Abstract
The T cell antigen receptor (TCR) is unique in that its affinity for ligand is unknown before encounter and can vary by orders of magnitude. How the immune system regulates individual T cells that display very different reactivity to antigen remains unclear. Here we found that activated CD4+ T cells, at the peak of clonal expansion, persistently downregulated their TCR expression in proportion to the strength of the initial antigen recognition. This programmed response increased the threshold for cytokine production and recall proliferation in a clone-specific manner and ultimately excluded clones with the highest antigen reactivity. Thus, programmed downregulation of TCR expression represents a negative feedback mechanism for constraining T cell effector function with a suitable time delay to thereby allow pathogen control while avoiding excess inflammatory damage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Nikolich-Zugich, J., Slifka, M.K. & Messaoudi, I. The many important facets of T-cell repertoire diversity. Nat. Rev. Immunol. 4, 123–132 (2004).
van Heijst, J.W. et al. Quantitative assessment of T cell repertoire recovery after hematopoietic stem cell transplantation. Nat. Med. 19, 372–377 (2013).
Busch, D.H. & Pamer, E.G. T cell affinity maturation by selective expansion during infection. J. Exp. Med. 189, 701–710 (1999).
Corse, E., Gottschalk, R.A. & Allison, J.P. Strength of TCR-peptide/MHC interactions and in vivo T cell responses. J. Immunol. 186, 5039–5045 (2011).
King, C.G. et al. T cell affinity regulates asymmetric division, effector cell differentiation, and tissue pathology. Immunity 37, 709–720 (2012).
Rees, W. et al. An inverse relationship between T cell receptor affinity and antigen dose during CD4+ T cell responses in vivo and in vitro. Proc. Natl. Acad. Sci. USA 96, 9781–9786 (1999).
Savage, P.A., Boniface, J.J. & Davis, M.M. A kinetic basis for T cell receptor repertoire selection during an immune response. Immunity 10, 485–492 (1999).
Varela-Rohena, A. et al. Control of HIV-1 immune escape by CD8 T cells expressing enhanced T-cell receptor. Nat. Med. 14, 1390–1395 (2008).
Zehn, D., Lee, S.Y. & Bevan, M.J. Complete but curtailed T-cell response to very low-affinity antigen. Nature 458, 211–214 (2009).
Corse, E., Gottschalk, R.A., Krogsgaard, M. & Allison, J.P. Attenuated T cell responses to a high-potency ligand in vivo. PLoS Biol. 8, e1000481 (2010).
Hebeisen, M. et al. SHP-1 phosphatase activity counteracts increased T cell receptor affinity. J. Clin. Invest. 123, 1044–1056 (2013).
Kalergis, A.M. et al. Efficient T cell activation requires an optimal dwell-time of interaction between the TCR and the pMHC complex. Nat. Immunol. 2, 229–234 (2001).
Viganò, S. et al. Functional avidity: a measure to predict the efficacy of effector T cells? Clin. Dev. Immunol. 2012, 153863 (2012).
O'Garra, A. et al. The immune response in tuberculosis. Annu. Rev. Immunol. 31, 475–527 (2013).
Ottenhoff, T.H. & Kaufmann, S.H. Vaccines against tuberculosis: where are we and where do we need to go? PLoS Pathog. 8, e1002607 (2012).
Lawn, S.D., Myer, L., Edwards, D., Bekker, L.G. & Wood, R. Short-term and long-term risk of tuberculosis associated with CD4 cell recovery during antiretroviral therapy in South Africa. AIDS 23, 1717–1725 (2009).
Mogues, T., Goodrich, M.E., Ryan, L., LaCourse, R. & North, R.J. The relative importance of T cell subsets in immunity and immunopathology of airborne Mycobacterium tuberculosis infection in mice. J. Exp. Med. 193, 271–280 (2001).
Hmama, Z., Gabathuler, R., Jefferies, W.A., de Jong, G. & Reiner, N.E. Attenuation of HLA-DR expression by mononuclear phagocytes infected with Mycobacterium tuberculosis is related to intracellular sequestration of immature class II heterodimers. J. Immunol. 161, 4882–4893 (1998).
Pai, R.K., Convery, M., Hamilton, T.A., Boom, W.H. & Harding, C.V. Inhibition of IFN-gamma-induced class II transactivator expression by a 19-kDa lipoprotein from Mycobacterium tuberculosis: a potential mechanism for immune evasion. J. Immunol. 171, 175–184 (2003).
Bold, T.D., Banaei, N., Wolf, A.J. & Ernst, J.D. Suboptimal activation of antigen-specific CD4+ effector cells enables persistence of M. tuberculosis in vivo. PLoS Pathog. 7, e1002063 (2011).
Egen, J.G. et al. Intravital imaging reveals limited antigen presentation and T cell effector function in mycobacterial granulomas. Immunity 34, 807–819 (2011).
Gallegos, A.M., Pamer, E.G. & Glickman, M.S. Delayed protection by ESAT-6-specific effector CD4+ T cells after airborne M. tuberculosis infection. J. Exp. Med. 205, 2359–2368 (2008).
Gallegos, A.M. et al. A γ interferon independent mechanism of CD4 T cell mediated control of M. tuberculosis infection in vivo. PLoS Pathog. 7, e1002052 (2011).
Crawford, F., Kozono, H., White, J., Marrack, P. & Kappler, J. Detection of antigen-specific T cells with multivalent soluble class II MHC covalent peptide complexes. Immunity 8, 675–682 (1998).
Egen, J.G. & Allison, J.P. Cytotoxic T lymphocyte antigen-4 accumulation in the immunological synapse is regulated by TCR signal strength. Immunity 16, 23–35 (2002).
Winslow, G.M., Roberts, A.D., Blackman, M.A. & Woodland, D.L. Persistence and turnover of antigen-specific CD4 T cells during chronic tuberculosis infection in the mouse. J. Immunol. 170, 2046–2052 (2003).
Badovinac, V.P., Porter, B.B. & Harty, J.T. Programmed contraction of CD8+ T cells after infection. Nat. Immunol. 3, 619–626 (2002).
Kaech, S.M. & Ahmed, R. Memory CD8+ T cell differentiation: initial antigen encounter triggers a developmental program in naive cells. Nat. Immunol. 2, 415–422 (2001).
Lee, W.T., Pasos, G., Cecchini, L. & Mittler, J.N. Continued antigen stimulation is not required during CD4+ T cell clonal expansion. J. Immunol. 168, 1682–1689 (2002).
Mercado, R. et al. Early programming of T cell populations responding to bacterial infection. J. Immunol. 165, 6833–6839 (2000).
van Stipdonk, M.J., Lemmens, E.E. & Schoenberger, S.P. Naive CTLs require a single brief period of antigenic stimulation for clonal expansion and differentiation. Nat. Immunol. 2, 423–429 (2001).
Lauer, P., Chow, M.Y., Loessner, M.J., Portnoy, D.A. & Calendar, R. Construction, characterization, and use of two Listeria monocytogenes site-specific phage integration vectors. J. Bacteriol. 184, 4177–4186 (2002).
Sussman, J.J. et al. Failure to synthesize the T cell CD3-ζ chain: structure and function of a partial T cell receptor complex. Cell 52, 85–95 (1988).
Cenciarelli, C. et al. Activation-induced ubiquitination of the T cell antigen receptor. Science 257, 795–797 (1992).
Naramura, M. et al. c-Cbl and Cbl-b regulate T cell responsiveness by promoting ligand-induced TCR down-modulation. Nat. Immunol. 3, 1192–1199 (2002).
Valitutti, S., Müller, S., Salio, M. & Lanzavecchia, A. Degradation of T cell receptor (TCR)-CD3-ζ complexes after antigenic stimulation. J. Exp. Med. 185, 1859–1864 (1997).
Labrecque, N. et al. How much TCR does a T cell need? Immunity 15, 71–82 (2001).
Schrum, A.G., Turka, L.A. & Palmer, E. Surface T-cell antigen receptor expression and availability for long-term antigenic signaling. Immunol. Rev. 196, 7–24 (2003).
Viola, A. & Lanzavecchia, A. T cell activation determined by T cell receptor number and tunable thresholds. Science 273, 104–106 (1996).
Itoh, Y., Hemmer, B., Martin, R. & Germain, R.N. Serial TCR engagement and down-modulation by peptide:MHC molecule ligands: relationship to the quality of individual TCR signaling events. J. Immunol. 162, 2073–2080 (1999).
Valitutti, S., Müller, S., Cella, M., Padovan, E. & Lanzavecchia, A. Serial triggering of many T-cell receptors by a few peptide-MHC complexes. Nature 375, 148–151 (1995).
Germain, R.N. Maintaining system homeostasis: the third law of Newtonian immunology. Nat. Immunol. 13, 902–906 (2012).
Baniyash, M. TCR zeta-chain downregulation: curtailing an excessive inflammatory immune response. Nat. Rev. Immunol. 4, 675–687 (2004).
Anderton, S.M. & Wraith, D.C. Selection and fine-tuning of the autoimmune T-cell repertoire. Nat. Rev. Immunol. 2, 487–498 (2002).
Weber, K.S. et al. Distinct CD4+ helper T cells involved in primary and secondary responses to infection. Proc. Natl. Acad. Sci. USA 109, 9511–9516 (2012).
Mandl, J.N., Monteiro, J.P., Vrisekoop, N. & Germain, R.N. T cell-positive selection uses self-ligand binding strength to optimize repertoire recognition of foreign antigens. Immunity 38, 263–274 (2013).
Persaud, S.P., Parker, C.R., Lo, W.L., Weber, K.S. & Allen, P.M. Intrinsic CD4+ T cell sensitivity and response to a pathogen are set and sustained by avidity for thymic and peripheral complexes of self peptide and MHC. Nat. Immunol. 15, 266–274 (2014).
Fassò, M. et al. T cell receptor (TCR)-mediated repertoire selection and loss of TCR vbeta diversity during the initiation of a CD4+ T cell response in vivo. J. Exp. Med. 192, 1719–1730 (2000).
McCue, D., Ryan, K.R., Wraith, D.C. & Anderton, S.M. Activation thresholds determine susceptibility to peptide-induced tolerance in a heterogeneous myelin-reactive T cell repertoire. J. Neuroimmunol. 156, 96–106 (2004).
McMahan, R.H. et al. Relating TCR-peptide-MHC affinity to immunogenicity for the design of tumor vaccines. J. Clin. Invest. 116, 2543–2551 (2006).
Acknowledgements
We thank M. Samstein, H. Yan and S. Reddy for technical assistance. Supported by the US National Institutes of Health (F32 grant AI074248 to A.M.G.; AI080619 to M.S.G. and E.G.P.; and P30 CA008748) and the Netherlands Organisation for Scientific Research (Veni grant 91614038 to J.W.J.v.H.).
Author information
Authors and Affiliations
Contributions
A.M.G., M.S.G., E.G.P. and J.W.J.v.H. designed the study; A.M.G., H.X. and J.W.J.v.H. performed experiments and analyzed data; A.M.G. generated the C7 and C24 mice and the L. monocytogenes–ESAT6 strain; I.M.L. and B.S. bred and genotyped the C7 and C24 mice and provided technical assistance; J.W.J.v.H. first identified programmed TCR downregulation; and E.G.P. and J.W.J.v.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 C24 T cells display greater upregulation of CTLA-4 expression during activation than do C7 T cells.
Flow cytometry showing CTLA-4 expression of C7 and C24 TH1 cells, during in vitro activation for 4 days. CD4 expression is shown as a control. Data are from one experiment representative of two independent experiments.
Supplementary Figure 2 Naive C24 T cells display lower expression of CD5 but greater basal phosphorylation of TCRζ than that of naive C7 T cells.
(a) Flow cytometry showing blood CD5 expression of C7 and C24 CD4+ T cells, as well as B6 CD4+ T cells. (b) Immunoblot showing basal TCRζ phosphorylation in whole-cell lysates of naive C7 and C24 T cells. Numbers below lanes indicate TCRζ pTyr142 density normalized to β-actin and presented relative to C7 T cells. Data are from one experiment representative of two independent experiments.
Supplementary Figure 3 Programmed TCR downregulation does not require TCR stimulation beyond initial T cell activation.
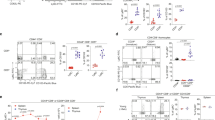
(a) Flow cytometry showing blood frequency of wild-type (WT) and MHC class II-deficient (MHC II−/–) mice that received 106 CD90.1+ C7 or C24 TH1 cells, which had been activated in vitro for 4 days. Blood samples were analyzed on day 7 post T cell activation. Numbers depict the percentage of gated cells. Note the clear absence of endogenous CD4+ T cells in MHC II−/– recipients. (b,c) Flow cytometry showing TCRβ expression kinetics of blood C7 and C24 TH1 cells from the recipient mice described in a. Individual flow plots are shown in b and aggregate data are shown in c. Endogenous CD4+ T cells were identified as CD90.1− CD4+ T cells in WT recipients. Data are from one experiment representative of two independent experiments (mean + s.d. of n = 3 male mice per group (c)).
Supplementary Figure 4 Surface marker and Foxp3 expression of activated C7 and C24 T cells undergoing programmed TCR downregulation.
(a) Flow cytometry showing splenocyte surface marker expression of mice that received 104 naive CD90.1+ C7 or C24 CD4+ T cells, and were infected with recombinant L. monocytogenes-ESAT6. Cells were analyzed either before TCR downregulation (day 6) or after TCR downregulation (day 9). A representative flow plot for each surface marker is shown. Endogenous CD4+ T cells were identified as CD90.1− CD4+ T cells. (b) Flow cytometry showing Foxp3 and CD25 expression of splenic C7 and C24 CD4+ T cells, as well as endogenous CD4+ T cells, obtained on day 9 from the recipient mice described in a. Numbers depict the percentage of gated cells. Data are from one experiment representative of two independent experiments.
Supplementary Figure 5 Programmed TCR downregulation of naive C7, C24 and endogenous CD4+ T cells activated by M. tuberculosis infection.
(a) Blood frequency of mice that received 104 naive CD90.1+ C7 or C24 CD4+ T cells, or did not receive any cells (no transfer), and were infected with M. tuberculosis. Endogenous activated CD4+ T cells were defined as CD44hi CD62L− CD4+ T cells in ‘no transfer’ recipients. (b,c) Flow cytometry showing TCRβ expression kinetics of blood C7 and C24 CD4+ T cells, as well as endogenous activated and naive CD4+ T cells from the recipient mice described in a. Individual flow plots are shown in b and aggregate data are shown in c. Endogenous naive CD4+ T cells were defined as CD44lo CD62L+ CD4+ T cells in ‘no transfer’ recipients. (d) Flow cytometry showing ESAT6(1–20) tetramer binding and TCRβ expression of endogenous lung CD4+ T cells, 28 days after M. tuberculosis infection. TCRβ expression of the 10% lowest and 10% highest tetramer binding cells is shown. (e) Pearson correlation of ESAT6(1–20) tetramer binding and TCRβ expression (r = 0.89; P < 0.001). Shown are the 10% lowest and 10% highest tetramer binding cells of each mouse. Data are from one experiment representative of two independent experiments (mean + s.d. of n = 4 female mice per group (a,c) or mean of n = 5 male mice per group (e)).
Supplementary Figure 6 Programmed TCR downregulation is not mediated by a transcriptional change in CD3 components or the TCRζ chain.
(a,b) Flow cytometry showing splenocyte TCRβ expression of mice that received 104 naive CD90.1+ C7 or C24 CD4+ T cells, and were infected with recombinant L. monocytogenes-ESAT6. On day 9 post infection, CD90.1+ C7 and C24 T cells were purified from spleens by magnetic separation. Individual flow plot is shown in a and aggregate data are shown in b. (c) Real-time PCR showing relative gene expression of splenic C7 and C24 T cells, obtained from the recipient mice described in a. Signals were corrected using Actb as endogenous control. Relative mRNA expression of the indicated genes was determined using the comparative CT method and normalized to the signal obtained for C7 T cells. Data are from one experiment representative of two independent experiments (mean + s.d. of n = 3 male mice per group (b,c)).
Supplementary Figure 7 Programmed TCR downregulation is a shared feature of clonally expanded CD4+ T cells and CD8+ T cells.
(a) Flow cytometry showing blood frequency of mice that were infected with recombinant L. monocytogenes-ESAT6 on day 0 and reinfected with the same bacteria on day 28. Blood samples were analyzed on day 31 post infection. Gating on CD4+ T cells revealed the presence of two subpopulations, with cells expressing either an activated (CD44hiCD62L−) or a naive (CD44loCD62L+) phenotype. Similar to CD4+ T cells, gating on CD8+ T cells revealed the presence of both activated and naive T cells, in addition to a third subpopulation that was CD44hiCD62L+, consistent with a central memory phenotype. Numbers depict the percentage of gated cells. (b) Blood frequency of endogenous activated CD4+ and CD8+ T cells from the mice described in a. (c,d) Flow cytometry showing TCRβ expression kinetics of endogenous activated CD4+ and CD8+ T cells from the mice described in a. Individual flow plots are shown in c and aggregate data are shown in d. Data are from one experiment representative of two independent experiments (mean + s.d. of n = 5 male mice per group (b,d)).
Supplementary Figure 8 Programmed TCR downregulation is relatively insensitive to the dose of activating peptide.
(a) Flow cytometry showing CFSE dilution of C7 and C24 CD4+ T cells activated with either 5,000 ng mL−1 or 50 ng mL−1 ESAT6(1–20) peptide for 3 days. (b) Blood frequency of mice that received 106 CD90.1+ C24 TH1 cells, which had been activated in vitro with either 5,000 ng mL−1 or 50 ng mL−1 ESAT6(1–20) peptide for 3 days. (c,d) Flow cytometry showing TCRβ expression kinetics of blood C24 TH1 cells from the recipient mice described in b. Individual flow plots are shown in c and aggregate data are shown in d. Endogenous CD4+ T cells were identified as CD90.1− CD4+ T cells. Data are from one experiment representative of two independent experiments (mean + s.d. of n = 5 female mice per group (b,d)).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1090 kb)
Supplementary Table 1
Table showing validation of antibodies used in the study (XLS 27 kb)
Rights and permissions
About this article
Cite this article
Gallegos, A., Xiong, H., Leiner, I. et al. Control of T cell antigen reactivity via programmed TCR downregulation. Nat Immunol 17, 379–386 (2016). https://doi.org/10.1038/ni.3386
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3386
This article is cited by
-
T cell receptor therapeutics: immunological targeting of the intracellular cancer proteome
Nature Reviews Drug Discovery (2023)
-
Immune evasion and provocation by Mycobacterium tuberculosis
Nature Reviews Microbiology (2022)
-
TIM-3 drives temporal differences in restimulation-induced cell death sensitivity in effector CD8+ T cells in conjunction with CEACAM1
Cell Death & Disease (2021)
-
ADAM12 is a costimulatory molecule that determines Th1 cell fate and mediates tissue inflammation
Cellular & Molecular Immunology (2021)
-
Proximal and distal effects of genetic susceptibility to multiple sclerosis on the T cell epigenome
Nature Communications (2021)