Abstract
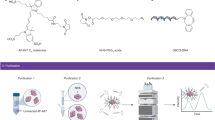
Precise control over interfacial chemistry between nanoparticles and other materials remains a major challenge that limits broad application of nanotechnology in biology. To address this challenge, we used 'steric exclusion' to completely convert commercial quantum dots (QDs) into monovalent imaging probes by wrapping each QD with a functionalized oligonucleotide. We demonstrated the utility of these QDs as modular and nonperturbing imaging probes by tracking individual Notch receptors on live cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
24 October 2013
In the version of this article initially published online, Figure 3c was mislabeled. The error has been corrected for the print, PDF and HTML versions of this article.
References
Pons, T., Medintz, I.L., Wang, X., English, D.S. & Mattoussi, H. J. Am. Chem. Soc. 128, 15324–15331 (2006).
Shmueli, G., Minka, T.P., Kadane, J.B., Borle, S. & Boatwright, P. J. R. Stat. Soc. Ser. C Appl. Stat. 54, 127–142 (2005).
Saxton, M.J. & Jacobson, K. Annu. Rev. Biophys. Biomol. Struct. 26, 373–399 (1997).
Banerjee, D., Liu, A.P., Voss, N.R., Schmid, S.L. & Finn, M.G. ChemBioChem 11, 1273–1279 (2010).
Howarth, M. et al. Nat. Methods 5, 397–399 (2008).
Zanchet, D., Micheel, C.M., Parak, W.J., Gerion, D. & Alivisatos, A.P. Nano Lett. 1, 32–35 (2001).
Clarke, S. et al. Nano Lett. 10, 2147–2154 (2010).
Carstairs, H.M.J., Lymperopoulos, K., Kapanidis, A.N., Bath, J. & Turberfield, A.J. ChemBioChem 10, 1781–1783 (2009).
You, C. et al. Angew. Chem. Int. Edn Engl. 49, 4108–4112 (2010).
Tikhomirov, G. et al. Nat. Nanotechnol. 6, 485–490 (2011).
You, C. et al. ACS Chem. Biol. 8, 320–326 (2013).
Wilson, R., Chen, Y. & Aveyard, J. Chem. Commun. (Camb.) 1156–1157 (2004).
Montenegro, J.-M. et al. Adv. Drug Deliv. Rev. 65, 677–688 (2013).
Jiang, L. et al. Chem. Phys. Lett. 380, 29–33 (2003).
Claridge, S.A., Liang, H.W., Basu, S.R., Fréchet, J.M.J. & Alivisatos, A.P. Nano Lett. 8, 1202–1206 (2008).
Bray, S.J. Nat. Rev. Mol. Cell Biol. 7, 678–689 (2006).
Gordon, W.R. et al. Blood 113, 4381–4390 (2009).
Douglass, A.D. & Vale, R.D. Cell 121, 937–950 (2005).
Triantafilou, M., Morath, S., Mackie, A., Hartung, T. & Triantafilou, K. J. Cell Sci. 117, 4007–4014 (2004).
Cairo, C.W. et al. J. Biol. Chem. 285, 11392–11401 (2010).
Jana, N.R., Gearheart, L. & Murphy, C.J. Langmuir 17, 6782–6786 (2001).
Grabolle, M. et al. Anal. Chem. 81, 6285–6294 (2009).
Selden, N.S. et al. J. Am. Chem. Soc. 134, 765–768 (2012).
Acknowledgements
We thank B. van Lengerich, L.D. Hughes and P. Haggie for assistance with single-particle-tracking software. S. Blacklow (Harvard University) provided human Notch1 constructs. R. Vale (University of California San Francisco) provided feedback on the single-molecule imaging. A.P. Alivisatos (University of California Berkeley) provided helpful discussion. J. Taunton and members of Cardiovascular Research Institute Core provided TIRF microscopes. N. Sturman helped with localization microscopy plugin. B. Liang, K. Southard and M. Todhunter generated some preliminary data. J.F. was supported by the University of California San Francisco Center for Synthetic and Systems Biology (P50 GM081879). D.S. was supported by Human Frontier Science Program Cross-disciplinary postdoc research fellowship. K.E.B. was supported by US National Institutes of Health National Research Service Award grant (5F32CA165620). Z.J.G. was supported by Kimmel Family Foundation. Y.J. was partly supported by 1R21EB015088-01 from the US National Institutes of Health NIBIB, and the Bryan Hemming Fellowship.
Author information
Authors and Affiliations
Contributions
Z.J.G., J.F. and Y.J. conceived the study; J.F., Z.J.G. and Y.J. designed experiments; J.F., D.S., K.E.B., M.J.T., Z.J.G. and Y.J. performed experiments; J.F. and D.S. analyzed and interpreted the data; and J.F., Z.J.G. and Y.J. wrote the manuscript. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–24, Supplementary Tables 1 and 2, and Supplementary Notes 1–4 (PDF 1580 kb)
Time lapse of SNAP proteins on supported lipid bilayers.
Timelapse TIRF movie taken of SNAP protein embeded in a supported lipid bilayer labeled with either Streptavidin QDots linked via BG-DNA-biotin, or mQDs. SNAP protein labeled with NHS-Atto488 shown to the right. Images taken at 20 Hz; scale bar is 10 μm. (AVI 19548 kb)
AF647- and mQD-labeled SNAP-Notch on a live U2OS cell.
Timelapse TIRF movie taken of SNAP.hN1 on a single cell. The cell's AF647 stained receptors were imaged for 25 seconds and immediately followed by imaging the mQD stained receptors. The AF647 & mQD regions are identical, separated by ~30 s in time. The white boxed region is shown in Figure 4c. Images taken at 20 Hz; scale bar is 1 μm. (AVI 11210 kb)
Rights and permissions
About this article
Cite this article
Farlow, J., Seo, D., Broaders, K. et al. Formation of targeted monovalent quantum dots by steric exclusion. Nat Methods 10, 1203–1205 (2013). https://doi.org/10.1038/nmeth.2682
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2682
This article is cited by
-
Nanoscale 3D spatial addressing and valence control of quantum dots using wireframe DNA origami
Nature Communications (2022)
-
Improved resolution in single-molecule localization microscopy using QD-PAINT
Experimental & Molecular Medicine (2021)
-
Enhanced Delta-Notch Lateral Inhibition Model Incorporating Intracellular Notch Heterogeneity and Tension-Dependent Rate of Delta-Notch Binding that Reproduces Sprouting Angiogenesis Patterns
Scientific Reports (2018)
-
Single-cell mRNA cytometry via sequence-specific nanoparticle clustering and trapping
Nature Chemistry (2018)
-
Single-cell mechanogenetics using monovalent magnetoplasmonic nanoparticles
Nature Protocols (2017)