Abstract
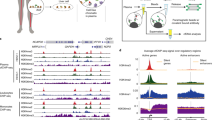
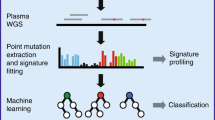
The analysis of cell-free DNA (cfDNA) in plasma represents a rapidly advancing field in medicine. cfDNA consists predominantly of nucleosome-protected DNA shed into the bloodstream by cells undergoing apoptosis. We performed whole-genome sequencing of plasma DNA and identified two discrete regions at transcription start sites (TSSs) where nucleosome occupancy results in different read depth coverage patterns for expressed and silent genes. By employing machine learning for gene classification, we found that the plasma DNA read depth patterns from healthy donors reflected the expression signature of hematopoietic cells. In patients with cancer having metastatic disease, we were able to classify expressed cancer driver genes in regions with somatic copy number gains with high accuracy. We were able to determine the expressed isoform of genes with several TSSs, as confirmed by RNA-seq analysis of the matching primary tumor. Our analyses provide functional information about cells releasing their DNA into the circulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
NCBI Reference Sequence
Sequence Read Archive
References
Schwarzenbach, H., Hoon, D.S. & Pantel, K. Cell-free nucleic acids as biomarkers in cancer patients. Nat. Rev. Cancer 11, 426–437 (2011).
Heitzer, E., Auer, M., Ulz, P., Geigl, J.B. & Speicher, M.R. Circulating tumor cells and DNA as liquid biopsies. Genome Med. 5, 73 (2013).
Crowley, E., Di Nicolantonio, F., Loupakis, F. & Bardelli, A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 10, 472–484 (2013).
Diaz, L.A. Jr. & Bardelli, A. Liquid biopsies: genotyping circulating tumor DNA. J. Clin. Oncol. 32, 579–586 (2014).
Heitzer, E., Ulz, P. & Geigl, J.B. Circulating tumor DNA as a liquid biopsy for cancer. Clin. Chem. 61, 112–123 (2015).
Diehl, F. et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc. Natl. Acad. Sci. USA 102, 16368–16373 (2005).
Lo, Y.M. et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci. Transl. Med. 2, 61ra91 (2010).
Ramachandran, S. & Henikoff, S. Replicating nucleosomes. Sci. Adv. 1, e1500587 (2015).
Snyder, M.W., Kircher, M., Hill, A.J., Daza, R.M. & Shendure, J. Cell-free DNA comprises an in vivo nucleosome footprint that informs its tissues-of-origin. Cell 164, 57–68 (2016).
Gaffney, D.J. et al. Controls of nucleosome positioning in the human genome. PLoS Genet. 8, e1003036 (2012).
Schones, D.E. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008).
Venkatesh, S. & Workman, J.L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 16, 178–189 (2015).
Valouev, A. et al. Determinants of nucleosome organization in primary human cells. Nature 474, 516–520 (2011).
Chandrananda, D., Thorne, N.P. & Bahlo, M. High-resolution characterization of sequence signatures due to non-random cleavage of cell-free DNA. BMC Med. Genomics 8, 29 (2015).
Eisenberg, E. & Levanon, E.Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Lui, Y.Y. et al. Predominant hematopoietic origin of cell-free DNA in plasma and serum after sex-mismatched bone marrow transplantation. Clin. Chem. 48, 421–427 (2002).
Sun, K. et al. Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments. Proc. Natl. Acad. Sci. USA 112, E5503–E5512 (2015).
Koh, W. et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl. Acad. Sci. USA 111, 7361–7366 (2014).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Heitzer, E., Ulz, P., Geigl, J.B. & Speicher, M.R. Non-invasive detection of genome-wide somatic copy number alterations by liquid biopsies. Mol. Oncol. 10, 494–502 (2016).
Heidary, M. et al. The dynamic range of circulating tumor DNA in metastatic breast cancer. Breast Cancer Res. 16, 421 (2014).
Heitzer, E. et al. Tumor-associated copy number changes in the circulation of patients with prostate cancer identified through whole-genome sequencing. Genome Med. 5, 30 (2013).
Mohan, S. et al. Changes in colorectal carcinoma genomes under anti-EGFR therapy identified by whole-genome plasma DNA sequencing. PLoS Genet. 10, e1004271 (2014).
Carter, S.L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Murtaza, M. et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature 497, 108–112 (2013).
Ulz, P., Heitzer, E. & Speicher, M.R. Co-occurrence of MYC amplification and TP53 mutations in human cancer. Nat. Genet. 48, 104–106 (2016).
Giordano, S.H. et al. Systemic therapy for patients with advanced human epidermal growth factor receptor 2–positive breast cancer: American Society of Clinical Oncology clinical practice guideline. J. Clin. Oncol. 32, 2078–2099 (2014).
Helsten, T. et al. The FGFR landscape in cancer: analysis of 4,853 tumors by next-generation sequencing. Clin. Cancer Res. 22, 259–267 (2016).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra24 (2014).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Ulz, P. et al. Whole-genome plasma sequencing reveals focal amplifications as a driving force in metastatic prostate cancer. Nat. Commun. 7, 12008 (2016).
Adelman, K. & Lis, J.T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Ivanov, M., Baranova, A., Butler, T., Spellman, P. & Mileyko, V. Non-random fragmentation patterns in circulating cell-free DNA reflect epigenetic regulation. BMC Genomics 16 (Suppl. 13), S1 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Lai, W., Choudhary, V. & Park, P.J. CGHweb: a tool for comparing DNA copy number segmentations from multiple algorithms. Bioinformatics 24, 1014–1015 (2008).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Acknowledgements
We are grateful to S. Perakis for critical reading and editing of this manuscript. This work was supported by CANCER-ID, a project funded by the Innovative Medicines Joint Undertaking (IMI JU).
Author information
Authors and Affiliations
Contributions
P.U. and M.R.S. designed the study. M.A. and R.G. performed the experiments. P.U., G.G.T., J.B.G., E.H., and M.R.S. analyzed data. E.P. and G.P. provided clinical samples and clinical information. S.W.J. and L.A. performed pathology analyses. K.K. conducted RNA-seq. PU., E.H., and M.R.S. supervised the study. P.U., J.B.G., E.H., and M.R.S. wrote the manuscript. All authors revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Mapping of the nucleosome-depleted region.
Localization of the NDR, which was mapped by analyses of 100 (red) and 1,000 (orange) highly expressed genes in the 104 plasma samples from healthy donors and which was most often observed in a –150 bp to +50 bp window with respect to the TSS (blue, 1,000 most weakly expressed genes).
Supplementary Figure 2 Classification of the 5,000 most highly and least expressed genes.
Support vector machine (SVM) classification based on normalized 2K-TSS and NDR coverage for the 5,000 most highly and least expressed genes. Red and dark blue circles represent genes correctly assigned to the expressed and unexpressed clusters, respectively, whereas light blue and orange circles represent incorrectly assigned genes (as in Fig. 3b).
Supplementary Figure 3 Quantitative relationship between nucleosome occupancy and gene expression.
(a) Correlation between 2K-TSS (left) and NDR (right) coverage and FPKM percentiles. (b) Means and distribution of the 2K-TSS and NDR coverage parameters of genes grouped into deciles. (c) Average FPKM percentile of binned 2K-TSS and NDR coverage parameters.
Supplementary Figure 4 Comparison of copy number profiles of the matching primary tumor with plasma DNA.
The copy number profiles of the matching primary tumors B7 (top) and B13 (bottom) were obtained by whole-genome sequencing with a shallow sequencing depth. Pairwise comparisons of genomic position–mapped profiles revealed high correlations between the copy number profiles (Pearson correlation coefficients = 0.74 (B7) and 0.88 (B13)).
Supplementary Figure 5 Reconstruction of the 12p11.1 nucleosome array with high-coverage sequenced plasma samples.
Assembly of the 12p11.1 nucleosome arrays in plasma samples from B7, B13, controls, and GM12878 for comparison.
Supplementary Figure 6 TSS nucleosome occupancy of unexpressed and housekeeping genes in high-coverage sequenced plasma samples in B7 and B13.
(a,b) Nucleosome occupancy at TSSs of unexpressed genes (fantom.gsc.riken.jp/5/) and housekeeping genes2 had the expected different pattern for B7 (a) and B13 (b).
Supplementary Figure 7 Distribution of the prediction consent.
Histogram of prediction consent in merged control (n=104) data. For the majority of genes, the prediction consent was above 95%; there are only a few genes with a prediction consent below 75%.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note. (PDF 1610 kb)
Supplementary Table 1
List of the 100 most highly expressed genes based on plasma RNA-seq data provided by Koh et al. but predicted to be unexpressed (n = 12). (XLSX 11 kb)
Supplementary Table 2
List of the 1,000 most highly expressed genes based on plasma RNA-seq data provided by Koh et al. but predicted to be unexpressed (n = 245). (XLSX 18 kb)
Supplementary Table 3
Subsampling of sequencing data to establish a lower boundary of necessary sequencing coverage. (XLSX 10 kb)
Supplementary Table 4
Prediction and FPKM values for genes in focal amplifications having log2 ratio > 1 in breast cancer case B7. (XLSX 19 kb)
Supplementary Table 5
Prediction and FPKM values for genes in focal amplifications having log2 ratio > 1 in breast cancer case B13. (XLSX 25 kb)
Rights and permissions
About this article
Cite this article
Ulz, P., Thallinger, G., Auer, M. et al. Inferring expressed genes by whole-genome sequencing of plasma DNA. Nat Genet 48, 1273–1278 (2016). https://doi.org/10.1038/ng.3648
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3648
This article is cited by
-
Multimodal epigenetic sequencing analysis (MESA) of cell-free DNA for non-invasive colorectal cancer detection
Genome Medicine (2024)
-
A multiparameter liquid biopsy approach allows to track melanoma dynamics and identify early treatment resistance
npj Precision Oncology (2024)
-
The landscape of cell-free mitochondrial DNA in liquid biopsy for cancer detection
Genome Biology (2023)
-
Multidimensional fragmentomic profiling of cell-free DNA released from patient-derived organoids
Human Genomics (2023)
-
Circulating tumor nucleic acids: biology, release mechanisms, and clinical relevance
Molecular Cancer (2023)