Abstract
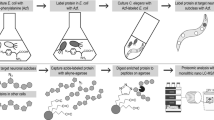
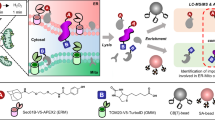
In this protocol we describe the incorporation of bio-orthogonal amino acids as a versatile method for visualizing and identifying de novo–synthesized proteins in the roundworm Caenorhabditis elegans. This protocol contains directions on implementing three complementary types of analysis: 'click chemistry' followed by western blotting, click chemistry followed by immunofluorescence, and isobaric tags for relative and absolute quantification (iTRAQ) quantitative mass spectrometry. The detailed instructions provided herein enable researchers to investigate the de novo proteome, an analysis that is complicated by the fact that protein molecules are chemically identical to each other, regardless of the timing of their synthesis. Our protocol circumvents this limitation by identifying de novo–synthesized proteins via the incorporation of the chemically modifiable azidohomoalanine instead of the natural amino acid methionine in the nascent protein, followed by facilitating the visualization of the resulting labeled proteins in situ. It will therefore be an ideal tool for studying de novo protein synthesis in physiological and pathological processes including learning and memory. The protocol requires 10 d for worm growth, liquid culture and synchronization; 1–2 d for bio-orthogonal labeling; and, with regard to analysis, 3–4 d for western blotting, 5–6 d for immunofluorescence or ∼3 weeks for mass spectrometry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Change history
12 September 2014
In the version of this article initially published, one of the two affiliations of one of the authors (Michael Kassiou) was incorrect. The mistaken affiliation read: “6Faculty of Health Sciences, Macquarie University, Sydney, New South Wales, Australia.” The correct affiliation is: “6Faculty of Health Sciences, University of Sydney, Sydney, New South Wales, Australia.” The error has been corrected in the HTML and PDF versions of the article.
References
Taylor, R.C. & Dillin, A. Aging as an event of proteostasis collapse. Cold Spring Harb. Perspect. Biol. 3 (2011).
Kaeberlein, M. & Kennedy, B.K. Hot topics in aging research: protein translation and TOR signaling, 2010. Aging Cell 10, 185–190 (2011).
Giboureau, N., Som, I.M., Boucher-Arnold, A., Guilloteau, D. & Kassiou, M. PET radioligands for the vesicular acetylcholine transporter (VAChT). Curr. Top Med. Chem. 10, 1569–1583 (2010).
De Strooper, B. Proteases and proteolysis in Alzheimer disease: a multifactorial view on the disease process. Physiol. Rev. 90, 465–494 (2010).
Chen, C.C. et al. Visualizing long-term memory formation in two neurons of the Drosophila brain. Science 335, 678–685 (2012).
Götz, J., Chen, F., van Dorpe, J. & Nitsch, R.M. Formation of neurofibrillary tangles in P301L tau transgenic mice induced by Aβ42 fibrils. Science 293, 1491–1495 (2001).
Ittner, L.M. et al. Dendritic function of tau mediates amyloid-β toxicity in Alzheimer's disease mouse models. Cell 142, 387–397 (2010).
Cajigas, I.J., Will, T. & Schuman, E.M. Protein homeostasis and synaptic plasticity. EMBO J. 29, 2746–2752 (2010).
Antonov, I., Kandel, E.R. & Hawkins, R.D. Presynaptic and postsynaptic mechanisms of synaptic plasticity and metaplasticity during intermediate-term memory formation in Aplysia. J. Neurosci. 30, 5781–5791 (2010).
Holt, C.E. & Schuman, E.M. The central dogma decentralized: new perspectives on RNA function and local translation in neurons. Neuron 80, 648–657 (2013).
David, D.C. et al. Proteomic and functional analysis reveal a mitochondrial dysfunction in P301L tau transgenic mice. J. Biol. Chem. 280, 23802–23814 (2005).
David, D.C. et al. β-Amyloid treatment of two complementary P301L tau-expressing Alzheimer's disease models reveals similar deregulated cellular processes. Proteomics 6, 6566–6577 (2006).
Lim, Y.-A. et al. Aβ and human amylin share a common toxicity pathway via mitochondrial dysfunction. Proteomics 10, 1621–1633 (2010).
Rhein, V. et al. Amyloid-β and tau synergistically impair the oxidative phosphorylation system in triple transgenic Alzheimer's disease mice. Proc. Natl. Acad. Sci. USA 106, 20057–20062 (2009).
Chen, F. et al. Role for glyoxalase I in Alzheimer's disease. Proc. Natl. Acad. Sci. USA 101, 7687–7692 (2004).
Hoerndli, F.J., Pelech, S., Papassotiropoulos, A. & Götz, J. Aβ treatment and P301L tau expression in an Alzheimer's disease tissue culture model act synergistically to promote aberrant cell cycle re-entry. Eur. J. Neurosci. 26, 60–72 (2007).
Schonrock, N. et al. Neuronal microRNA deregulation in response to Alzheimer's disease amyloid-β. PLoS ONE 5, e11070 (2010).
Götz, J. & Ittner, L.M. Animal models of Alzheimer's disease and frontotemporal dementia. Nat. Rev. Neurosci. 9, 532–544 (2008).
Chew, Y.L., Fan, X., Götz, J. & Nicholas, H.R. Protein with tau-like repeats regulates neuronal integrity and lifespan in C. elegans. J. Cell Sci. 126, 2079–2091 (2013).
Pienaar, I.S., Götz, J. & Feany, M.B. Parkinson's disease: insights from non-traditional model organisms. Prog. Neurobiol. 92, 558–571 (2010).
Duboff, B., Götz, J. & Feany, M.B. Tau promotes neurodegeneration via DRP1 mislocalization. Neuron 75, 618–632 (2012).
Liang, V. et al. Altered proteostasis in aging and heat shock response in C. elegans revealed by analysis of the global and de novo synthesized proteome. Cell Mol. Life Sci. 10.1007/s00018-014-1558-7 (2014).
Kaletta, T. & Hengartner, M.O. Finding function in novel targets: C. elegans as a model organism. Nat. Rev. Drug Discov. 5, 387–398 (2006).
Gautier, A., Nakata, E., Lukinavicius, G., Tan, K.T. & Johnsson, K. Selective cross-linking of interacting proteins using self-labeling tags. J. Am. Chem. Soc. 131, 17954–17962 (2009).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Machleidt, T., Robers, M. & Hanson, G.T. Protein labeling with FlAsH and ReAsH. Methods Mol. Biol. 356, 209–220 (2007).
Hughes, A.J. & Herr, A.E. Microfluidic western blotting. Proc. Natl. Acad. Sci. USA 109, 21450–21455 (2012).
David, D., Hoerndli, F. & Götz, J. Functional genomics meets neurodegenerative disorders. Part I: transcriptomic and proteomic technology. Prog. Neurobiol. 76, 153–168 (2005).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Husi, H. et al. Selective chemical intervention in the proteome of Caenorhabditis elegans. J. Proteome Res. 9, 6060–6070 (2010).
Fredens, J. et al. Quantitative proteomics by amino acid labeling in C. elegans. Nat. Methods 8, 845–847 (2011).
Larance, M. et al. Stable-isotope labeling with amino acids in nematodes. Nat. Methods 8, 849–851 (2011).
Boisvert, F.M. et al. A quantitative spatial proteomics analysis of proteome turnover in human cells. Mol. Cell. Proteomics 11, M111.011429 (2012).
Seyfried, N.T. et al. Multiplex SILAC analysis of a cellular TDP-43 proteinopathy model reveals protein inclusions associated with SUMOylation and diverse polyubiquitin chains. Mol. Cell. Proteomics 9, 705–718 (2010).
Huh, K.H. & Wenthold, R.J. Turnover analysis of glutamate receptors identifies a rapidly degraded pool of the N-methyl-D-aspartate receptor subunit, NR1, in cultured cerebellar granule cells. J. Biol. Chem. 274, 151–157 (1999).
Shi, H.J., Stubbs, R. & Hood, K. Characterization of de novo synthesized proteins released from human colorectal tumour explants. Electrophoresis 30, 2442–2453 (2009).
Ong, S.E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660 (2006).
Prescher, J.A. & Bertozzi, C.R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Beatty, K.E. & Tirrell, D.A. Two-color labeling of temporally defined protein populations in mammalian cells. Bioorg. Med. Chem. Lett. 18, 5995–5999 (2008).
Kolb, H.C., Finn, M.G. & Sharpless, K.B. Click chemistry: diverse chemical function from a few good reactions. Angew Chem. Int. Ed. Engl. 40, 2004–2021 (2001).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Kiick, K.L., Saxon, E., Tirrell, D.A. & Bertozzi, C.R. Incorporation of azides into recombinant proteins for chemoselective modification by the Staudinger ligation. Proc. Natl. Acad. Sci. USA 99, 19–24 (2002).
Havrylenko, S., Legouis, R., Negrutskii, B. & Mirande, M. Methionyl-tRNA synthetase from Caenorhabditis elegans: a specific multidomain organization for convergent functional evolution. Protein Sci. 19, 2475–2484 (2010).
Laughlin, S.T., Baskin, J.M., Amacher, S.L. & Bertozzi, C.R. In vivo imaging of membrane-associated glycans in developing zebrafish. Science 320, 664–667 (2008).
Kho, Y. et al. A tagging-via-substrate technology for detection and proteomics of farnesylated proteins. Proc. Natl. Acad. Sci. USA 101, 12479–12484 (2004).
Bruckman, M.A. et al. Surface modification of tobacco mosaic virus with 'click' chemistry. Chembiochem 9, 519–523 (2008).
Weisbrod, S.H. & Marx, A. Novel strategies for the site-specific covalent labelling of nucleic acids. Chem. Commun. (Camb.) 2008, 5675–5685 (2008).
Hinz, F.I., Dieterich, D.C., Tirrell, D.A. & Schuman, E.M. Non-canonical amino acid labeling in vivo to visualize and affinity purify newly synthesized proteins in larval zebrafish. ACS Chem. Neurosci. 3, 40–49 (2012).
Laughlin, S.T. & Bertozzi, C.R. Imaging the glycome. Proc. Natl. Acad. Sci. USA 106, 12–17 (2009).
Laughlin, S.T. & Bertozzi, C.R. In vivo imaging of Caenorhabditis elegans glycans. ACS Chem. Biol. 4, 1068–1072 (2009).
Dieterich, D.C. et al. Labeling, detection and identification of newly synthesized proteomes with bioorthogonal non-canonical amino-acid tagging. Nat. Protoc. 2, 532–540 (2007).
Dieterich, D.C. et al. In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat. Neurosci. 13, 897–905 (2010).
Putz, S.M., Boehm, A.M., Stiewe, T. & Sickmann, A. iTRAQ analysis of a cell culture model for malignant transformation, including comparison with 2D-PAGE and SILAC. J. Proteome Res. 11, 2140–2153 (2012).
Yoon, B.C. et al. Local translation of extranuclear lamin B promotes axon maintenance. Cell 148, 752–764 (2012).
Nicholas, H.R. & Hodgkin, J. The ERK MAP kinase cascade mediates tail swelling and a protective response to rectal infection in C. elegans. Curr. Biol. 14, 1256–1261 (2004).
Nicholas, H.R. & Hodgkin, J. Responses to infection and possible recognition strategies in the innate immune system of Caenorhabditis elegans. Mol. Immunol. 41, 479–493 (2004).
Kramer, G., Kasper, P.T., de Jong, L. & de Koster, C.G. Quantitation of newly synthesized proteins by pulse labeling with azidohomoalanine. Methods Mol. Biol. 753, 169–181 (2011).
Nicholas, H.R., Lowry, J.A., Wu, T. & Crossley, M. The Caenorhabditis elegans protein CTBP-1 defines a new group of THAP domain-containing CtBP co-repressors. J. Mol. Biol. 375, 1–11 (2008).
de Bono, M. & Maricq, A.V. Neuronal substrates of complex behaviors in C. elegans. Annu. Rev. Neurosci. 28, 451–501 (2005).
David, D.C. et al. Widespread protein aggregation as an inherent part of aging in C. elegans. PLoS Biol. 8, e1000450 (2010).
Ngo, J.T. et al. Cell-selective metabolic labeling of proteins. Nat. Chem. Biol. 5, 715–717 (2009).
Altun, Z.F. & Hall, D.H. Handbook of C. elegans anatomy. WormAtlas http://www.wormatlas.org/hermaphrodite/hermaphroditehomepage.htm (2012).
Ghafouri, S. & McGhee, J.D. Bacterial residence time in the intestine of Caenorhabditis elegans. Nematology 9, 87–91 (2007).
Haenni, S. et al. Analysis of C. elegans intestinal gene expression and polyadenylation by fluorescence-activated nuclei sorting and 3′-end-seq. Nucleic Acids Res. 40, 6304–6318 (2012).
Finney, M. & Ruvkun, G. The unc-86 gene product couples cell lineage and cell identity in C. elegans. Cell 63, 895–905 (1990).
Duerr, J.S. Immunohistochemistry. Misc: In WormBook C. elegans Research Community 1–61 10.1895/wormbook.1.105.1 (19 June 2006).
Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Link, A.J. & Tirrell, D.A. Cell surface labeling of Escherichia coli via copper(I)-catalyzed [3+2] cycloaddition. J. Am. Chem. Soc. 125, 11164–11165 (2003).
Gravato-Nobre, M.J. et al. Multiple genes affect sensitivity of Caenorhabditis elegans to the bacterial pathogen Microbacterium nematophilum. Genetics 171, 1033–1045 (2005).
You, Y.J., Kim, J., Cobb, M. & Avery, L. Starvation activates MAP kinase through the muscarinic acetylcholine pathway in Caenorhabditis elegans pharynx. Cell Metab. 3, 237–245 (2006).
Keith, S.A., Amrit, F.R., Ratnappan, R. & Ghazi, A. The C. elegans healthspan and stress-resistance assay toolkit. Methods 68, 476–486 (2014).
Minniti, A.N. et al. Intracellular amyloid formation in muscle cells of Aβ-transgenic Caenorhabditis elegans: determinants and physiological role in copper detoxification. Mol. Neurodegener. 4, 2 (2009).
Zhang, T., Mullane, P.C., Periz, G. & Wang, J. TDP-43 neurotoxicity and protein aggregation modulated by heat shock factor and insulin/IGF-1 signaling. Hum. Mol. Genet. 20, 1952–1965 (2011).
Yoneda, T. et al. Compartment-specific perturbation of protein handling activates genes encoding mitochondrial chaperones. J. Cell. Sci. 117, 4055–4066 (2004).
Calfon, M. et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 415, 92–96 (2002).
Acknowledgements
This study was supported by the Estate of Dr. Clem Jones, AO, and by grants from the Australian Research Council (DP13300101932) and the National Health and Medical Research Council of Australia (APP1037746 and APP1003150) to J.G. Mass spectrometry was undertaken at The Australian Proteome Facility, with the infrastructure provided by the Australian Government through the National Collaborative Research Infrastructure Strategy. Some strains were provided by the CGC, which is funded by the US National Institutes of Health Office of Research Infrastructure Programs (P40 OD010440).
Author information
Authors and Affiliations
Contributions
M.U., V.L., Y.L.C., S.B., X.S., T.Z., H.L. and S.B. performed the experiments; M.U., V.L., Y.L.C., S.B., X.S., T.Z., H.L., S.B., M.K., H.R.N. and J.G. analyzed the data; and M.U., V.L., Y.L.C., H.R.N. and J.G. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ullrich, M., Liang, V., Chew, Y. et al. Bio-orthogonal labeling as a tool to visualize and identify newly synthesized proteins in Caenorhabditis elegans. Nat Protoc 9, 2237–2255 (2014). https://doi.org/10.1038/nprot.2014.150
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.150
This article is cited by
-
Extracellular matrix dynamics: tracking in biological systems and their implications
Journal of Biological Engineering (2022)
-
Neuronal subclass-selective proteomic analysis in Caenorhabditis elegans
Scientific Reports (2020)
-
Non-canonical amino acid labeling in proteomics and biotechnology
Journal of Biological Engineering (2019)
-
Nonradioactive quantification of autophagic protein degradation with L-azidohomoalanine labeling
Nature Protocols (2017)
-
Plasma biomarker proteins for detection of human growth hormone administration in athletes
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.