Abstract
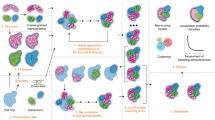
We describe a method that integrates data derived from different mass spectrometry (MS)-based techniques with a modeling strategy for structural characterization of protein assemblies. We encoded structural data derived from native MS, bottom-up proteomics, ion mobility–MS and chemical cross-linking MS into modeling restraints to compute the most likely structure of a protein assembly. We used the method to generate near-native models for three known structures and characterized an assembly intermediate of the proteasomal base.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Robinson, C.V., Sali, A. & Baumeister, W. Nature 450, 973–982 (2007).
Alber, F. et al. Nature 450, 683–694 (2007).
Stengel, F., Aebersold, R. & Robinson, C.V. Mol. Cell. Proteomics 11, R111.014027 (2012).
Lasker, K. et al. Proc. Natl. Acad. Sci. USA 109, 1380–1387 (2012).
Hall, Z., Politis, A. & Robinson, C.V. Structure 20, 1596–1609 (2012).
Politis, A. et al. PLoS ONE 5, e12080 (2010).
Leitner, A. et al. Mol. Cell. Proteomics 9, 1634–1649 (2010).
Walzthoeni, T., Leitner, A., Stengel, F. & Aebersold, R. Curr. Opin. Struct. Biol. 23, 252–260 (2013).
Ruotolo, B.T. et al. Nat. Protoc. 3, 1139–1152 (2008).
Leitner, A., Walzthoeni, T. & Aebersold, R. Nat. Protoc. 9, 120–137 (2014).
Walzthoeni, T. et al. Nat. Methods 9, 901–903 (2012).
Rinner, O. et al. Nat. Methods 5, 315–318 (2008).
Lander, G.C. et al. Nature 482, 186–191 (2012).
Kao, A. et al. Mol. Cell. Proteomics 11, 1566–1577 (2012).
Bohn, S. et al. Biochem. Biophys. Res. Commun. 435, 250–254 (2013).
Nakamura, Y. et al. Biochem. Biophys. Res. Commun. 359, 503–509 (2007).
Barrault, M.B. et al. Proc. Natl. Acad. Sci. USA 109, E1001–E1010 (2012).
Saeki, Y. et al. Cell 137, 900–913 (2009).
Roelofs, J. et al. Nature 459, 861–865 (2009).
Tomko, R.J. Jr. et al. Mol. Cell 38, 393–403 (2010).
McCormick, M.S., Sazinsky, M.H., Condon, K.L. & Lippard, S.J. J. Am. Chem. Soc. 128, 15108–15110 (2006).
Jabri, E. & Karplus, A. Biochemistry 35, 10616–10626 (1996).
Sakata, E. et al. Mol. Cell 42, 637–649 (2011).
Ghaemmaghami, S. et al. Nature 425, 737–741 (2003).
Sobott, F., Hernández, H., McCammon, M.G., Tito, M.A. & Robinson, C.V. Anal. Chem. 74, 1402–1407 (2002).
Hernández, H. & Robinson, C.V. Nat. Protoc. 2, 715–726 (2007).
Pringle, S.D. et al. Int. J. Mass Spectrom. 261, 1–12 (2007).
Kemper, P.R., Dupuis, N.F. & Bowers, M.T. Int. J. Mass Spectrom. 287, 46–57 (2009).
Bush, M.F. et al. Anal. Chem. 82, 9557–9565 (2010).
Russel, D. et al. PLoS Biol. 10, e1001244 (2012).
Alber, F., Kim, M.F. & Sali, A. Structure 13, 435–445 (2005).
Alber, F., Forster, F., Korkin, D., Topf, M. & Sali, A. Annu. Rev. Biochem. 77, 443–477 (2008).
Johnson, S.C. Psychometrika 32, 241–254 (1967).
Phillips, J.C. et al. J. Comput. Chem. 26, 1781–1802 (2005).
Pettersen, E.F. et al. J. Comput. Chem. 25, 1605–1612 (2004).
Shen, M.-y. & Sali, A. Protein Sci. 15, 2507–2524 (2006).
Šali, A. & Blundell, T.L. J. Mol. Biol. 234, 779–815 (1993).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
MMOH and ToMOH were a gift of S.J. Lippard (Massachusetts Institute of Technology). Urease from K. aerogenes was a gift from R.P. Hausinger (Michigan State University). This work was supported by funding from PROSPECTS (Proteomics Specification in Space and Time Grant HEALTH-F4-2008-201648) within the European Union 7th Framework Program (A.P., C.V.R. and R.A.) and from European Research Council advanced grants “Proteomics v3.0” (233226) and “IMPRESS” (268851) to R.A. and C.V.R. H.H. is funded by Medical Research Council programme grant (G1000819). F.S. is a Sir Henry Wellcome Fellow funded by the Wellcome Trust (grant 095951), and C.V.R. is funded by the Royal Society.
Author information
Authors and Affiliations
Contributions
F.S. and A.P. conceived the study; F.S., A.P., C.V.R. and R.A. designed the research; A.P. performed all modeling and developed the software; F.S. carried out the experiments; Z.H. and H.H. performed part of the IM-MS and native MS experiments. A.L. and T.W. supported CX-MS experiments and analysis; F.S. and A.P. analyzed the data; A.P., F.S., C.V.R. and R.A. wrote the paper; all authors commented on and edited the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–30, Supplementary Tables 1–12, Supplementary Results and Supplementary Notes 1–3 (PDF 14099 kb)
Supplementary Software
Software package for modeling protein complexes from hybrid mass spectrometry data (ZIP 217 kb)
Source data
Rights and permissions
About this article
Cite this article
Politis, A., Stengel, F., Hall, Z. et al. A mass spectrometry–based hybrid method for structural modeling of protein complexes. Nat Methods 11, 403–406 (2014). https://doi.org/10.1038/nmeth.2841
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2841
This article is cited by
-
Advances in mass spectrometry to unravel the structure and function of protein condensates
Nature Protocols (2023)
-
On the Effect of Sphere-Overlap on Super Coarse-Grained Models of Protein Assemblies
Journal of the American Society for Mass Spectrometry (2019)
-
Structural prediction of protein models using distance restraints derived from cross-linking mass spectrometry data
Nature Protocols (2018)
-
A cross-linking/mass spectrometry workflow based on MS-cleavable cross-linkers and the MeroX software for studying protein structures and protein–protein interactions
Nature Protocols (2018)
-
Disentangling constraints using viability evolution principles in integrative modeling of macromolecular assemblies
Scientific Reports (2017)