation in mice. Ann. Allergy Asthma Immunol. 109, 201–207 (2012).
Segal, L. N. et al. Enrichment of lung microbiome with supraglottic taxa is associated with increased pulmonary inflammation. Microbiome 1, 19 (2013).
Pragman, A. A., Kim, H. B., Reilly, C. S., Wendt, C. & Isaacson, R. E. The lung microbiome in moderate and severe chronic obstructive pulmonary disease. PLoS ONE 7, e47305 (2012).
Sze, M. A. et al. The lung tissue microbiome in chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 185, 1073–1080 (2012).
Benjamin, J. L. et al. Smokers with active Crohn's disease have a clinically relevant dysbiosis of the gastrointestinal microbiota. Inflamm. Bowel Dis. 18, 1092–1100 (2012).
Biedermann, L. et al. Smoking cessation alters intestinal microbiota: insights from quantitative investigations on human fecal samples. Inflamm. Bowel Dis. 20, 1496–1501 (2014).
Kabeerdoss, J., Jayakanthan, P., Pugazhendhi, S. & Ramakrishna, B. S. Alterations of mucosal microbiota in the colon of patients with inflammatory bowel disease revealed by real time polymerase chain reaction amplification of 16S ribosomal ribonucleic acid. Indian J. Med. Res. 142, 23–32 (2015).
Schwab, C. et al. Longitudinal study of murine microbiota activity and interactions with the host during acute inflammation and recovery. ISME J. 8, 1101–1114 (2014).
Khonsari, S. et al. A comparative study of bifidobacteria in human babies and adults. Biosci. Microbiota Food Health 35, 97–103 (2016).
Verschuere, S. et al. Cigarette smoking alters epithelial apoptosis and immune composition in murine GALT. Lab. Invest. 91, 1056–1067 (2011).
Allais, L. et al. Chronic cigarette smoke exposure induces microbial and inflammatory shifts and mucin changes in the murine gut. Environ. Microbiol. 18, 1352–1363 (2015).
Hammadi, M., Adi, M., John, R., Khoder, G. A. & Karam, S. M. Dysregulation of gastric H,K-ATPase by cigarette smoke extract. World J. Gastroenterol. 15, 4016–4022 (2009).
Sapkota, A. R., Berger, S. & Vogel, T. M. Human pathogens abundant in the bacterial metagenome of cigarettes. Environ. Health Perspect. 118, 351–356 (2010).
Kulkarni, R. et al. Cigarette smoke increases Staphylococcus aureus biofilm formation via oxidative stress. Intect Immun. 80, 3804–3811 (2012).
Semlali, A., Killer, K., Alanazi, H., Chmielewski, W. & Rouabhia, M. Cigarette smoke condensate increases C. albicans adhesion, growth, biofilm formation, and EAP1, HWP1 and SAP2 gene expression. BMC Microbiol. 14, 61 (2014).
Hu, J., Wei, T., Sun, S., Zhao, A. & Xu, C. Effects of cigarette smoke condensate on the production and characterization of exopolysaccharides by Bifidobacterium. An. Acad. Bras. Cienc. 87, 997–1005 (2015).
Reale, M. et al. Daily intake of Lactobacillus casei Shirota increases natural killer cell activity in smokers. Br. J. Nutr. 108, 308–314 (2012).
Verheijden, K. A. T. et al. Treatment with specific prebiotics or probiotics prevents the development of lung emphysema in a mouse model of COPD. Eur. J. Pharmacol. 668, e12–e13 (2011).
Mortaz, E. et al. Anti-inflammatory effects of Lactobacillus rahmnosus and Bifidobacterium breve on cigarette smoke activated human macrophages. PLoS ONE 10, e0136455 (2015).
Tomoda, K. et al. Whey peptide-based enteral diet attenuated elastase-induced emphysema with increase in short chain fatty acids in mice. BMC Pulm. Med. 15, 64 (2015).
Tomoda, K. et al. Cigarette smoke decreases organic acids levels and population of Bifidobacterium in caecum of rats. J. Toxicol. Sci. 36, 261–266 (2011).
Kish, L. et al. Environmental particulate matter induces murine intestinal inflammatory responses and alters the gut microbiome. PLoS ONE 8, e62220 (2013).
Zuo, L. et al. Cigarette smoking is associated with intestinal barrier dysfunction in the small intestine but not in the large intestine of mice. J. Crohns Colitis 8, 1710–1722 (2014).
Huvenne, W. et al. Exacerbation of cigarette smoke-induced pulmonary inflammation by Staphylococcus aureus enterotoxin in mice. Respir. Res. 12, 69 (2011).
Brass, D. M. et al. Chronic LPS inhalation causes emphysema-like changes in mouse lung that are associated with apoptosis. Am. J. Respir. Cell. Mol. Biol. 39, 584–590 (2008).
Kelly, C. J. et al. Crosstalk between microbiota-derived short-chain fatty acids and intestinal epithelial HIF augments tissue barrier function. Cell Host Microbe 17, 662–671 (2015).
Suzuki, T., Yoshida, S. & Hara, H. Physiological concentrations of short-chain fatty acids immediately suppress colonic epithelial permeability. Br. J. Nutr. 100, 297–305 (2008).
Chen, L. W., Chen, P. H. & Hsu, C. M. Commensal microflora contribute to host defense against Escherichia coli pneumonia through toll-like receptors. Shock 36, 67–75 (2011).
Ichinohe, T. et al. Microbiota regulates immune defence against respiratory tract influenza A virus infection. Proc. Natl Acad. Sci. USA 108, 5354–5359 (2011).
Wu, S. et al. Microbiota regulates the TLR7 signaling pathway against respiratory tract influenza A virus infection. Curr. Microbiol. 67, 414–422 (2013).
Vieira, A. T. et al. Control of Klebsiella pneumoniae pulmonary infection and immunomodulaation by oral treatment with commensal probiotic Bifidobacterium longum 51A. Microbes Infect. 18, 180–189 (2016).
Kawahara, T. et al. Consecutive oral administration of Bifidobacterium longum MM-2 improves the defense system against influenza virus infection by enhancing natural killer cell activity in a murine model. Microbiol. Immunol. 59, 1–12 (2015).
Luoto, R. et al. Prebiotic and probiotic supplementation prevents rhinovirus infections in preterm infants: a randomized placebo-controlled trial. J. Allergy Clin. Immunol. 133, 405–413 (2014).
Jespersen, L. et al. Effect of Lactobacillus paracasei subsp. paracasei, L. casei 431 on immune response to influenza vaccination and upper respiratory tract infections in healthy adult volunteers: a randomized, double-blind, placebo-controlled, parallel-group study. Am. J. Clin. Nutr. 101, 1188–1196 (2015).
King, S., Glanville, J., Sanders, M. E., Fitzgerald, A. & Varley, D. Effectiveness of probiotics on the duration of illness in healthy children and adults who develop common acute respiratory infectious conditions: a systematic review and meta-analysis. Br. J. Nutr. 112, 41–54 (2014).
West, N. P. et al. Probiotic supplementation for respiratory and gastrointestinal illness symptoms in healthy physically active individuals. Clin. Nutr. 33, 581–587 (2014).
Clarke, T. B. Early innate immunity to bacterial infection in the lung is regulated systemically by the commensal microbiota via Nod-like receptor ligands. Infect. Immun. 82, 4596–4606 (2014).
Oh, K. Z. et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity 41, 478–492 (2014).
Bernard, H. et al. Dietary pectin-derived acidic oligosaccharides improve the pulmonary bacterial clearance of Pseudomonas aeruginosa lung infection in mice by modulating intestinal microbiota and immunity. J. Infect. Dis. 211, 156–165 (2015).
Kishino, E., Takemura, N., Masaki, H., Ito, T. & Nakazawa, M. Dietary lactosucrose suppresses influenza A (H1N1) virus infection in mice. Biosci. Microbiota Food Health 34, 67–76 (2015).
Wang, J. et al. Respiratory influenza virus infection induces intestinal immune injury via microbiota-mediated TH17 cell–dependent inflammation. J. Exp. Med. 211, 2397–2410 (2014).
Huang, Y. J. & LiPuma, J. J. The microbiome in cystic fibrosis. Clin. Chest Med. 37, 59–67 (2016).
Strachan, D. P. Hay fever, hygiene, and household size. BMJ 299, 1259–1260 (1989).
Riedler, J. et al. Exposure to farming in early life and development of asthma and allergy: a cross-sectional survey. Lancet 358, 1129–1133 (2001).
Ball, T. M. et al. Siblings, day-care attendance, and the risk of asthma and wheezing during childhood. N. Engl. J. Med. 343, 538–543 (2000).
Rook, G. A., Martinelli, R. & Brunet, L. R. Innate immune responses to mycobacteria and the downregulation of atopic responses. Curr. Opin. Allergy Clin. Immunol. 3, 337–342 (2003).
Bieber, T. Atopic dermatitis. N. Engl. J. Med. 358, 1483–1494 (2008).
Gale, E. A. The rise of childhood type 1 diabetes in the 20th century. Diabetes 51, 3353–3361 (2002).
Alonso, A. & Hernan, M. A. Temporal trends in the incidence of multiple sclerosis: a systematic review. Neurology 71, 129–135 (2008).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Ottman, N., Smidt, H., de Vos, W. M. & Belzer, C. The function of our microbiota: who is out there and what do they do? Front. Cell. Infect. Microbiol. 2, 104 (2012).
Okada, H., Kuhn, C., Feillet, H. & Bach, J. F. The 'hygiene hypothesis' for autoimmune and allergic diseases: an update. Clin. Exp. Immunol. 160, 1–9 (2010).
Maizels, R. M., McSorley, H. J. & Smyth, D. J. Helminths in the hygiene hypothesis: sooner or later? Clin. Exp. Immunol. 177, 38–46 (2014).
Sun, X., Fiala, J. L. & Lowery, D. Patent watch: modulating the human microbiome with live biotherapeutic products: intellectual property landscape. Nat. Rev. Drug Discov. 15, 224–225 (2016).
Brown, A. J. et al. Pharmacological properties of acid N-thiazolylamide FFA2 agonists. Pharmacol. Res. Perspect. 3, e00141 (2015).
Hudson, B. D. et al. Defining the molecular basis for the first potent and selective orthosteric agonists of the FFA2 free fatty acid receptor. J. Biol. Chem. 288, 17296–17312 (2013).
Schmidt, J. et al. Selective orthosteric free fatty acid receptor 2 (FFA2) agonists: identification of the structural and chemical requirements for selective activation of FFA2 versus FFA3. J. Biol. Chem. 286, 10628–10640 (2011).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Acknowledgements
The authors acknowledge the support of fellowships from the Australian National Health and Medical Research Council (NHMRC; to M.A.C. and P.M.H.), the Australian Research Council (ARC; to P.H.), the Brawn Foundation, the Faculty of Health and Medicine at the University of Newcastle, Australia, and grants from the NHMRC and the Rainbow Foundation (to P.M.H.). The authors thank F. Thomson and M. Thomson for their continued support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Budden, K., Gellatly, S., Wood, D. et al. Emerging pathogenic links between microbiota and the gut–lung axis. Nat Rev Microbiol 15, 55–63 (2017). https://doi.org/10.1038/nrmicro.2016.142
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2016.142
This article is cited by
-
Intact lung tissue and bronchoalveolar lavage fluid are both suitable for the evaluation of murine lung microbiome in acute lung injury
Microbiome (2024)
-
Vitamin D and the microbiota connection: understanding its potential to improve COPD outcomes
The Egyptian Journal of Bronchology (2024)
-
Impact of hyperoxia on the gut during critical illnesses
Critical Care (2024)
-
Pathophysiology of acute lung injury in patients with acute brain injury: the triple-hit hypothesis
Critical Care (2024)
-
Lung microbiome: new insights into the pathogenesis of respiratory diseases
Signal Transduction and Targeted Therapy (2024)