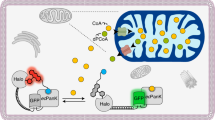
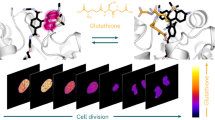
Abstract
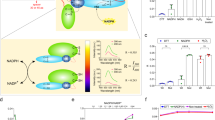
Reduced nicotinamide adenine dinucleotide phosphate (NADPH) is essential for biosynthetic reactions and antioxidant functions; however, detection of NADPH metabolism in living cells remains technically challenging. We develop and characterize ratiometric, pH-resistant, genetically encoded fluorescent indicators for NADPH (iNap sensors) with various affinities and wide dynamic range. iNap sensors enabled quantification of cytosolic and mitochondrial NADPH pools that are controlled by cytosolic NAD+ kinase levels and revealed cellular NADPH dynamics under oxidative stress depending on glucose availability. We found that mammalian cells have a strong tendency to maintain physiological NADPH homeostasis, which is regulated by glucose-6-phosphate dehydrogenase and AMP kinase. Moreover, using the iNap sensors we monitor NADPH fluctuations during the activation of macrophage cells or wound response in vivo. These data demonstrate that the iNap sensors will be valuable tools for monitoring NADPH dynamics in live cells and gaining new insights into cell metabolism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
26 July 2017
In the html version of this article initially published online, Haiyan Liu was incorrectly listed as the sole corresponding author. Haiyan Liu and Yi Yang should be listed as corresponding authors. The error has been corrected in the html version as of 26 July 2017.
References
Gorrini, C., Harris, I.S. & Mak, T.W. Modulation of oxidative stress as an anticancer strategy. Nat. Rev. Drug Discov. 12, 931–947 (2013).
Schulze, A. & Harris, A.L. How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature 491, 364–373 (2012).
Vallance, P. & Leiper, J. Blocking NO synthesis: how, where and why? Nat. Rev. Drug Discov. 1, 939–950 (2002).
Trachootham, D., Alexandre, J. & Huang, P. Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nat. Rev. Drug Discov. 8, 579–591 (2009).
Pollak, N., Niere, M. & Ziegler, M. NAD kinase levels control the NADPH concentration in human cells. J. Biol. Chem. 282, 33562–33571 (2007).
Nóbrega-Pereira, S. et al. G6PD protects from oxidative damage and improves healthspan in mice. Nat. Commun. 7, 10894 (2016).
MacDonald, M.J. Feasibility of a mitochondrial pyruvate malate shuttle in pancreatic islets. Further implication of cytosolic NADPH in insulin secretion. J. Biol. Chem. 270, 20051–20058 (1995).
Blacker, T.S. et al. Separating NADH and NADPH fluorescence in live cells and tissues using FLIM. Nat. Commun. 5, 3936 (2014).
Zhao, Y. et al. Genetically encoded fluorescent sensors for intracellular NADH detection. Cell Metab. 14, 555–566 (2011).
Zhao, Y. et al. SoNar, a highly responsive NAD+/NADH sensor, allows high-throughput metabolic screening of anti-tumor agents. Cell Metab. 21, 777–789 (2015).
Bilan, D.S. et al. Genetically encoded fluorescent indicator for imaging NAD(+)/NADH ratio changes in different cellular compartments. Biochim. Biophys. Acta 1840, 951–957 (2014).
Hung, Y.P., Albeck, J.G., Tantama, M. & Yellen, G. Imaging cytosolic NADH-NAD(+) redox state with a genetically encoded fluorescent biosensor. Cell Metab. 14, 545–554 (2011).
Cambronne, X.A. et al. Biosensor reveals multiple sources for mitochondrial NAD+. Science 352, 1474–1477 (2016).
Cameron, W.D. et al. Apollo-NADP+: a spectrally tunable family of genetically encoded sensors for NADP+. Nat. Methods 13, 352–358 (2016).
Zhao, Y. et al. In vivo monitoring of cellular energy metabolism using SoNar, a highly responsive sensor for NAD+/NADH redox state. Nat. Protoc. 11, 1345–1359 (2016).
Zhao, Y. & Yang, Y. Real-time and high-throughput analysis of mitochondrial metabolic states in living cells using genetically encoded NAD+/NADH sensors. Free Radic. Biol. Med. 100, 43–52 (2016).
Powers, R. et al. Comparison of protein active site structures for functional annotation of proteins and drug design. Proteins 65, 124–135 (2006).
Yoshikawa, S. et al. Structure of archaeal glyoxylate reductase from Pyrococcus horikoshii OT3 complexed with nicotinamide adenine dinucleotide phosphate. Acta Crystallogr. D Biol. Crystallogr. 63, 357–365 (2007).
Nagai, T., Sawano, A., Park, E.S. & Miyawaki, A. Circularly permuted green fluorescent proteins engineered to sense Ca2+. Proc. Natl. Acad. Sci. USA 98, 3197–3202 (2001).
Hedeskov, C.J., Capito, K. & Thams, P. Cytosolic ratios of free [NADPH]/[NADP+] and [NADH]/[NAD+] in mouse pancreatic islets, and nutrient-induced insulin secretion. Biochem. J. 241, 161–167 (1987).
Veech, R.L., Eggleston, L.V. & Krebs, H.A. The redox state of free nicotinamide-adenine dinucleotide phosphate in the cytoplasm of rat liver. Biochem. J. 115, 609–619 (1969).
Ohashi, K., Kawai, S. & Murata, K. Identification and characterization of a human mitochondrial NAD kinase. Nat. Commun. 3, 1248 (2012).
Tantama, M., Martínez-François, J.R., Mongeon, R. & Yellen, G. Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio. Nat. Commun. 4, 2550 (2013).
Noda, T. & Amano, F. Differences in nitric oxide synthase activity in a macrophage-like cell line, RAW264.7 cells, treated with lipopolysaccharide (LPS) in the presence or absence of interferon-γ (IFN-γ): possible heterogeneity of iNOS activity. J. Biochem. 121, 38–46 (1997).
Takanaga, H., Chaudhuri, B. & Frommer, W.B. GLUT1 and GLUT9 as major contributors to glucose influx in HepG2 cells identified by a high sensitivity intramolecular FRET glucose sensor. Biochim. Biophys. Acta 1778, 1091–1099 (2008).
Dooley, C.T. et al. Imaging dynamic redox changes in mammalian cells with green fluorescent protein indicators. J. Biol. Chem. 279, 22284–22293 (2004).
Pandolfi, P.P. et al. Targeted disruption of the housekeeping gene encoding glucose 6-phosphate dehydrogenase (G6PD): G6PD is dispensable for pentose synthesis but essential for defense against oxidative stress. EMBO J. 14, 5209–5215 (1995).
Au, S.W., Gover, S., Lam, V.M. & Adams, M.J. Human glucose-6-phosphate dehydrogenase: the crystal structure reveals a structural NADP+ molecule and provides insights into enzyme deficiency. Structure 8, 293–303 (2000).
Woods, A. et al. Ca2+/calmodulin-dependent protein kinase kinase-β acts upstream of AMP-activated protein kinase in mammalian cells. Cell Metab. 2, 21–33 (2005).
Tsou, P., Zheng, B., Hsu, C.H., Sasaki, A.T. & Cantley, L.C. A fluorescent reporter of AMPK activity and cellular energy stress. Cell Metab. 13, 476–486 (2011).
Oakhill, J.S. et al. AMPK is a direct adenylate charge-regulated protein kinase. Science 332, 1433–1435 (2011).
Ermakova, Y.G. et al. Red fluorescent genetically encoded indicator for intracellular hydrogen peroxide. Nat. Commun. 5, 5222 (2014).
Niethammer, P., Grabher, C., Look, A.T. & Mitchison, T.J. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 459, 996–999 (2009).
Belousov, V.V. et al. Genetically encoded fluorescent indicator for intracellular hydrogen peroxide. Nat. Methods 3, 281–286 (2006).
Berg, J., Hung, Y.P. & Yellen, G. A genetically encoded fluorescent reporter of ATP:ADP ratio. Nat. Methods 6, 161–166 (2009).
Pomorski, A., Kochanńczyk, T., Miłoch, A. & Kręzżel, A. Method for accurate determination of dissociation constants of optical ratiometric systems: chemical probes, genetically encoded sensors, and interacting molecules. Anal. Chem. 85, 11479–11486 (2013).
Pavlova, N.N. & Thompson, C.B. The emerging hallmarks of cancer metabolism. Cell Metab. 23, 27–47 (2016).
Jeon, S.M., Chandel, N.S. & Hay, N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature 485, 661–665 (2012).
Cool, B. et al. Identification and characterization of a small molecule AMPK activator that treats key components of type 2 diabetes and the metabolic syndrome. Cell Metab. 3, 403–416 (2006).
Yavari, A. et al. Chronic activation of γ2 AMPK induces obesity and reduces β cell function. Cell Metab. 23, 821–836 (2016).
Burkewitz, K., Zhang, Y. & Mair, W.B. AMPK at the nexus of energetics and aging. Cell Metab. 20, 10–25 (2014).
Shackelford, D.B. & Shaw, R.J. The LKB1-AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009).
Thliveris, A.T., Little, J.W. & Mount, D.W. Repression of the E. coli recA gene requires at least two LexA protein monomers. Biochimie 73, 449–456 (1991).
Zhao, Y. et al. An expanded palette of genetically encoded Ca2+ indicators. Science 333, 1888–1891 (2011).
Llopis, J., McCaffery, J.M., Miyawaki, A., Farquhar, M.G. & Tsien, R.Y. Measurement of cytosolic, mitochondrial, and Golgi pH in single living cells with green fluorescent proteins. Proc. Natl. Acad. Sci. USA 95, 6803–6808 (1998).
Tian, W.N. et al. Importance of glucose-6-phosphate dehydrogenase activity for cell growth. J. Biol. Chem. 273, 10609–10617 (1998).
Outten, C.E. & Culotta, V.C. A novel NADH kinase is the mitochondrial source of NADPH in Saccharomyces cerevisiae. EMBO J. 22, 2015–2024 (2003).
Wagner, T.C. & Scott, M.D. Single extraction method for the spectrophotometric quantification of oxidized and reduced pyridine nucleotides in erythrocytes. Anal. Biochem. 222, 417–426 (1994).
Acknowledgements
We thank S.J. Remington for the roGFP1 vector; G. Yellen for the PercevalHR vector; V.V. Belousov for the HyPerRed vector; C. Fan, J. Ye, Y. Tang, T. Wang, Z. Wu, H. Hu, Z. Du, T. Li, Y. Xia, L. Zhou and Y. Wang for technical assistance; and S. Tribuna for secretarial assistance. This research was supported by the National Natural Science Foundation of China (31225008, 91313301 and 31470833 to Y.Y.; 31370755 to H.L.; 91649123 and 31671484 to Y. Zhao), 973 Program (2013CB531200 to Y.Y.), the Shanghai Science and Technology Commission (14XD1401400 and 16430723100 to Y.Y. and 15YF1402600 to Y. Zhao), the Specialized Research Fund for the Doctoral Program of Higher Education (20100074110010 to Y.Y.), the Lift Engineering for Young Talent of China Association for Science and Technology (to Y. Zhao), Shanghai Young Top-notch Talent (to Y. Zhao), the State Key Laboratory of Bioreactor Engineering (to Y.Y.), the 111 Project (B07023 to Y.Y.), the Fundamental Research Funds for the Central Universities (to Y.Y. and Y. Zhao) and the US National Institutes of Health (HL061795, HL007690 and GM107618 to J.L.).
Author information
Authors and Affiliations
Contributions
Y.Y., H.L., H.C. and R.T. designed and validated the sensors in vitro. Y.Y., R.T. and Y. Zhao designed the live cell imaging, metabolic assays and signaling study experiments. R.T., Y. Zhao, A.W., J.Z., X.C., Y. Zou, M.S., R.L. and N.S. performed experiments. J.D., H.-M.Z., L.Z., X.Q., and J.L. gave technical support and conceptual advice. Y.Y., H.L., Y. Zhao, R.T., H.C. and J.L. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Generation of SoNar-based NADPH sensors.
(a) The proportion(%) of charged amino acid residues around the 2'-hydroxyl group of the ligand in NADP(H) and NAD(H) binding proteins. Most NADP(H) binding proteins contain positively charged residues around the phosphate, while the NAD(H) binding proteins favor a negatively charged environment at the corresponding site. The proteins selected for analysis are given in Supplementary Table 1. (b) The NADH binding site of T-Rex (carbon atoms in yellow, PDB ID:1xcb) superimposed with the NADP binding site of glyoxylate reductase from P. horikoshii (carbon atoms in cyan, PDB ID: 2dbr). The structure of glyoxylate reductase has been selected because the geometrical arrangement of its ligand binding site is highly similar to that of T-Rex (according to CPASS), despite that it’s binding to NADPH rather than NADH. Ligands and side chains are shown as sticks, with oxygen atoms in red, nitrogen in blue, and phosphorous in orange. (c) The proportion(%) of nonpolar residues in the adenine binding pocket of protein structures containing bound NAD(H) and NADP(H), respectively. For a given protein structure, amino acid residues with at least one atom within 5 Å from any adenine atom were identified to calculate the proportion of nonpolar residues. Val, Pro, Trp, Gly, Ala, Met, Phe, Leu, Ile or Cys are counted as nonpolar residues. The proteins used for analysis are listed in Supplementary Table 1.
Supplementary Figure 2 Fluorescence properties of iNap sensors.
(a) Fluorescence response of SoNar and 40 mutants listed in Supplementary Table 3 to 1.0 mM NADPH and its analogs, 4 mM ADP, 4 mM ATP, 4 mM AMP, 0.5 mM NAD+, 50 μM NADH, and 10 μM NADP+. The 3rd, 8th, 31st, 39th and 40th mutants were denoted iNap3, iNap2, iNap1, iNap4 and iNapc, respectively. The ratio of fluorescence excited at 420 nm and 485 nm shown as a heatmap. (b) The initial excitation ratios of different iNap sensors at pH 7.3. (c and d) Emission spectra of purified iNap1 in the control condition (black), and after addition of 0.2 mM NADPH (orange), normalized to the peak intensity in the control condition. Excitation was fixed at 420 nm (c) and 490 nm (d), respectively. (e) Fluorescence response of iNap1 corrected by iNapc to 0.2 mM NADPH at various temperatures. pH correction is determined by dividing the fluorescence ratio (R420/485) of iNap1 by that of iNapc. (f) iNap1 fluorescence plotted against NADPH in the presence of NADH (50 μM), NAD+ (500 μM), and NADP+ (10 μM). (g) iNap1 fluorescence plotted against NADPH in the presence of ADP (1 mM, 2 mM or 4 mM) and ATP (1 mM, 2 mM or 4 mM). (h) iNap1 fluorescence plotted against NADPH at the indicated ATP: ADP ratios; the total adenine nucleotide concentration was 2 or 4 mM. In supplementary Fig. 2f-h, fluorescence ratios were normalized to the value at the saturating concentration of NADPH at pH 7.3, experiments were performed in triplicate, and the data were from the three independent detections, error bars represent SEM.
Supplementary Figure 3 The pH sensitivity of iNap1 sensors.
(a) Fluorescence ratios (R420/590) of mCherry-iNap1 plotted against NADPH at the indicated pH. The fusion construct allows ratiometric and pH-resistant measurement of NADPH. (b) Fluorescence intensities of iNap1 and iNapc with excitation at 420 nm or 485 nm, and emission at 528 nm. Data were normalized to the fluorescence at pH 7.0. (c) Fluorescence intensity of iNap1 when excited at 485 nm plotted against the NADPH concentration at the indicated pH. (d) NADPH titration curves of iNap1 corrected by iNapc when excited at 485 nm at the indicated pH. (e) NADPH titration curves of iNap1 normalized by iNapc at the indicated pH. (f and g) pH-correction of the excitation ratio 420/485 nm of iNap1 with iNapc in the absence (f) and presence (g) of 0.2 mM NADPH. Data were normalized to the fluorescence ratio at pH 7.0. (h) Fluorescence intensity of iNapc when excited at 485 nm plotted against the NADPH concentration at the indicated pH. In supplementary Fig. 3a-h, experiments were performed in triplicate and the data were from the three independent detections, error bars represent SEM.
Supplementary Figure 4 Fluorescence intensity and homogeneity of iNap1 in mammalian cells.
(a) Comparison of the fluorescence intensity of different genetically encoded fluorescent sensors in HeLa cells. (b and c) The excitation spectra of HeLa expressing iNap1 cells (b) or control cells (c) with or without 160 μM diamide. (d-f) The homogeneity of iNap1 (d), iNapc (e) and mCherry-iNap1(f) fluorescence in individual cells. Quantitative data of the sensor’s fluorescence were obtained from at least 200 cells. Error bars represent SD. Scale bar 20 μm.
Supplementary Figure 5 Subcellular distribution and regulation of NADPH in mammalian cells.
(a and b) Quantification of iNap1 (a) and iNap3-Mit (b) fluorescence in NADK overexpressing or knockdown cells. Data are normalized to digitonin-permeabilized cell samples in the absence of NADPH. Error bars represent SEM. (c and d) NADK knockdown or overexpression (OE) in HeLa cells was assessed by Western blot (c) or activity analysis (d). (e and f) Effect of NADK knockdown or overexpression (OE) on whole-cell (e) and mitochondrial (f) NADPH levels in HeLa cells. Data obtained by a single extraction method and in vitro biochemical assay. (g and h) Ratiometric fluorescence imaging (g) and quantification by microplate reader (h) of mCherry-iNap1 or Mit-mCherry-iNap3 in wide-type and NADK-overexpressing HeLa cells. Scale bar, 10 μm. (i) Fluorescence response of cells expressing iNap1 or PercevalHR to the glycolysis inhibitor 2-DG (2 mM). Fluorescence was measured 1 hr after 2-DG treatment and normalized to the control condition. (j) G6PD knockdown or PGD knockdown in HeLa cells was assessed by Western blot. (k) Effect of G6PD knockdown or PGD knockdown on cytosolic NADPH levels assessed by iNap1 fluorescence. (l and m) NADPH detection of in resting (Ctrl) or activated (LPS plus IFN-γ stimulation) RAW264.7 cells by flow cytometry (l) or microplate reader (m). In supplementary Fig. 5 a, b, i and k, R is normalized by iNapc. In supplementary Fig. 5a, b, d, e, f, k, m, experiments were performed in triplicate and the data were from the three independent detections, unpaired t-test. *p < 0.05, **p < 0.01, and ***p < 0.001. All experiments were performed in HeLa cells; error bars represent SEM unless otherwise indicated.
Supplementary Figure 6 NADPH dynamics of glucose-deprived cells in the setting of oxidative stress.
(a) Kinetics of FLII12 Pglu-700μ, SoNar, iNap1, or roGFP1 fluorescence in HeLa cells in response to glucose deprivation. (b) Kinetics of iNap1 and roGFP1 fluorescence in glucose-deprived HeLa cells in response to 80 μM H2O2. (c) Effect of the glutathione reductase and thioredoxin reductase inhibitor BCNU on diamide-induced NADPH oxidation in glucose-deprived HeLa cells. Cells were pre-treated with BCNU for 30 min. (d) Effect of DHEA on G6PD activity and PGD activity. The cell lysate was pre-incubated with DHEA for 1 hr. (e) PGD knockdown would not affect NADPH recovery under oxidation. In supplementary Fig. 6 a, b, c and e, R is normalized by iNapc. All experiments were performed in glucose-deprivation conditions with triplicate and the data were from the three independent detections. Error bars represent SEM.
Supplementary Figure 7 NADPH dynamics of glucose-fed cells in the setting of oxidative stress.
(a) Quantification of iNap1 or iNapc fluorescence in HeLa cells before and 40 min after treatment with 40 μM diamide; data derived from Fig. 4a. (b) Kinetics of iNap1 or roGFP1 fluorescence in glucose-fed HeLa cells in response to 80 μM H2O2. (c) Effect of the glutathione reductase and thioredoxin reductase inhibitor BCNU on diamide-induced NADPH oxidation in glucose-fed HeLa cells. Cells were pre-treated with BCNU for 30 min. (d) Effect of different metabolites (glutamine or pyruvate, 2 mM; isocitrate, malate or glucose, 10 mM) on diamide-induced NADPH oxidation in glucose-fed HeLa cells. Data were normalized to samples in the absence of metabolite. (e) Dose-dependent fluorescence response of cytosolic iNap1 and SoNar to different concentrations of glucose in diamide-treated HeLa cells. Fluorescence was measured immediately after diamide addition. Data were normalized to diamide-treated samples in the absence of glucose. In supplementary Fig. 7 b, c, d and e, R is normalized by iNapc, experiments were performed in triplicate and the data were from the three independent detections. Error bars represent SEM.
Supplementary Figure 8 NADPH metabolism in diamide-treated cells.
(a) The effect of ionomycin on diamide-induced NADPH oxidation in glucose-fed cells at 35 min. Data from Fig. 5c. R is normalized by iNapc. Unpaired t-test. *p < 0.05. (b) Effect of dorsomorphin on NADPH recovery in glucose-deprived cells under oxidative stress. (c) Working models for NADPH regulation in glucose-deprived cells under oxidative stress. In supplementary Fig. 8a, b, experiments were performed in triplicate and the data were from the three independent detections.
Supplementary Figure 9 Imaging of iNapc in zebrafish larvae.
No pH variation was observed in wound margin of iNapc-expressing zebrafish larvae. Tail-fin tip amputation was performed at the 0 min time point. Scale bar, 50 μm.
Supplementary Figure 10 The central nodes for cellular NADPH metabolism.
NADPH is mainly consumed through redox defense, reductive biosynthesis, and mutant isocitrate dehydrogenase (mIDH1/2) in cells. NADPH is regenerated primarily via the oxidative pentose phosphate pathway (PPP) and other redox reactions catalyzed by malic enzyme (ME1/3), IDH1/2, methylene tetrahydrofolate dehydrogenase (MTHFD1/2) and nicotinamide nucleotide transhydrogenase (NNT). NADP+ can be generated from NAD+ by cytosolic (NADK) or mitochondrial (NADK2) forms of NAD kinases. GSSG, oxidized glutathione; GSH, reduced glutathione; Arg, arginine; Glu, glutamate; Pro, proline; NTP, nucleotide triphosphate; dNTP, deoxynucleoside triphosphate; α-KG, alpha-ketoglutarate; 2-HG, 2-hydroxyglutarate; G-6-P, Glucose 6-phosphate; R-5-P, Ribulose 5-phosphate; THF, tetrahydrofolate; me-THF, methenyl/methylene-THF; for-THF, formyl-THF; GR, glutathione reductase (GR); FASN, fatty acid synthase; G6PD, glucose-6-phosphate dehydrogenase; 6PGD, 6-phosphogluconate dehydrogenase.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–4
Rights and permissions
About this article
Cite this article
Tao, R., Zhao, Y., Chu, H. et al. Genetically encoded fluorescent sensors reveal dynamic regulation of NADPH metabolism. Nat Methods 14, 720–728 (2017). https://doi.org/10.1038/nmeth.4306
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4306
This article is cited by
-
Comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo using highly responsive biosensors
Nature Protocols (2024)
-
Surplus fatty acid synthesis increases oxidative stress in adipocytes and induces lipodystrophy
Nature Communications (2024)
-
Visualizing subcellular changes in the NAD(H) pool size versus redox state using fluorescence lifetime imaging microscopy of NADH
Communications Biology (2024)
-
A short guide on blue fluorescent proteins: limits and perspectives
Applied Microbiology and Biotechnology (2024)
-
Electrochemiluminescent imaging of a NADH-based enzymatic reaction confined within giant liposomes
Analytical and Bioanalytical Chemistry (2024)