Abstract
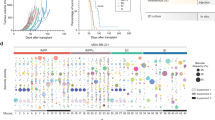
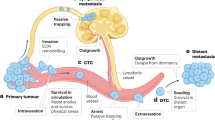
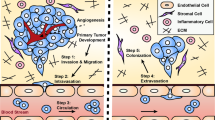
We describe a sensitive, robust, high-throughput method for quantifying the ability of metastatic tumor cells to colonize a secondary organ. Metastasis is the leading cause of death in cancer patients, and successful colonization of the secondary organ is the rate-limiting step in the metastatic process; thus, experimental methods that can be used to interrogate the key factors required for this critical step are of great importance. The experimental metastasis assay we detail here includes tail-vein injection of cancer cells into the mouse and determination of the resulting secondary organ colonization, primarily in the lung, 10 d post dosing. This assay can be used to investigate factors that regulate metastatic colonization both at the tumor-cell-intrinsic level (via manipulation of the tumor cells before injection) and at the tumor-cell-extrinsic level (such as the tissue microenvironment, via the use of genetically modified (GM) mice or agents such as antibodies or drugs). Using this method, we have robustly screened more than 950 GM mouse lines to identify novel microenvironmental regulators of metastatic colonization. The experimental details discussed here include choosing of appropriate cell numbers, handling of the cells, selection of recipient animals and injection techniques. Furthermore, we discuss key experimental design considerations, including the choice of the method used to determine metastatic burden and statistical analysis of the results, as well as provide troubleshooting tips and identification of factors that contribute to experimental variability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chambers, A.F., Naumov, G.N., Vantyghem, S.A. & Tuck, A.B. Molecular biology of breast cancer metastasis. Clinical implications of experimental studies on metastatic inefficiency. Breast Cancer Res. 2, 400–407 (2000).
Koop, S. et al. Independence of metastatic ability and extravasation: metastatic ras-transformed and control fibroblasts extravasate equally well. Proc. Natl. Acad. Sci. USA 93, 11080–11084 (1996).
Luzzi, K.J. et al. Multistep nature of metastatic inefficiency: dormancy of solitary cells after successful extravasation and limited survival of early micrometastases. Am. J. Pathol. 153, 865–873 (1998).
Zhao, Z.M. et al. Early and multiple origins of metastatic lineages within primary tumors. Proc. Natl. Acad. Sci. USA 113, 2140–2145 (2016).
Hurst, D.R. & Welch, D.R. Metastasis suppressor genes at the interface between the environment and tumor cell growth. Int. Rev. Cell Mol. Biol. 286, 107–180 (2011).
Park, Y.G. et al. Sipa1 is a candidate for underlying the metastasis efficiency modifier locus Mtes1. Nat. Genet. 37, 1055–1062 (2005).
van der Weyden, L. et al. Genome-wide in vivo screen identifies novel host regulators of metastatic colonization. Nature 541, 233–236 (2017).
Fidler, I.J. & Radinsky, R. Genetic control of cancer metastasis. J. Natl. Cancer Inst. 82, 166–168 (1990).
Goldberg, S.F., Harms, J.F., Quon, K. & Welch, D.R. Metastasis-suppressed C8161 melanoma cells arrest in lung but fail to proliferate. Clin. Exp. Metastasis 17, 601–607 (1999).
Chekmareva, M.A. et al. Chromosome 17-mediated dormancy of AT6.1 prostate cancer micrometastases. Cancer Res. 58, 4963–4969 (1998).
Quigley, J.P. & Armstrong, P.B. Tumor cell intravasation alu-cidated: the chick embryo opens the window. Cell 94, 281–284 (1998).
Heilmann, S. et al. A quantitative system for studying metastasis using transparent zebrafish. Cancer Res. 75, 4272–4282 (2015).
Khanna, C. & Hunter, K. Modeling metastasis in vivo. Carcinogenesis 26, 513–523 (2005).
Hoffman, R.M. Orthotopic metastatic mouse models for anticancer drug discovery and evaluation: a bridge to the clinic. Investig. New Drugs 17, 343–359 (1999).
Kuo, T.H. et al. Site-specific chemosensitivity of human small-cell lung carcinoma growing orthotopically compared to subcutaneously in SCID mice: the importance of orthotopic models to obtain relevant drug evaluation data. Anticancer Res. 13, 627–630 (1993).
Lucke, B., Breedis, C., Woo, Z.P., Berwick, L. & Nowell, P. Differential growth of metastatic tumors in liver and lung; experiments with rabbit V2 carcinoma. Cancer Res. 12, 734–738 (1952).
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989).
Fidler, I.J. & Kripke, M.L. Metastasis results from preexisting variant cells within a malignant tumor. Science 197, 893–895 (1977).
Hart, I.R. & Fidler, I.J. Role of organ selectivity in the determination of metastatic patterns of B16 melanoma. Cancer Res. 40, 2281–2287 (1980).
Steeg, P.S. Targeting metastasis. Nat. Rev. Cancer 16, 201–218 (2016).
Fidler, I.J. Metastasis: quantitative analysis of distribution and fate of tumor emboli labeled with 125 I-5-iodo-2′-deoxyuridine. J. Natl. Cancer Inst. 45, 773–782 (1970).
Seo, J.B., Im, J.G., Goo, J.M., Chung, M.J. & Kim, M.Y. Atypical pulmonary metastases: spectrum of radiologic findings. Radiographics 21, 403–417 (2001).
Rodero, M.P., Auvynet, C., Poupel, L., Combadiere, B. & Combadiere, C. Control of both myeloid cell infiltration and angiogenesis by CCR1 promotes liver cancer metastasis development in mice. Neoplasia 15, 641–648 (2013).
Orosz, P. et al. Promotion of experimental liver metastasis by tumor necrosis factor. Int. J. Cancer 60, 867–871 (1995).
Brunson, K.W. & Nicolson, G.L. Selection and biologic properties of malignant variants of a murine lymphosarcoma. J. Natl. Cancer Inst. 61, 1499–1503 (1978).
Edel, G. Mechanism of liver-specific metastatic tumor spread in a murine tumor model. J. Cancer Res. Clin. Oncol. 114, 47–58 (1988).
Karp, N.A., Melvin, D., Sanger Mouse Genetics Project & Mott, R.F. Robust and sensitive analysis of mouse knockout phenotypes. PLoS One 7, e52410 (2012).
Klerk, C.P. et al. Validity of bioluminescence measurements for noninvasive in vivo imaging of tumor load in small animals. Biotechniques 43, 7–13 (2007).
Nogawa, M. et al. Monitoring luciferase-labeled cancer cell growth and metastasis in different in vivo models. Cancer Lett. 217, 243–253 (2005).
Bouvet, M. & Hoffman, R.M. Tumor imaging technologies in mouse models. Methods Mol. Biol. 1267, 321–348 (2015).
Wessels, J.T. et al. In vivo imaging in experimental preclinical tumor research--a review. Cytometry A 71, 542–549 (2007).
Curran, P.J. & Hussong, A.M. Integrative data analysis: the simultaneous analysis of multiple data sets. Psychol. Methods 14, 81–100 (2009).
van der Weyden, L., Karp, N.A., Swiatkowska, A., Adams, D.J. & Speak, A.O. Genome wide in vivo mouse screen data from studies to assess host regulation of metastatic colonisation. Sci. Data http://dx.doi.org/10.1038/sdata.2017.129 (2017).
Acknowledgements
This work was supported by grants from Cancer Research UK (C20510/A13031 to D.J.A.), the Wellcome Trust (WT098051 to D.J.A.), ERC Combat Cancer (319661 to D.J.A.) and the National Institutes of Health (U54HG004028 to N.A.K.). The authors thank the members of the Research Support Facility (Wellcome Trust Genome Campus) for their care of the mice, and M. Nohadani for histological processing of the lung samples.
Author information
Authors and Affiliations
Contributions
L.v.d.W. and D.J.A. conceived the idea of screening mice from large-scale mouse phenotyping pipelines for metastatic colonization. L.v.d.W., A.S. and A.O.S. performed the experiments. N.A.K. performed statistical analysis. M.J.A. performed histopathology analysis. L.v.d.W. and A.O.S. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Speak, A., Swiatkowska, A., Karp, N. et al. A high-throughput in vivo screening method in the mouse for identifying regulators of metastatic colonization. Nat Protoc 12, 2465–2477 (2017). https://doi.org/10.1038/nprot.2017.118
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.118
This article is cited by
-
New insights into the correlations between circulating tumor cells and target organ metastasis
Signal Transduction and Targeted Therapy (2023)
-
CRISPR activation screen in mice identifies novel membrane proteins enhancing pulmonary metastatic colonisation
Communications Biology (2021)
-
Molecular alterations of cancer cell and tumour microenvironment in metastatic gastric cancer
Oncogene (2018)
-
Transparent reporting of experimental parameters in assays measuring phenotypic steps in metastasis
Clinical & Experimental Metastasis (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.