Abstract
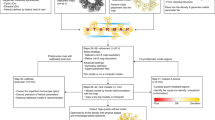
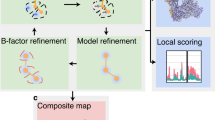
Integrative modeling enables structure determination of macromolecular complexes by combining data from multiple experimental sources such as X-ray crystallography, electron microscopy or cross-linking mass spectrometry. It is particularly useful for complexes not amenable to high-resolution electron microscopy—complexes that are flexible, heterogeneous or imaged in cells with cryo-electron tomography. We have recently developed an integrative modeling protocol that allowed us to model multi-megadalton complexes as large as the nuclear pore complex. Here, we describe the Assembline software package, which combines multiple programs and libraries with our own algorithms in a streamlined modeling pipeline. Assembline builds ensembles of models satisfying data from atomic structures or homology models, electron microscopy maps and other experimental data, and provides tools for their analysis. Compared with other methods, Assembline enables efficient sampling of conformational space through a multistep procedure, provides new modeling restraints and includes a unique configuration system for setting up the modeling project. Our protocol achieves exhaustive sampling in less than 100–1,000 CPU-hours even for complexes in the megadalton range. For larger complexes, resources available in institutional or public computer clusters are needed and sufficient to run the protocol. We also provide step-by-step instructions for preparing the input, running the core modeling steps and assessing modeling performance at any stage.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All datasets needed for the step-by-step tutorials (i.e., for yeast NPC and Elongator complex modeling) are provided at https://git.embl.de/rantos/scnpc_tutorial.git (for the yeast NPC) and https://git.embl.de/kosinski/elongator_tutorial.git (for the Elongator complex).
Code availability
The Assembline software is freely available as an open-source Python package (https://www.embl-hamburg.de/Assembline/), which can be installed from source code (https://git.embl.de/kosinski/Assembline) or from the Anaconda repository (https://anaconda.org/kosinskilab/assembline). Documentation is provided in the Supplementary Manual and can also be found online (https://assembline.readthedocs.io/en/latest/), and in the form of the step-by-step tutorials for modeling the yeast NPC (Supplementary Tutorial 1, online version: https://scnpc-tutorial.readthedocs.io/en/latest/) and Elongator complex (Supplementary Tutorial 2, online version: https://elongator-tutorial.readthedocs.io/en/latest/).
References
Marsh, J. A. & Teichmann, S. A. Structure, dynamics, assembly, and evolution of protein complexes. Ann. Rev. Biochem. 84, 551–575 (2015).
Patel, A. B. et al. Structure of human TFIID and mechanism of TBP loading onto promoter DNA. Science 362, eaau8872 (2018).
Vos, S. M., Farnung, L., Urlaub, H. & Cramer, P. Structure of paused transcription complex Pol II–DSIF–NELF. Nature 560, 601–606 (2018).
Lyumkis, D. Challenges and opportunities in cryo-EM single-particle analysis. J. Biol. Chem. 294, 5181–5197 (2019).
Beck, M. & Baumeister, W. Cryo-electron tomography: can it reveal the molecular sociology of cells in atomic detail? Trends Cell Biol. 26, 825–837 (2016).
Allegretti, M. et al. In-cell architecture of the nuclear pore and snapshots of its turnover. Nature 586, 796–800 (2020).
O’Reilly, F. J. et al. In-cell architecture of an actively transcribing-translating expressome. Science 369, 554–557 (2020).
Turk, M. & Baumeister, W. The promise and the challenges of cryo‐electron tomography. FEBS Lett. 594, 3243–3261 (2020).
Wagner, F. R. et al. Preparing samples from whole cells using focused-ion-beam milling for cryo-electron tomography. Nat. Protoc. 15, 2041–2070 (2020).
Bharat, T. A. M. & Scheres, S. H. W. Resolving macromolecular structures from electron cryo-tomography data using subtomogram averaging in RELION. Nat. Protoc. 11, 2054–2065 (2016).
Fäßler, F., Dimchev, G., Hodirnau, V. V., Wan, W. & Schur, F. K. M. Cryo-electron tomography structure of Arp2/3 complex in cells reveals new insights into the branch junction. Nat. Commun. 11, 6437 (2020).
Malhotra, S., Träger, S., Dal Peraro, M. & Topf, M. Modelling structures in cryo-EM maps. Curr. Opin. Struct. Biol. 58, 105–114 (2019).
Ferber, M. et al. Automated structure modeling of large protein assemblies using crosslinks as distance restraints. Nat. Methods 13, 515–520 (2016).
Orbán-Németh, Z. et al. Structural prediction of protein models using distance restraints derived from cross-linking mass spectrometry data. Nat. Protoc. 13, 478–494 (2018).
Gräwert, T. W. & Svergun, D. I. Structural modeling using solution small-angle X-ray scattering (SAXS). J. Mol. Biol. 432, 3078–3092 (2020).
Koukos, P. I. & Bonvin, A. M. J. J. Integrative modelling of biomolecular complexes. J. Mol. Biol. 432, 2861–2881 (2020).
Braitbard, M., Schneidman-Duhovny, D. & Kalisman, N. Integrative structure modeling: overview and assessment. Annu. Rev. Biochem. 88, 113–135 (2019).
Rout, M. P. & Sali, A. Principles for integrative structural biology studies. Cell 177, 1384–1403 (2019).
Viswanath, S., Chemmama, I. E., Cimermancic, P. & Sali, A. Assessing exhaustiveness of stochastic sampling for integrative modeling of macromolecular structures. Biophys. J. 113, 2344–2353 (2017).
Zimmerli, C. E. et al. Nuclear pores constrict upon energy depletion. Preprint at bioRxiv https://doi.org/10.1101/2020.07.30.228585 (2020).
Webb, B. et al. Integrative structure modeling with the Integrative Modeling Platform. Protein Sci. 27, 245–258 (2018).
Bordoli, L. et al. Protein structure homology modeling using SWISS-MODEL workspace. Nat. Protoc. 4, 1–13 (2009).
Kosinski, J. et al. Molecular architecture of the inner ring scaffold of the human nuclear pore complex. Science 352, 363–365 (2016).
Kosinski, J. et al. Xlink analyzer: software for analysis and visualization of cross-linking data in the context of three-dimensional structures. J. Struct. Biol. 189, 177–183 (2015).
Pettersen, E. F. et al. UCSF Chimera-A visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Russel, D. et al. Putting the pieces together: Integrative Modeling Platform software for structure determination of macromolecular assemblies. PLoS Biol. 10, e1001244 (2012).
Saltzberg, D. et al. Modeling biological complexes using Integrative Modeling Platform. Methods Mol. Biol. 2022, 353–377 (2019).
Dauden, M. I. et al. Architecture of the yeast Elongator complex. EMBO Rep. 18, 264–279 (2017).
Dauden, M. I. et al. Molecular basis of tRNA recognition by the Elongator complex. Sci. Adv. 5, eaaw2326 (2019).
Beckham, K. S. H. et al. Structure of the mycobacterial ESX-5 type VII secretion system pore complex. Sci. Adv. 7, eabg9923 (2021).
Bui, K. H. et al. Integrated structural analysis of the human nuclear pore complex scaffold. Cell 155, 1233–1243 (2013).
Fisher, R. A. Frequency distribution of the values of the correlation coefficient in samples from an indefinitely large population. Biometrika 10, 507 (1915).
Strimmer, K. fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics 24, 1461–1462 (2008).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Kirkpatrick, S., Gelatt, C. D. & Vecchi, M. P. Optimization by simulated annealing. Science 220, 671–680 (1983).
Schneidman-Duhovny, D., Hammel, M. & Sali, A. FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res. 38, W540–W544 (2010).
Lasker, K., Topf, M., Sali, A. & Wolfson, H. J. Inferential optimization for simultaneous fitting of multiple components into a CryoEM map of their assembly. J. Mol. Biol. 388, 180–194 (2009).
Dominguez, C., Boelens, R. & Bonvin, A. M. J. J. HADDOCK: a protein–protein docking approach based on biochemical or biophysical information. J. Am. Chem. Soc. 125, 1731–1737 (2003).
Karaca, E., Rodrigues, J. P. G. L. M., Graziadei, A., Bonvin, A. M. J. J. & Carlomagno, T. M3: an integrative framework for structure determination of molecular machines. Nat. Methods 14, 897–902 (2017).
Marrink, S. J., Risselada, H. J., Yefimov, S., Tieleman, D. P. & De Vries, A. H. The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 111, 7812–7824 (2007).
DiMaio, F., Tyka, M. D., Baker, M. L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using Rosetta. J. Mol. Biol. 392, 181–190 (2009).
Wriggers, W. Conventions and workflows for using Situs. Acta Crystallogr. D. Biol. Crystallogr. 68, 344–351 (2012).
Topf, M. et al. Protein structure fitting and refinement guided by cryo-EM density. Structure 16, 295–307 (2008).
Trabuco, L. G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Pandurangan, A. P., Vasishtan, D., Alber, F. & Topf, M. γ-TEMPy: simultaneous fitting of components in 3D-EM maps of their assembly using a genetic algorithm. Structure 23, 2365–2376 (2015).
Vitalis, A. & Caflisch, A. Equilibrium sampling approach to the interpretation of electron density maps. Structure 22, 156–167 (2014).
Lopéz-Blanco, J. R. & Chacón, P. IMODFIT: efficient and robust flexible fitting based on vibrational analysis in internal coordinates. J. Struct. Biol. 184, 261–270 (2013).
Ratje, A. H. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010).
Saha, M. & Morais, M. C. FOLD-EM: automated fold recognition in medium-and low-resolution (4–15 Å) electron density maps. Bioinformatics 28, 3265–3273 (2012).
de Vries, S. J. & Zacharias, M. ATTRACT-EM: a new method for the computational assembly of large molecular machines using cryo-EM maps. PLoS One 7, e49733 (2012).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Gil, V. A. & Guallar, V. pyRMSD: a Python package for efficient pairwise RMSD matrix calculation and handling. Bioinformatics 29, 2363–2364 (2013).
McInnes, L., Healy, J. & Astels, S. hdbscan: hierarchical density based clustering. J. Open Source Softw. 2, 205 (2017).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Beck, M. & Hurt, E. The nuclear pore complex: understanding its function through structural insight. Nat. Rev. Mol. Cell Biol. 18, 73–89 (2017).
Lin, D. H. et al. Architecture of the symmetric core of the nuclear pore. Science 352, aaf1015 (2016).
Drin, G. et al. A general amphipathic α-helical motif for sensing membrane curvature. Nat. Struct. Mol. Biol. 14, 138–146 (2007).
Kim, S. J. et al. Integrative structure and functional anatomy of a nuclear pore complex. Nature 555, 475–482 (2018).
Dauden, M. I., Jaciuk, M., Müller, C. W. & Glatt, S. Structural asymmetry in the eukaryotic Elongator complex. FEBS Lett. 592, 502–515 (2018).
Acknowledgements
We thank coauthors of the publications in which the Assembline protocol has been applied. We are grateful to D. Ziemianowicz and K. Kaszuba for comments on the manuscript and the protocol, and A. Obarska for feedback on the protocol. The work has been supported by the Federal Ministry of Education and Research of Germany (FKZ 031L0100)
Author information
Authors and Affiliations
Contributions
V.R., K.K. and J.K. developed the protocol. V.R and J.K wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks André Hoelz, Peter J. Peters and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Beckham, K. S. H. et al. Sci. Adv. 7, eabg9923 (2021): https://doi.org/10.1126/sciadv.abg9923
Allegretti, M. et al. Nature 586, 796–800 (2020): https://doi.org/10.1038/s41586-020-2670-5
Zimmerli, C. E. et al. Preprint at bioRxiv (2020): https://doi.org/10.1101/2020.07.30.228585
Dauden, M. I. et al. EMBO Rep. 18, 264–279 (2017): https://doi.org/10.15252/embr.201643353
Kosinski, J. et al. Science 352, 363–365 (2016): https://doi.org/10.1126/science.aaf0643
Essential data used in this protocol
Allegretti, M. et al. Nature 586, 796–800 (2020): https://doi.org/10.1038/s41586-020-2670-5
Dauden, M. I. et al. EMBO Rep. 18, 264–279 (2017): https://doi.org/10.15252/embr.201643353
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Supplementary Manual and Supplementary Tutorials 1 and 2.
Supplementary Video 1
CR Y-complex trajectory from global optimization. Animation video depicting the trajectory of the best scoring CR Y-complex model of the wild-type in-cell ScNPC6 produced with global optimization step. The model is shown in the coarse-grained representation inside the respective EM map.
Supplementary Video 2
NR Y-complex trajectory from refinement. Animation video depicting the trajectory of the best scoring NR Y-complex model of the in-cell wild-type ScNPC6 produced with refinement step. The model (and the symmetrical copies) is shown in the coarse-grained representation inside the respective EM map.
Rights and permissions
About this article
Cite this article
Rantos, V., Karius, K. & Kosinski, J. Integrative structural modeling of macromolecular complexes using Assembline. Nat Protoc 17, 152–176 (2022). https://doi.org/10.1038/s41596-021-00640-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00640-z
This article is cited by
-
The palisade layer of the poxvirus core is composed of flexible A10 trimers
Nature Structural & Molecular Biology (2024)
-
CombFold: predicting structures of large protein assemblies using a combinatorial assembly algorithm and AlphaFold2
Nature Methods (2024)
-
Integrated modeling of the Nexin-dynein regulatory complex reveals its regulatory mechanism
Nature Communications (2023)
-
Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.