Abstract
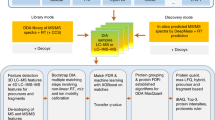
Targeted proteomics is a method of choice for accurate and high-throughput quantification of predefined sets of proteins. Many workflows use isotope-labeled reference peptides for every target protein, which is time consuming and costly. We report a statistical approach for quantifying full protein panels with a reduced set of reference peptides. This label-sparse approach achieves accurate quantification while reducing experimental cost and time. It is implemented in the software tool SparseQuant.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Picotti, P., Bodenmiller, B., Mueller, L.N., Domon, B. & Aebersold, R. Cell 138, 795–806 (2009).
Picotti, P. et al. Nat. Methods 7, 43–46 (2010).
Cima, I. et al. Proc. Natl. Acad. Sci. USA 108, 3342–3347 (2011).
Sabidó, E. et al. Mol. Cell. Proteomics 11, M111.014746 (2012).
Ferguson, R.E. et al. Proteomics 5, 566–571 (2005).
Greer, S., Honeywell, R., Geletu, M., Arulanandam, R. & Raptis, L. J. Immunol. Methods 355, 76–79 (2010).
Picotti, P. & Aebersold, R. Nat. Methods 9, 555–566 (2012).
Ong, S.-E. & Mann, M. Nat. Protoc. 1, 2650–2660 (2006).
Zhang, H. et al. Mol. Cell. Proteomics 10, M110.006593 (2011).
Sabidó, E. et al. Mol. Syst. Biol. 9, 681 (2013).
Geiger, T., Cox, J., Ostasiewicz, P., Wisniewski, J.R. & Mann, M. Nat. Methods 7, 383–385 (2010).
Gillet, L.C. et al. Mol. Cell. Proteomics 11, O111.016717 (2012).
Peterson, A.C., Russell, J.D., Bailey, D.J., Westphall, M.S. & Coon, J.J. Mol. Cell. Proteomics 11, O112.020131 (2012).
Chang, C.-Y. et al. Mol. Cell. Proteomics 11, M111.014662 (2012).
McCulloch, C.E. & Searle, S.R. Generalized, Linear, and Mixed Models (Wiley, 2001).
Montgomery, D. Design and Analysis of Experiments (Wiley, 2005).
Paulovich, A.G., Whiteaker, J.R., Hoofnagle, A.N. & Wang, P. Proteomics Clin. Appl. 2, 1386–1402 (2008).
Kim, J.-S. et al. Mol. Cell. Proteomics 10, M110.007302 (2011).
Addona, T.A. et al. Nat. Biotechnol. 27, 633–641 (2009).
Kersten, S. EMBO Rep. 2, 282–286 (2001).
Farrah, T. et al. Proteomics 12, 1170–1175 (2012).
Shannon, P. et al. Genome Res. 13, 2498–2504 (2003).
Szklarczyk, D. et al. Nucleic Acids Res. 39, D561–D568 (2011).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. Nucleic Acids Res. 32, D277–D280 (2004).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
The work was supported in part by the LiverX program of the Swiss Initiative for Systems Biology (SystemsX) to E.S., by the European Research Council (ERC) via grant “Proteomics v3.0” (grant 233226) and the Swiss National Science Foundation (grant 31-147086) to R.A., and by the US National Science Foundation CAREER award DBI-1054826 to O.V. We thank M. Choi for assistance with the manuscript.
Author information
Authors and Affiliations
Contributions
C.-Y.C. developed and implemented the statistical analysis framework, analyzed the data sets and wrote the manuscript. E.S. designed and conducted the mouse liver study and wrote the manuscript. R.A. supervised the experimental aspects of the work and wrote the manuscript. O.V. designed and supervised the statistical aspects of the work and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Data 1 and Supplementary Notes 1–3 (PDF 4597 kb)
Supplementary Data 2
Identified and quantified SRM transitions in the mouse liver study. These data are the input to the label-sparse quantification approach. (XLS 1793 kb)
Supplementary Data 3
The raw data in the Skyline format, generated by the initial screening experiment. These data are evidence that some transitions lack their reference counterpart, and require the label-sparse quantification approach. (ZIP 14258 kb)
Supplementary Data 4
The raw data in the Skyline format, generated by the actual experiment. These data can be used to assess the quality of the individual transitions. These transitions were identified and quantified, and the result is stored in Supplementary Data 2. (ZIP 17652 kb)
Supplementary Software
SparseQuant (ZIP 102 kb)
Source data
Rights and permissions
About this article
Cite this article
Chang, CY., Sabidó, E., Aebersold, R. et al. Targeted protein quantification using sparse reference labeling. Nat Methods 11, 301–304 (2014). https://doi.org/10.1038/nmeth.2806
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2806