Abstract
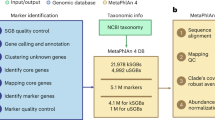
Metagenomic shotgun sequencing data can identify microbes populating a microbial community and their proportions, but existing taxonomic profiling methods are inefficient for increasingly large data sets. We present an approach that uses clade-specific marker genes to unambiguously assign reads to microbial clades more accurately and >50× faster than current approaches. We validated our metagenomic phylogenetic analysis tool, MetaPhlAn, on terabases of short reads and provide the largest metagenomic profiling to date of the human gut. It can be accessed at http://huttenhower.sph.harvard.edu/metaphlan/.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
DeLong, E.F. Nat. Rev. Microbiol. 3, 459–469 (2005).
Daniel, R. Nat. Rev. Microbiol. 3, 470–478 (2005).
The Human Microbiome Project Consortium. Nature advance online publication, doi:10.1038/nature11209 (14 June 2012).
Qin, J. et al. Nature 464, 59–65 (2010).
Ravel, J. et al. Proc. Natl. Acad. Sci. USA 108, 4680–4687 (2011).
Veiga, P. et al. Proc. Natl. Acad. Sci. USA 107, 18132–18137 (2010).
Turnbaugh, P.J. et al. Nature 457, 480–484 (2009).
Markowitz, V.M. et al. Nucleic Acids Res. 38, D382–D390 (2010).
Fredricks, D.N., Fiedler, T.L. & Marrazzo, J.M. N. Engl. J. Med. 353, 1899–1911 (2005).
Stewart, F.J., Ulloa, O. & DeLong, E.F. Environ. Microbiol. 14, 23–40 (2012).
Arumugam, M. et al. Nature 473, 174–180 (2011).
Brady, A. & Salzberg, S. Nat. Methods 8, 367 (2011).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. J. Mol. Biol. 215, 403–410 (1990).
Parks, D.H., MacDonald, N. & Beiko, R. BMC Bioinformatics 12, 328 (2011).
Rosen, G.L., Reichenberger, E.R. & Rosenfeld, A.M. Bioinformatics 27, 127–129 (2011).
Segata, N. & Huttenhower, C. PLoS ONE 6, e24704 (2011).
Bohlin, J. et al. BMC Evol. Biol. 10, 249 (2010).
Edgar, R.C. Bioinformatics 26, 2460–2461 (2010).
Wu, M. & Eisen, J.A. Genome Biol. 9, R151 (2008).
Ciccarelli, F.D. et al. Science 311, 1283–1287 (2006).
Mavromatis, K. et al. Nat. Methods 4, 495–500 (2007).
Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. Nucleic Acids Res. 38, D355–D360 (2010).
Li, H., Ruan, J. & Durbin, R. Genome Res. 18, 1851–1858 (2008).
Pruitt, K.D., Tatusova, T., Klimke, W. & Maglott, D.R. Nucleic Acids Res. 37, D32–D36 (2009).
Huson, D.H., Auch, A.F., Qi, J. & Schuster, S.C. Genome Res. 17, 377–386 (2007).
Huson, D.H., Mitra, S., Ruscheweyh, H.J., Weber, N. & Schuster, S.C. Genome Res. 21, 1552–1560 (2011).
Gori, F., Folino, G., Jetten, M.S.M. & Marchiori, E. Bioinformatics 27, 196–203 (2011).
Berger, S.A. & Stamatakis, A. Bioinformatics 27, 2068–2075 (2011).
Gerlach, W. & Stoye, J. Nucleic Acids Res. 39, e91 (2011).
McHardy, A.C., Rigoutsos, I., Hugenholtz, P., Tsirigos, A. & Martin, H.G. Nat. Methods 4, 63–72 (2007).
Patil, K.R. et al. Nat. Methods 8, 191–192 (2011).
Brady, A. & Salzberg, S.L. Nat. Methods 6, 673–676 (2009).
Rosen, G., Garbarine, E., Caseiro, D., Polikar, R. & Sokhansanj, B. Adv. Bioinformatics 2008, 205969 (2008).
Nalbantoglu, O.U., Way, S.F., Hinrichs, S.H. & Sayood, K. BMC Bioinformatics 12, 41 (2011).
Leung, H.C. et al. Bioinformatics 27, 1489–1495 (2011).
Schloss, P.D. et al. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Cole, J.R. et al. Nucleic Acids Res. 37, D141–D145 (2009).
Acknowledgements
We would like to thank F. Stewart and E. DeLong for their helpful input during this study; D. Gevers, S. Sykes and K. Huang for their feedback on the methodology and J. Reyes and G. Weingart for their assistance with the implementation. This work was supported by US National Institutes of Health grant 1R01HG005969 and National Science Foundation grant DBI-1053486 to C.H.
Author information
Authors and Affiliations
Contributions
N.S., A.B., O.J. and C.H. conceived the method; N.S. implemented the software; N.S. and C.H. performed the experiments; N.S., L.W., V.N. and C.H. analyzed the data; and N.S. and C.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1 and 2 and Supplementary Notes 1–3 (PDF 2101 kb)
Rights and permissions
About this article
Cite this article
Segata, N., Waldron, L., Ballarini, A. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods 9, 811–814 (2012). https://doi.org/10.1038/nmeth.2066
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2066
This article is cited by
-
MetaPro: a scalable and reproducible data processing and analysis pipeline for metatranscriptomic investigation of microbial communities
Microbiome (2023)
-
Exploring the microbial diversity and characterization of cellulase and hemicellulase genes in goat rumen: a metagenomic approach
BMC Biotechnology (2023)
-
Short-term feeding of defatted bovine colostrum mitigates inflammation in the gut via changes in metabolites and microbiota in a chicken animal model
Animal Microbiome (2023)
-
Differential responses of the gut microbiome and resistome to antibiotic exposures in infants and adults
Nature Communications (2023)
-
Modulation of the gut microbiome with nisin
Scientific Reports (2023)