Abstract
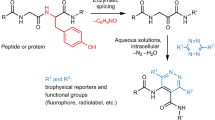
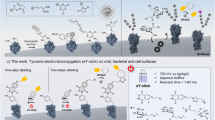
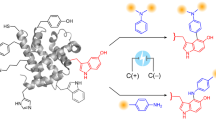
This protocol describes an efficient method to site-specifically label cell-surface or purified proteins with chemical probes in two steps: probe incorporation mediated by enzymes (PRIME) followed by chelation-assisted copper-catalyzed azide-alkyne cycloaddition (CuAAC). In the PRIME step, Escherichia coli lipoic acid ligase (LplA) site-specifically attaches a picolyl azide (pAz) derivative to a 13-aa recognition sequence that has been genetically fused onto the protein of interest. Proteins bearing pAz are chemoselectively derivatized with an alkyne-probe conjugate by chelation-assisted CuAAC in the second step. We describe herein the optimized protocols to synthesize pAz to perform PRIME labeling and to achieve CuAAC derivatization of pAz on live cells, fixed cells and purified proteins. Reagent preparations, including synthesis of pAz probes and expression of LplA, take 12 d, whereas the procedure for performing site-specific pAz ligation and CuAAC on cells or on purified proteins takes 40 min–3 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
04 December 2013
In the version of this article initially published, Katharine A. White, Scott Grecian, Scott Clarke and Kyle R. Gee were not present in the author list. The error has been corrected in the HTML and PDF versions of the article. The author contributions section and competing financial interests statement have also been modified to include information relevant to these authors.
References
Uttamapinant, C. et al. Fast, cell-compatible click chemistry with copper-chelating azides for biomolecular labeling. Angew. Chem. Int. Ed. Engl. 51, 5852–5856 (2012).
Uttamapinant, C. et al. A fluorophore ligase for site-specific protein labeling inside living cells. Proc. Natl. Acad. Sci. USA 107, 10914–10919 (2010).
Liu, D.S. et al. Diels-Alder cycloaddition for fluorophore targeting to specific proteins inside living cells. J. Am. Chem. Soc. 134, 792–795 (2012).
Cohen, J.D., Thompson, S. & Ting, A.Y. Structure-guided engineering of a Pacific Blue fluorophore ligase for specific protein imaging in living cells. Biochemistry 50, 8221–8225 (2011).
Jin, X., Uttamapinant, C. & Ting, A.Y. Synthesis of 7-aminocoumarin by Buchwald-Hartwig cross coupling for specific protein labeling in living cells. Chembiochem 12, 65–70 (2011).
Baruah, H. et al. An engineered aryl azide ligase for site-specific mapping of protein-protein interactions through photo-cross-linking. Angew. Chem. Int. Ed. Engl. 47, 7018–7021 (2008).
Yao, J.Z. et al. Fluorophore targeting to cellular proteins via enzyme-mediated azide ligation and strain-promoted cycloaddition. J. Am. Chem. Soc. 134, 3720–3728 (2012).
Liu, D.S. et al. Quantum dot targeting with lipoic acid ligase and HaloTag for single-molecule imaging on living cells. ACS Nano. 6, 11080–11087 (2012).
Liu, D.S. et al. Imaging trans-cellular neurexin-neuroligin interactions by enzymatic probe ligation. PLoS ONE 8, e52823 (2013).
Hong, V. et al. Analysis and optimization of copper-catalyzed azide-alkyne cycloaddition for bioconjugation. Angew. Chem. Int. Ed. Engl. 48, 9879–9883 (2009).
Sako, Y., Minoghchi, S. & Yanagida, T. Single-molecule imaging of EGFR signalling on the surface of living cells. Nat. Cell Biol. 2, 168–172 (2000).
Dani, A. et al. Superresolution imaging of chemical synapses in the brain. Neuron 68, 843–856 (2010).
Kellner, R.R. et al. Nanoscale organization of nicotinic acetylcholine receptors revealed by stimulated emission depletion microscopy. Neuroscience 144, 135–143 (2007).
Zou, P. & Ting, A.Y. Imaging LDL receptor oligomerization during endocytosis using a co-internalization assay. ACS Chem. Biol. 6, 308–313 (2011).
Marsh, D.R. et al. Distribution of an NMDA receptor:GFP fusion protein in sensory neurons is altered by a C-terminal construct. J. Neurochem. 77, 23–33 (2001).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Howarth, M. et al. A monovalent streptavidin with a single femtomolar biotin binding site. Nat. Methods 3, 267–273 (2006).
Klemm, J.D., Schreiber, S.L. & Crabtree, G.R. Dimerization as a regulatory mechanism in signal transduction. Annu. Rev. Immunol. 16, 569–592 (1998).
Shi, X. et al. Quantitative fluorescence labeling of aldehyde-tagged proteins for single-molecule imaging. Nat. Methods 9, 499–503 (2012).
Dirksen, A. et al. Nucleophilic catalysis of hydrazone formation and transimination: implications for dynamic covalent chemistry. J. Am. Chem. Soc. 128, 15602–15603 (2006).
Martin, B.R. et al. Mammalian cell-based optimization of the biarsenical-binding tetracysteine motif for improved fluorescence and affinity. Nat. Biotechnol. 23, 1308–1314 (2005).
Beatty, K.E. et al. Live-cell imaging of cellular proteins by a strain-promoted azide-alkyne cycloaddition. Chembiochem 11, 2092–2095 (2010).
Puthenveetil, S. et al. Yeast display evolution of a kinetically efficient 13-amino acid substrate for lipoic acid ligase. J. Am. Chem. Soc. 131, 16430–16438 (2009).
Howarth, M. et al. Monovalent, reduced-size quantum dots for imaging receptors on living cells. Nat. Methods 5, 397–399 (2008).
Carrington, J.C. & Dougherty, W.G. A viral cleavage site cassette: identification of amino acid sequences required for tobacco etch virus polyprotein processing. Proc. Natl. Acad. Sci. USA 85, 3391–3395 (1988).
Besanceney-Webler, C. et al. Increasing the efficacy of bioorthogonal click reactions for bioconjugation: a comparative study. Angew. Chem. Int. Ed. Engl. 50, 8051–8056 (2011).
Chan, T.R. et al. Polytriazoles as copper(I)-stabilizing ligands in catalysis. Org. Lett. 6, 2853–2855 (2004).
Howarth, M. & Ting, A.Y. Imaging proteins in live mammalian cells with biotin ligase and monovalent streptavidin. Nat. Protoc. 3, 534–545 (2008).
Woehlke, G. et al. Microtubule interaction site of the kinesin motor. Cell 90, 207–216 (1997).
Presolski, S.I., Hong, V.P. & Finn, M.G. Copper-catalyzed azide-alkyne click chemistry for bioconjugation. Curr. Protoc. Chem. Biol. 3, 153–162 (2011).
Acknowledgements
We thank Ting laboratory members who have contributed to PRIME-related protocols, particularly M. Fernández-Suárez. We also thank A. Karunakaran (University of California, San Francisco (UCSF)) and R.D. Vale (UCSF) for LAP-kinesin K560 and kinesin K420 proteins, C. Garner (Stanford University) for the neuron fixative recipe and M.G. Finn (Scripps Research Institute) for helpful advice on CuAAC. Funding was provided by the US National Institutes of Health (R01 GM072670) and the Dreyfus Foundation.
Author information
Authors and Affiliations
Contributions
C.U., M.I.S., D.S.L., J.Z.Y., K.A.W., S.G., S.C., K.R.G. and A.Y.T. developed protocols. C.U. contributed all the data. C.U., M.I.S. and A.Y.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
A.Y.T. holds a patent (publication number US8137925 B2) related to methods that use lipoic acid ligase for specific protein labeling. Life Technologies has applied for a patent related to picolyl azide reagents.
Supplementary information
Supplementary Figure 1
Estimation of cell-surface labeling yield by streptavidin gel shift assay. HEK cells expressing LAP-tagged cyan fluorescent protein fused to the transmembrane domain of the platelet-derived growth factor receptor (LAP-CFP-TM1) were labeled with 200 μM picolyl azide for 20 min using exogenously supplied W37VLplA, then by live-cell CuAAC for 5 min with 20 μM biotin-alkyne and 50 μM CuSO4. Cells were then lysed under hypotonic conditions and incubated with ∼10 μM streptavidin for 30 min at room temperature. The reaction mixture was analyzed on 12% SDS-PAGE without boiling the samples, to preserve biotin-streptavidin binding. CFP-containing species were visualized by in-gel CFP fluorescence. The shift in the mobility of a population of LAP-CFP-TM upon streptavidin addition gives the yield of biotin-alkyne modification on LAP-CFP-TM. Negative controls with streptavidin omitted (lane 1) or with AP-CFP-TM2 replacing LAP-CFP-TM (lanes 3-4) showed no change in mobility. We previously estimated that ∼50% of TM fusion proteins (such as LAP-CFP-TM) are located at the cell surface, with the rest in the ER, Golgi, or endosomes. Therefore, if ∼38% of total LAP-CFP-TM is biotinylated by our standard labeling protocol (average of two independent replicates), we estimate that ∼76% of its cell surface pool is labeled. (PDF 571 kb)
Rights and permissions
About this article
Cite this article
Uttamapinant, C., Sanchez, M., Liu, D. et al. Site-specific protein labeling using PRIME and chelation-assisted click chemistry. Nat Protoc 8, 1620–1634 (2013). https://doi.org/10.1038/nprot.2013.096
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.096
This article is cited by
-
Further assessments of ligase LplA-mediated modifications of proteins in vitro and in cellulo
Molecular Biology Reports (2022)
-
A far-red hybrid voltage indicator enabled by bioorthogonal engineering of rhodopsin on live neurons
Nature Chemistry (2021)
-
A quantitative thiol reactivity profiling platform to analyze redox and electrophile reactive cysteine proteomes
Nature Protocols (2020)
-
Genetic code expansion in the mouse brain
Nature Chemical Biology (2016)
-
Labeling proteins on live mammalian cells using click chemistry
Nature Protocols (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.