Abstract
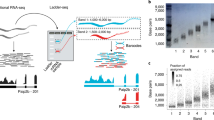
De novo assembly of RNA-seq data enables researchers to study transcriptomes without the need for a genome sequence; this approach can be usefully applied, for instance, in research on 'non-model organisms' of ecological and evolutionary importance, cancer samples or the microbiome. In this protocol we describe the use of the Trinity platform for de novo transcriptome assembly from RNA-seq data in non-model organisms. We also present Trinity-supported companion utilities for downstream applications, including RSEM for transcript abundance estimation, R/Bioconductor packages for identifying differentially expressed transcripts across samples and approaches to identify protein-coding genes. In the procedure, we provide a workflow for genome-independent transcriptome analysis leveraging the Trinity platform. The software, documentation and demonstrations are freely available from http://trinityrnaseq.sourceforge.net. The run time of this protocol is highly dependent on the size and complexity of data to be analyzed. The example data set analyzed in the procedure detailed herein can be processed in less than 5 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Wang, Z., Gerstein, M. & Snyder, M. RNA-seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Haas, B.J. & Zody, M.C. Advancing RNA-seq analysis. Nat. Biotechnol. 28, 421–423 (2010).
Martin, J.A. & Wang, Z. Next-generation transcriptome assembly. Nat. Rev. Genet. 12, 671–682 (2011).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Guttman, M. et al. Ab initio reconstruction of cell type–specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat. Biotechnol. 28, 503–510 (2010).
Robertson, G. et al. De novo assembly and analysis of RNA-seq data. Nat. Methods 7, 909–912 (2010).
Schulz, M.H., Zerbino, D.R., Vingron, M. & Birney, E. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28, 1086–1092 (2012).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Duan, J., Xia, C., Zhao, G., Jia, J. & Kong, X. Optimizing de novo common wheat transcriptome assembly using short-read RNA-seq data. BMC Genomics 13, 392 (2012).
Xu, D.L. et al. De novo assembly and characterization of the root transcriptome of Aegilops variabilis during an interaction with the cereal cyst nematode. BMC Genomics 13, 133 (2012).
Zhao, Q.Y. et al. Optimizing de novo transcriptome assembly from short-read RNA-seq data: a comparative study. BMC Bioinformatics 12 (suppl. 14), S2 (2011).
Henschel, R. et al. Trinity RNA-seq assembler performance optimization. XSEDE '12 Proceedings of the 1st Conference of the Extreme Science and Engineering Discovery Environment: bridging from the eXtreme to the campus and beyond (Chicago, Illinois, USA, July 16–20, 2012) http://dx.doi.org/10.1145/2335755.2335842 (2012).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Bullard, J.H., Purdom, E., Hansen, K.D. & Dudoit, S. Evaluation of statistical methods for normalization and differential expression in mRNA-seq experiments. BMC Bioinformatics 11, 94 (2010).
Fang, Z. & Cui, X. Design and validation issues in RNA-seq experiments. Briefi. Bioinform. 12, 280–287 (2011).
Auer, P.L. & Doerge, R.W. Statistical design and analysis of RNA sequencing data. Genetics 185, 405–416 (2010).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Roberts, A. & Pachter, L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nat. Methods 10, 71–73 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Robinson, M.D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Dillies, M.A. et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform. http://dx.doi.org/10.1093/bib/bbs046 (17 September 2012).
Marioni, J.C., Mason, C.E., Mane, S.M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Abeel, T., Van Parys, T., Saeys, Y., Galagan, J. & Van de Peer, Y. GenomeView: a next-generation genome browser. Nucleic Acids Res. 40, e12 (2012).
Liu, L. et al. Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012, 251364 (2012).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Rothberg, J.M. et al. An integrated semiconductor device enabling non-optical genome sequencing. Nature 475, 348–352 (2011).
Van Belleghem, S.M., Roelofs, D., Van Houdt, J. & Hendrickx, F. De novo transcriptome assembly and SNP discovery in the wing polymorphic salt marsh beetle Pogonus chalceus (Coleoptera, Carabidae). PLoS ONE 7, e42605 (2012).
Kleinman, C.L. & Majewski, J. Comment on “Widespread RNA and DNA sequence differences in the human transcriptome”. Science 335, 1302 (2012).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Pounds, S.B., Gao, C.L. & Zhang, H. Empirical Bayesian selection of hypothesis testing procedures for analysis of sequence count expression data. Stat. Appl. Genet. Mol. Biol. http://dx.doi.org/10.1515/1544-6115.1773 (2012).
Tarazona, S., Garcia-Alcalde, F., Dopazo, J., Ferrer, A. & Conesa, A. Differential expression in RNA-seq: a matter of depth. Genome Res. 21, 2213–2223 (2011).
Cumbie, J.S. et al. GENE-counter: a computational pipeline for the analysis of RNA-seq data for gene expression differences. PLoS ONE 6, e25279 (2011).
Hardcastle, T.J. & Kelly, K.A. baySeq: empirical Bayesian methods for identifying differential expression in sequence count data. BMC Bioinformatics 11, 422 (2010).
Leng, N. et al. An empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29, 1035–1043 (2012).
Tuna, M. & Amos, C.I. Genomic sequencing in cancer. Cancer Lett. http://dx.doi.org/doi:10.1016/j.canlet.2012.11.004 (2012).
Rhind, N. et al. Comparative functional genomics of the fission yeasts. Science 332, 930–936 (2011).
Kumar, S. & Blaxter, M.L. Comparing de novo assemblers for 454 transcriptome data. BMC Genomics 11, 571 (2010).
Papanicolaou, A., Stierli, R., Ffrench-Constant, R.H. & Heckel, D.G. Next generation transcriptomes for next generation genomes using est2assembly. BMC Bioinformatics 10, 447 (2009).
Lohse, M. et al. RobiNA: a user-friendly, integrated software solution for RNA-seq–based transcriptomics. Nucleic Acids Res. 40, W622–W627 (2012).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17 http://journal.embnet.org/index.php/embnetjournal/article/view/200/479 (2011).
Haas, B.J., Chin, M., Nusbaum, C., Birren, B.W. & Livny, J. How deep is deep enough for RNA-seq profiling of bacterial transcriptomes? BMC Genomics 13, 734 (2012).
Brown, C.T., Howe, A., Zhang, Q., Pryrkosz, A.B. & Brom, T.H. A reference-free algorithm for computational normalization of shotgun sequencing data. arXiv:1203.4802 [q-bio.GN] (2012).
Borodina, T., Adjaye, J. & Sultan, M. A strand-specific library preparation protocol for RNA sequencing. Methods Enzymol. 500, 79–98 (2011).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 37, e123 (2009).
Sung, W.K. et al. Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat. Genet. 44, 765–769 (2012).
Acknowledgements
We are grateful to D. Jaffe and S. Young for access to additional computing resources, to Z. Chen for help in R-scripting, to L. Gaffney for help with figure illustrations, to C. Titus Brown for essential discussions and inspiration related to digital normalization strategies, to G. Marcais and C. Kingsford for supporting the use of their Jellyfish software in Trinity and to B. Walenz for supporting our earlier use of Meryl. We are grateful to our users and their feedback, in particular J. Wortman and P. Bain for comments on earlier drafts of the manuscript. This project has been funded in part (B.J.H.) with Federal funds from the National Institute of Allergy and Infectious Diseases (NIAID), US National Institutes of Health (NIH), Department of Health and Human Services (DHHS), under contract no. HHSN272200900018C. Work was supported by Howard Hughes Medical Institute (HHMI), a NIH PIONEER award, a Center for Excellence in Genome Science grant no. 5P50HG006193-02 from the National Human Genome Research Institute (NHGRI) and the Klarman Cell Observatory at the Broad Institute (A.R.). A.P. was supported by the CSIRO Office of the Chief Executive (OCE). M.Y. was supported by the Clore Foundation. P.B. was supported by the National Science Foundation (NSF) grant no. OCI-1053575 for the Extreme Science and Engineering Discovery Environment (XSEDE) project. B.L. and C.D. were partially supported by NIH grant no.1R01HG005232-01A1. In addition, B.L. was partially funded by J. Thomson's MacArthur Professorship and by the Morgridge Institute for Research support for Computation and Informatics in Biology and Medicine. M.L. was supported by the Bundesministerium für Bildung und Forschung via the project 'NGSgoesHPC'. N.P. was funded by the Fund for Scientific Research, Flanders (Fonds Wetenschappelijk Onderzoek (FWO) Vlaanderen), Belgium. R.H. and R.D.L. were funded by the NSF under grant nos. ABI-1062432 and CNS-0521433 to Indiana University, and by Indiana METACyt Initiative, which is supported in part by Lilly Endowment, Inc. J.B. was supported through a CSIRO eResearch Accelerated Computing Project. Any opinions, findings and conclusions or recommendations expressed in this article are those of the authors and do not necessarily reflect the views of any of the funding bodies and institutions including the National Science Foundation, the National Center for Genome Analysis Support and Indiana University.
Author information
Authors and Affiliations
Contributions
B.J.H. is the current lead developer of Trinity and is additionally responsible for the development of the companion in silico normalization and TransDecoder utilities described herein. M.Y. contributed to Butterfly software enhancements, generating figures and to the manuscript text. B.L. and C.N.D. developed RSEM and are responsible for enhancements related to improved Trinity support. B.J.H. and A.P. wrote the initial draft of the manuscript. A.R. is the Principal Investigator. All authors contributed to Trinity development and/or writing of the final manuscript, and all authors approved the final text.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Note
Supplementary materials for de novo transcript sequence reconstruction from RNA-seq: reference generation and analysis with Trinity. (PDF 699 kb)
Supplementary Figure 1
Defining minimum edge thresholds during initial Butterfly graph pruning. (PDF 554 kb)
Supplementary Figure 2
Butterfly's minimum support requirement for path extension during transcript reconstruction. (PDF 551 kb)
Supplementary Figure 3
Merging of insufficiently different path sequences. (PDF 530 kb)
Supplementary Figure 4
Enforcing path restrictions via triplet locking. (PDF 536 kb)
Supplementary Figure 5
Restrictions on the number of paths to be extended at each node. (PDF 540 kb)
Supplementary Figure 6
Evaluating assembly completeness for the S. pombe transcriptome. (PDF 636 kb)
Supplementary Figure 7
Evaluating assembly completeness for the mouse dendritic cell transcriptome. (PDF 584 kb)
Supplementary Figure 8
Correlation of expression values between reference transcripts and Trinity transcript components according to percent length agreement in S. pombe. (PDF 551 kb)
Supplementary Figure 9
Agreement between expression profiles calculated based on reference transcripts and trinity components at different S. pombe samples. (PDF 584 kb)
Rights and permissions
About this article
Cite this article
Haas, B., Papanicolaou, A., Yassour, M. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8, 1494–1512 (2013). https://doi.org/10.1038/nprot.2013.084
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.084
This article is cited by
-
Roast: a tool for reference-free optimization of supertranscriptome assemblies
BMC Bioinformatics (2024)
-
Genome-wide identification and expression pattern analysis of the kiwifruit GRAS transcription factor family in response to salt stress
BMC Genomics (2024)
-
Myrosinase isogenes in wasabi (Wasabia japonica Matsum) and their putative roles in glucosinolate metabolism
BMC Plant Biology (2024)
-
Girdling behavior of the longhorn beetle modulates the host plant to enhance larval performance
BMC Ecology and Evolution (2024)
-
Near-gapless and haplotype-resolved apple genomes provide insights into the genetic basis of rootstock-induced dwarfing
Nature Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.