Abstract
Three strikingly different alternative male mating morphs (aggressive 'independents', semicooperative 'satellites' and female-mimic 'faeders') coexist as a balanced polymorphism in the ruff, Philomachus pugnax, a lek-breeding wading bird1,2,3. Major differences in body size, ornamentation, and aggressive and mating behaviors are inherited as an autosomal polymorphism4,5. We show that development into satellites and faeders is determined by a supergene6,7,8 consisting of divergent alternative, dominant and non-recombining haplotypes of an inversion on chromosome 11, which contains 125 predicted genes. Independents are homozygous for the ancestral sequence. One breakpoint of the inversion disrupts the essential CENP-N gene (encoding centromere protein N), and pedigree analysis confirms the lethality of homozygosity for the inversion. We describe new differences in behavior, testis size and steroid metabolism among morphs and identify polymorphic genes within the inversion that are likely to contribute to the differences among morphs in reproductive traits.
Similar content being viewed by others
Main
Stable genetic polymorphisms for alternative reproductive strategies, involving differences in body size, structure and behavior, have been identified in a range of taxa9,10. These traits are clearly associated with fitness and are putatively maintained by frequency-dependent selection. However, little is known about their genomic basis or mode of evolution or why examples of such strategies are relatively rare in contrast to the widespread phenotypic plasticity in reproductive strategy in response to environmental variation10.
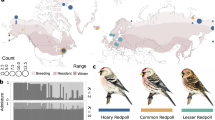
The ruff (P. pugnax) is a Eurasian sandpiper with a highly polygynous, lek-based mating system involving competition and cooperation among three male morphs (Fig. 1) and extensive female mate choice and genetic polyandry1,2,11,12. Territorial independent males, with hypervariable but predominantly dark ornamental plumage, defend small mating courts on leks. Non-territorial satellite males, with predominantly white ornamental plumage, join independents on courts, co-displaying to attract females but competing for matings. Rare small, female-mimicking faeder males attend the courts of independent males and attempt rapid copulations when females solicit matings from ornamented displaying males3.
(a) Representative breeding plumages. (b) Body size distributions5 (Online Methods). PC1, principal component 1. (c) Comparisons of morph behavioral profiles for aggression (gray), display (red), proximity (orange) and alert stance (green). Morph-specific box plots are shown for each behavior from the means for individual male ruffs, standardized across the sample population by subtracting the global mean and dividing by standard deviation. Morphs differed in their distributions of behavior (MANOVA on individual-bird mean values; multivariate heterogeneity among morphs Wilks's λ = 0.33, approximate F6,44 = 5.37, P < 0.0003). (d) Residuals of the logarithm of the testis volume index corrected for date differ among morphs (F2,37 = 6.31, P = 0.0045). Testes of independents (n = 29) are substantially smaller than those of satellites (n = 4; Tukey adjusted P = 0.006) and faeders (n = 6; Tukey adjusted P = 0.02). Box plots indicate the median (bold line), 25% and 75% quartiles (box), and range (whiskers). (e,f) Seasonal patterns of circulating steroid concentrations in the blood plasma of male ruffs, including for androstenedione (A4) (e) and testosterone plus dihydrotestosterone (T + DHT) (f). Points are daily means ± s.d. Data are shown for independents (n = 11; red, jittered by −1 d), satellites (n = 9; blue) and faeders (n = 2; black, jittered by +1 d). Independents have higher date-specific levels of testosterone than satellites (F1,97 = 10.01, P = 0.002), and satellites have higher levels of androstenedione than independents (F1,97 = 17.03, P < 0.0001; Supplementary Table 1); faeders appear similar to satellites but were not tested statistically owing to a sample size of n = 2.
Two autosomal Mendelian factors, Faeder and Satellite, have been shown to be dominant to independent, but the precise genetic architecture of morph determination remains unclear4,5. We have maintained a pedigreed captive breeding population of several hundred ruffs since 1985. Adult males were phenotyped for mating behavior (independent, satellite or faeder) and the presence of ornamental plumage (ornamented versus faeder). Some adult females were also phenotyped, with small size indicating carriers of Faeder5 and aggressive behavior following testosterone implantation permitting classification as independents or satellites13.
We explored the reproductive physiology of the morphs, with a particular focus on steroid metabolites, known to affect these phenotypes13. The testes of satellites, like those of faeders3, were larger than those of independents, despite their smaller body sizes (Fig. 1b and Supplementary Fig. 1), presumably to increase the efficiency of less frequent or effective copulations by producing larger numbers of sperm. Breeding independents had higher circulating testosterone concentrations, whereas satellites and faeders had higher concentrations of androstenedione (Fig. 1e,f and Supplementary Table 1), suggesting fundamental differences among the morphs in the regulation of the hypothalamus-pituitary-gonad system that drives seasonal reproduction14,15.
To identify and characterize the loci controlling morph divergence, we used a combination of genetics and genomics, on the basis of both our captive population and wild ruffs. We generated a reference ruff genome from a single independent male in the captive colony, using Illumina paired-end 3-kb and 5-kb mate-pair data and long-read Pacific Biosciences (PacBio) data (Supplementary Fig. 2 and Supplementary Table 2). The assembled genome has a span of ∼1.17 Gb in 12,085 scaffolds longer than 1 kb, with 292 scaffolds longer than 877 kb (Supplementary Table 3). Genes were predicted using extensive new transcriptome data (Supplementary Table 4). Assessments of assembly quality using genomic read mapping, transcriptome read and assembly mapping, and comparison to the high-quality chicken (Gallus gallus) genome assembly suggest high completeness and contiguity (Supplementary Fig. 3 and Supplementary Table 5). The draft genome was ordered and orientated using the chicken reference to yield a chromosome-level assembly. We identified and typed SNPs in the pedigree population using reduced-representation restriction site–associated DNA (RAD) sequencing16 at low density (on the basis of SbfI restriction sites), identifying 1,068,556 SNPs. We mapped Faeder and Satellite to a genetic map based on 3,948 biallelic SNPs with minor allele frequency ≥ 0.1, which had been typed in 286 individuals with a success rate ≥ 0.99.
Faeder and Satellite each mapped significantly and uniquely (logarithm of odds (LOD) >3) to the same region of chromosome 11 (Fig. 2 and Supplementary Fig. 4), which is coincident with the region previously identified for Faeder using microsatellite markers17. The region of linkage encompasses about one-fifth of the chromosome. Independent estimation of genetic maps for cohorts containing or excluding birds carrying Satellite or Faeder gave very different recombination lengths for this region (Fig. 2b), suggesting that it might contain a genomic rearrangement refractory to recombination between haplotypes.
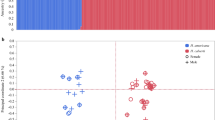
(a) The ruff linkage-mapping pedigree. Paternal links are shown as red lines, and maternal links are shown as blue lines. (b) Linkage maps (in cM) of the inversion region on ruff chromosome 11 support the presence of the inversion polymorphism, indicated by a ∼70% shorter linkage map when only independents are considered. SNPs on the same contig are shown in the same color. (c) Association between markers and morphs based on 41 unrelated males (ten faeder, ten satellite and 21 independent). Alternating shades of gray indicate different contigs, ordered on the basis of synteny with the chicken genome. The top panel shows −log10 (P values) of association across the entire ruff genome; the middle (Faeder) and bottom (Satellite) panels show enlarged views of the associated peak (0.703–0.713 Gb; indicated by dashed lines in the top panel) for comparisons with the independent locus. Red and blue dots represent genome-wide significant signals (P < 0.05, 1,000 permutations) for the Faeder and Satellite loci, respectively.
A separate genome-wide association study (GWAS), using densely sampled SNPs (from RAD sequencing at PstI restriction sites) in a sample of 41 unrelated faeder, satellite and independent birds, identified genome-wide significance of association with the mating morphs on contigs from the same section of chromosome 11 (P < 0.05, Fig. 2c). The high resolution provided by the unrelated individuals identified specific segments of this region uniquely associated with either Faeder or Satellite, indicating the presence of morph-specific haplotypes spanning several megabases of the chromosome.
To fine map the variation in the region, we performed whole-genome sequencing on an additional five males (one independent, two satellites and two faeders) at 80× genome coverage (Supplementary Table 2). Nucleotide variation (measured by Watterson's theta) was substantially greater in faeder and satellite individuals within the region of interest (faeder, 0.011; satellite, 0.011) than in regions adjacent to the inversion (faeder, 0.003; satellite, 0.002). The divergence (measured by Dxy) between faeder and the other morphs was high across the entire region (Fig. 3), with morph-specific lineage sorting occurring within the region (Fig. 3c). These patterns are consistent with the presence of a large non-recombining inversion, and this was confirmed by the orientations of read pairs across the breakpoints (Supplementary Fig. 5).
(a) Divergence (Dxy) among morphs across the inversion (4.4 Mb) and flanking regions. The vertical dashed lines represent the two inversion breakpoints. The CENP-N gene spans the breakpoint located on contig 3357. Each line is staggered by 15 kb to make the lines visible. (b–d) Evolutionary relationships among the three morphs. Maximum-likelihood trees show the relationship among the sequences from two resequenced satellites, two faeders, the non-inverted reference (Independent 1) and a resequenced independent (Independent 2) for sequence adjacent to the inversion region (contig 3913: 140,000–150,000) (b), sequence within the inversion region exhibiting high divergence from the reference (contig 3357: 280,000–290,000) (c) and sequence within the inversion region where satellites show low divergence from independents (contig 3208) (d). (e) Divergence patterns between the Faeder and Satellite haplotypes across regions containing candidate loci involved in steroid hormone metabolism (PLCG2, SDR42E1, HSD17B2 and CYB5B), sperm motility (GAS8) and pigmentation (MC1R), as well as loci for transcription factors expressed in the gonads (ZFPM1) and in proximity to MC1R showing Satellite-specific divergence (TCF25). Arrows indicate the presence of morph-specific deletions affecting the coding region of PLCG2 (Supplementary Fig. 6) and deletions in noncoding regions surrounding HSD17B2 (ref. 20). Dots indicate the presence of morph-specific nucleotide substitutions (blue, Satellite; red, Faeder).
One inversion breakpoint disrupts the CENP-N gene (encoding centromere protein N) between its fourth and fifth exons. CENP-N is essential for mitotic centromere assembly18. We therefore predicted that homozygosity for the inversion haplotypes would be lethal, and our breeding data confirmed a complete absence of inversion homozygotes (Table 1). The other breakpoint appears to be in a noncoding repeat sequence.
Because recombination is completely suppressed close to inversion breakpoints, regions in linkage disequilibrium (LD) with breakpoints are expected to have high divergence7,19. This expectation was supported by divergence analyses using our resequencing data. We identified 44,433 SNPs specific to one of the three haplotypes in the assembled reference (independent) inversion region. The inverted haplotypes showed several structural changes, including regions with large (>100-bp) deletions and duplications (Supplementary Fig. 5). Within some regions of the inversion, satellites showed greater similarity to independents than faeders (Fig. 3a,d), suggesting that the Satellite allele originated through recombination or gene conversion between the inverted Faeder and non-inverted independent alleles20.
We phased SNPs located in the inversion into longer haplotypic sequences and compared 100 gene sequences directly among the three morphs. Of these gene sequences, 78% showed consistent morph-specific differences, including encoded amino acid substitutions, insertions and deletions (Supplementary Table 6). In addition, two of six genes adjacent to the inversion encoded Faeder-specific protein sequence differences. Under neutral drift and in the absence of recombination, we would expect consistent divergence across the inverted region. However, the genes in the inversion varied widely in their divergence among morphs, with several showing high divergence (Fig. 3e).
Several divergent protein-coding genes have predicted functions in hormonal systems, such as androsteroid homeostasis and plumage development, relevant to ruff morph phenotypes (Fig. 3e and Supplementary Table 7). A key candidate is HSD17B2 (estradiol 17-β-dehydrogenase 2), encoding an enzyme that preferentially inactivates testosterone to androstenedione and estradiol to estrone21. Also showing divergence among morphs are SDR42E1 (short-chain dehydrogenase reductase), ZDHHC7 (palmitoyltransferase)22 and CYB5B (cytochrome b5) (Supplementary Table 7). The morph-specific alleles of these enzymes may alter steroid secretion levels and/or receptor responsiveness, driving morphological and neurological mechanisms responsible for contrasting anatomical, plumage and behavioral profiles (Fig. 1c). Genes involved in steroid metabolism have also been implicated in another polymorphic vertebrate—the white-throated sparrow, Zonotrichia albicollis15, in which the two morphs are similarly associated with an inversion. As in ruffs, the sparrow morphs differ in aggression and testosterone levels during the breeding season23. These similarities suggest that convergent molecular pathways may contribute to the evolution of behavioral variation during reproduction. Another gene located in the ruff inversion, MC1R (melanocortin-1 receptor), a locus that controls color polymorphisms in other birds24, might account for the reduced melanin levels in satellite display feathers (Fig. 1a). In the Faeder (but not Satellite) inversion, the PLCG2 gene (1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase γ-2) has experienced complex deletions and rearrangement, including loss of an exon encoding an SH3 protein-interaction domain, and is probably a loss-of-function allele (Supplementary Fig. 6). PLCG2 encodes a transmembrane signaling enzyme involved in cell receptor activation25 that interacts with epidermal growth factor receptor (EGFR)26. As EGFR is involved in the formation of feather arrays27, PLCG2 is a candidate for the loss of secondary sexual expression of display feathers and behavior in faeders. Additional genes with roles in sperm motility and gonadal expression are also present in the inversion (Supplementary Table 7).
As homozygosity for the inversion appears to be lethal, to maintain allelic frequencies, the fitness of individuals carrying the inversion, in one or both sexes, must exceed that of homozygotes for the ancestral independent haplotype. Heterozygous carriers of the inversion also have poor survival in crosses (Table 1). Higher reproductive success by satellite and faeder males is a likely explanation for how the survival disadvantage is offset, and the larger testes of these morphs suggest that they might be more successful in sperm competition, despite equal or lower mating rates2,11. Selection should also favor disassortative mating by individuals carrying the inversion, particularly by females. As some ruff females mate with multiple morphs12, morph discrimination of mates, if it occurs, is not ubiquitous. Strong disassortative mating is a key feature of the white-throated sparrow system, although the causative inversion in this species is not lethal15.
Alternative reproductive morphs are predicted to evolve when strong reproductive skews provide low thresholds for invading forms9. The lek mating system of ancestral independents would have satisfied this condition, but numerous other species that lack genetically polymorphic alternatives also do so10. The occurrence of genomic rearrangements that can provide viable substrates for differentiation must be rare, but, by suppressing recombination and combining the fates of loci within the same genomic region, the inversion in the ruff enabled a phenotypically complex alternative strategy to evolve through the coevolution of genes affecting male behavior, morphology and fertility. As shown here and independently20, the initial occurrence of one alternative can facilitate the evolution of multiple morphs, as has also occurred in other species9,10,28,29,30.
Methods
Samples.
DNA was extracted31 from samples obtained from the Simon Fraser University colony, which was founded with 110 ruffs hatched from wild eggs collected in Finland before 1990 (refs. 4,5) plus two faeder males from the Netherlands in 2006 (under permits from the Canadian Food Inspection Agency, Canadian Wildlife Service and Simon Fraser University Animal Care Committee). Additional blood samples were obtained from wild males in breeding plumage caught in the Netherlands (ten faeders and ten satellites) between 2004–2008 or Belarus (one independent and one satellite) in 2014 (under permits from the Dutch Ringing Centre, the Animal Experimentation Committee of the University of Groningen and the Belarus Bird Ringing Centre).
Male morph determination.
Behavioral phenotypes of captive male ruffs were determined during the breeding season1,3,13,32 (Fig. 1c). Classification of ornamented males was based on ethological displays13, with some wild males assigned from plumage32,33,34,35,36. Faeders were definitively identified by small size and lack of ornamental plumage, seasonal facial wattles and epigamic display3,37.
Behavioral profiles.
We quantified the behavior in captivity of 19 independents, six satellites and two wild-caught faeders (Fig. 1c). The faeders were housed with 61 females. Two independents and one satellite were visible in an adjacent pen. These ornamented males were introduced to the faeders and females during 1- to 2-h morning observation periods. We scanned the postures and relative positions of males at 2-min intervals and recorded all interactions in 55 sessions over 50 d. Independents and satellites were replaced at least every 5 or 10 d, respectively.
Four behavioral variables summarized differences among morphs1,3,13. Aggressive behavior included total chases, bill points, bill thrusts or fights per minute. Display was defined as the proportion of scans with squats, half-squats and obliques. Proximity was the proportion of scans in which a male was positioned <2 bird lengths from another displaying male. Alert stance was the proportion of scans in which a male was standing on the lek with its head up. Morph-specific means of rates calculated for each behavior from the means of each male, standardized across the sample population, are shown in Figure 1c.
Testis volume index.
Lengths (L) and widths (W) of both right and left testes were measured by T.P. with calipers, to the nearest 0.1 mm, for prebreeding and breeding ruffs that died during capture in the Netherlands from March to June in 1993–2005 (ref. 38) (Supplementary Fig. 1a). A volume index including both testes, in mm3, was calculated assuming testes were cylindrical (L × W2 × 0.785). A non-breeding baseline index, measured during late winter, was defined as <120 mm3. Residuals from the log-transformed index were calculated for males caught between 10 April and 15 May each year showing gonadal recrudescence above baseline, using a quadratic regression controlling for date (F2,36 = 7.62, P = 0.0002; Supplementary Fig. 1).
Steroid hormone measurement.
Hormone levels were measured in blood plasma samples collected in 2003 and 2006 (ref. 39). In 2003, we sampled 16 displaying males 3–9 years old held with four other males in two groups of ten (five independents and five satellites). Blood was sampled between 9 a.m. and 2 p.m. approximately every 2 weeks before, throughout and after the breeding season (14 March to 7 July) (n = 107), and plasma was separated and stored at −20 °C. In 2006, we sampled two groups of three independents and two satellites. Two faeder males (see “Behavioral profiles”) were also sampled. Males had constant visual access to females and physical access for 2–3 h between 6 a.m. and 11 a.m., when lek attendance is highest in the wild40. Blood samples (n = 50) were collected between 10 a.m. and 12 p.m., immediately after males had access to females, approximately every 2 weeks between 14 May and 2 July.
Plasma samples were analyzed at the University of Alaska, Fairbanks, in duplicate following established radioimmunoassay (RIA) procedures41,42. Thirty 2003 samples were extracted with HPLC-grade dichloromethane. Steroids were separated using diatomaceous earth/glycol chromatography ('column' RIA), such that testosterone, dihydrotestosterone (DHT) and androstenedione could be analyzed from a single 100-μl plasma sample. Testosterone and DHT titers were strongly correlated (r2 = 0.95, n = 30, P < 0.0001); thus, the remaining 2003 samples were analyzed in two assays without separation of steroids before RIA ('direct' RIA), using a 100-μl sample for testosterone and DHT ('total T') and a 50-μl sample for androstenedione. All 2006 samples were assayed for total testosterone and androstenedione using 100 μl of plasma, with steroids separated using diatomaceous earth/glycol chromatography ('short-column' RIA). An antibody with the same cross-reactivity for testosterone and DHT (T-3003, Research Diagnostics) was used for testosterone, DHT and total testosterone RIAs, and an androstenedione-specific antibody (A-1707, Wien Laboratories) was used for androstenedione RIAs. Mean ± s.d. percentage recoveries were 55.5 ± 8.7, 45.8 ± 6.9 and 45.6 ± 7.3 for androstenedione, testosterone and DHT, respectively, in column assays; 69.1 ± 5.2 for total testosterone and 67.7 ± 6.2 for androstenedione in direct assays; and 72.7 ± 7.0 for total testosterone and 64.0 ± 10.0 for androstenedione in short-column assays. The between-assay coefficient of variation (CV) was 21% for total testosterone and 22% for androstenedione (A4), and the within-assay CVs were <10%. The minimum detectable amount was 3.90 pg/sample for androstenedione and 1.95 pg/sample for testosterone, DHT and total testosterone.
We tested for morph-specific differences in levels and temporal patterns of circulating steroids (Fig. 1e,f) using generalized linear mixed models with log linked Poisson distribution. Date, its quadratic effect and two-way interactions with the morph were included as factors. Bird was a random factor, along with assay type and year (social situation); these had no detectable effects (Supplementary Table 1).
Pedigree construction.
Preliminary pedigree assignments from 1985–2013 were generated using 27 microsatellite markers43 in 756 ruffs. We assigned parentage, including all possible candidate parents, using Cervus44 and Colony45. Inconsistencies were resolved by manual inspection, incorporating housing information for the most likely candidates.
Genome sequencing.
We sequenced the genome of an independent male from the captive colony. Illumina HiSeq 2500 v4 150-bp paired reads were generated, using paired-end and mate-pair libraries with various insert sizes, resulting in 137× raw coverage (Supplementary Fig. 2 and Supplementary Table 2). We used PacBio RS II technology with P5-C3 chemistry to generate 8.8× coverage in long reads (mean length of 5,713 bp).
Genome sequence assembly.
The genome was de novo assembled using an integrated approach (Supplementary Fig. 2).
Cleaning and trimming of raw reads. The raw Illumina paired-end library reads were quality trimmed (>q30) and adaptor sequences were removed using fastqc-mcf (ea-utils.1.1.2-537; see URLs). Short reads (<50 bases) were discarded. Mate-pair reads were quality trimmed (>q30) and adaptor and linker sequences were removed using CutAdapt46 1.3. We retained only read pairs containing Nextera linker sequence and remaining read length >50 bases.
Removal of contaminant data. An initial assembly of raw paired-end reads was prepared using the CLC Bio assembler (CLC Bio 4.2.0; see URLs). Contaminant data deriving from bacterial, parasite and viral genomes were identified using BLAST47 against the NCBI nr database, reporting the best hits with E value <1 × 10−50. Contigs likely to be contaminants were extracted, and reads mapping to these were removed.
k-mer optimization. The optimal k-mer size for assembly was estimated with sga-preqc v0.10.13 (ref. 48) and kmergenie49 v1.5924 on one lane each for each paired-end library and a k-mer sweep with ABySS50 v1.3.7 using all paired-end and mate-pair reads. sga-preqc identified an optimum k value of around 35, whereas kmergenie identified k = 30 and 38 as the optimal values and ABySS suggested k = 38. We used k = 38.
Genome assembly. Preliminary assembly of all paired-end and mate-pair reads using ABySS resulted in unitigs that were masked using RepeatMasker (version open-4.0.5; see URLs), with mate-pair reads mapped using bwa-aln and bwa-sampe (v0.7.7; ref. 51). Reads for which both from the pair mapped to the masked unitigs were retained. Filtered paired-end reads were then used to scaffold the unitigs using SSPACE52 (Basic 2.0). Further scaffolding was performed with PacBio reads using PBJelly53 v14.1.14.
RNA sequencing and annotation.
Previous RNA sequencing (RNA-seq) data in ruffs were available for genes expressed in feather follicles54. We generated new RNA-seq data from egg, chick heart, lung and brain, female heart and brain, and male heart, brain and testes to obtain a wide variety of transcripts for gene annotation (Supplementary Table 2). Raw data were quality and adaptor trimmed using CutAdapt 1.3 and assembled using Trinity55 (version r20140413). The initial transcripts were filtered for abundance (Trinity align_and_estimate_abundance.pl, filter_fasta_by_rsem_values.pl; RSEM 1.2.7; ref. 56).
Gene prediction.
We predicted genes using the Assemblage gene prediction pipeline (see URLs) up until the second round of Maker, using Maker 2.31.7 (ref. 57), cegma 2.4 (ref. 58), SNAP version 2006-07-28 (ref. 59) and GeneMark-ES 2.3e60. Highly expressed, unique transcripts from the Trinity assembly and proteins from UniProt and/or SWISS-PROT were used as evidence. Predicted genes with an annotation edit distance (AED) <1 were selected from the Maker first-pass results and used as hints to train Augustus v3.02 (ref. 61) (Supplementary Table 4). Genes predicted by Augustus on the inversion-associated contigs were further annotated using blast2go62. Annotations were checked manually to identify incorrectly split or fused genes (n = 5 and 4, respectively). In total, we annotated 125 genes (101 with known homologs) within the inversion, with 89 (71%) supported by partial mRNA transcripts.
RAD sequencing.
We chose 300 ruffs from the pedigree for low-density RAD sequencing and genetic mapping. For the initial GWAS, we used high-density RAD sequencing of 41 unrelated males with established phenotypes originating from Finland (21 independents) and the Netherlands (ten faeders and ten satellites). For the pedigree analyses, we sampled to maximize pedigree completeness, morph representation and number of generations in our pedigree. Genomic DNA was digested using the SbfI (low density) or PstI (high density) restriction enzyme, following Baird et al.16. We pooled samples from different morphs during library preparation.
SNP calling.
For calling SNPs in the pedigreed, low-density RAD sequences and in the resequenced individuals, we used the Genome Analysis Toolkit (GATK) 3.2.2 variant-calling pipeline63. Reads were aligned to the reference genome with bwa-mem 0.7.10 and realigned using GATK RealignerTargetCreator and IndelRealigner to improve alignment quality before running GATK HaplotypeCaller. Genotypes were then called using GATK GenotypeGVCFs. For the analysis of high-density RAD sequences from unrelated individuals, demultiplexing and filtering were performed using Stacks v1.21. Mapping was performed using bwa-mem 0.7.10. SAMtools v0.1.19-44428cd64 was used for filtering bam files. We only included properly mapped and paired reads, removing reads with non-primary, supplementary or terminal alignments and with mapping quality >30. SNP calling was performed using bcftools v0.1.19-44428cd65.
Confirming linkage to the satellite- and faeder-determining loci.
We used two-point linkage mapping to locate the Faeder and Satellite loci. Fourteen individuals lacking pedigree links with the remaining 286 RAD-sampled birds were excluded, resulting in an eight-generation pedigree with 286 paternal and maternal links, 186 maternal grandmaternal or grandpaternal links, and 195 paternal grandmaternal or grandpaternal links (Fig. 2a). Phenotypes were known for 189 individuals (117 independents, 38 faeders and 34 satellites); 85 additional individuals were known to not be faeders. Twelve birds were unknowns.
Satellite and Faeder causal loci were both scored as segregating in the pedigree (independents as 1/1 homozygotes and satellites and faeders as 1/2 heterozygotes at the Satellite and Faeder loci, respectively). We followed a two-stage procedure. First, 3,948 informative SNPs (3,901 with 50 informative meioses3) were tested for linkage with Satellite and Faeder using CRI-MAP v2.503 (modified by J. Maddox, University of Melbourne; see URLs). We split the pedigree into smaller families, using the crigen function of Linkage Mapping Software (X. Liu, Monsanto). We then performed two-point linkage analysis. SNPs associated with Faeder or Satellite and with LOD score >3 were associated with chromosome 11. In the second stage, all low-density RAD SNPs from contigs spanning the inversion (n = 3,810) were tested for linkage with the two morphs. Two-point mapping usually finds relatively few markers cosegregating with a causal variant. However, numerous SNPs within the chromosome 11 contigs showed highly significant perfect cosegregation (LOD >3 and recombination fraction = 0) with both Faeder and Satellite (Fig. 2b), suggesting an inversion polymorphism.
Confirming the inversion with linkage mapping.
An inversion polymorphism produces alternative marker orders segregating within the mapping panel. Thus, any marker order tested will contain erroneous recombination events that lead to larger estimates of the map length. We tested whether maps from the complete data set (286 birds) and from a subset of the data (independents only; n = 215 birds) differed in map distances. SNPs on contigs spanning the inversion, with at least 70 informative meioses and separated by ≥50 kb, which caused no parent-offspring mismatches in the pedigree, were retained for analyses, leaving 35 markers. Because of the conserved synteny between the ruff and chicken genomes, linkage map lengths were initially tested using the contig and SNP order inferred from homology to the chicken genome. Alternative orders attempted with the FLIPS option showed lower likelihoods and longer maps than the initial marker order. The sex-averaged map distances, estimated using the CRI-MAP command CHROMPIC, were 74.9 cM for the complete data set and 20.7 cM (28%) for the reduced data set (Fig. 2b). The latter measure is consistent with other avian maps of chromosome 11 (refs. 66,67,68,69), supporting the presence of an inversion polymorphism in this region.
Genome-wide association study.
Tests of association between markers and morphs were performed using GenABEL70. Correction for population stratification was performed by first calculating identity-by-state values for all SNPs in the data set and adding these values as cofactors in the model. Genome-wide significance was assessed by performing 1,000 permutations of the data.
Inversion mapping with paired-end reads.
Following Corbett-Detig et al.71, we searched for an inversion in the region of interest identified by linkage mapping and GWAS. We identified two breakpoints exhibiting morph-specific clustering of reads mapping in parallel to the (independent) reference genome at position 185,694 on contig 3357 and position 821,901 on contig 1270 (Supplementary Fig. 7). Resequenced faeders and satellites had reads mapping in parallel orientation at these breakpoints and reduced coverage (∼50%) of properly mapped reads (Supplementary Fig. 7). Using allele-specific primers, we confirmed the predicted sequence across one breakpoint (contig 1270) in three satellite males. The inverted sequence at one breakpoint showed an inserted repetitive motif in Faeder that was absent in independent, mapping to a non-LTR (long terminal repeat) retrotransposon. These two breakpoints coincided with sharp changes in between-morph divergence (Dxy; Supplementary Fig. 7) and increased heterozygosity in faeder and satellite individuals, indicating that the inversion is heterozygous in faeders and satellites.
Confirming that the inversion haplotype is lethal recessive.
The pedigree enables tests of the lethality of the inversion haplotype(s) carrying the Faeder and Satellite alleles. Because satellite and faeder morphs are imperfectly identified phenotypically, we first identified SNPs cosegregating with Faeder or Satellite. Eighteen SNPs cosegregated with Faeder with recombination fraction = 0 and an LOD score >15. For two of these SNPs (contig 3047: 314,715 and contig 3047: 314,697), we identified >30 progeny where both parents were heterozygous for the inversion haplotype. Under Mendelian segregation, we expected a 1:2:1 ratio of the AA, AB and BB genotypes among the progeny, where A is the ancestrally ordered allele and B is the inversion-associated allele. At both SNPs, the ratio of genotypes deviated significantly from expectation (Table 1), suggesting that homozygosity for the inversion is lethal. Furthermore, the observed AA:AB ratio also exceeded Mendelian expectations of 1:2, suggesting that heterozygotes also carry a viability cost (Table 1).
These results were supported by data for two further SNPs that perfectly cosegregated with Satellite and for which >30 progeny of heterozygous × heterozygous matings were observed (Table 1). For both SNPs, the lack of inversion homozygotes and the deficit of heterozygotes were significant. The lethality of the inversion was not sex specific and is probably not morph specific, although this is difficult to confirm owing to some unknown female phenotypes. In both sexes, the absence of inversion homozygotes was statistically significant, and we produced more non-inversion homozygotes than heterozygotes, although the departure from the expected 1:2 ratio was only significant in females.
We did not specify family structure in Table 1, as fertilizations in birds are independent events. A more conservative test estimated a G-test statistic for each of five paternal half-sibships with four or more offspring (27 offspring across five families) and compared the summed G-test statistics against a χ2 distribution. The complete absence of inversion homozygotes remained statistically significant (χ2 = 12.2, degrees of freedom = 5, P = 0.03).
Population genomics and phylogenomics.
Divergence (Dxy) and heterozygosity were evaluated using Evolib (see URLs). Sliding-window analyses were carried out calculated using Slide (see URLs). Maximum-likelihood reconstruction of morph phylogenies was performed using RAxML72 under the generalized time-reversible substitution model with a gamma model of rate heterogeneity (GTRGAMMA), with maximum-likelihood searches performed on 50 randomized stepwise-addition parsimony trees. Orthologous regions of the outgroup species, killdeer (Charadrius vociferus)73, were obtained through BLAST search and performing multiple alignment with the phased haplotypes.
Haplotype calling and analysis of morph-specific amino acid differences.
We established haplotypes of alleles differing from the reference for five wild ruffs resequenced to 80× coverage for all inversion contigs. We used read-backed phasing, implemented in GATK63 (version 3.3.0), phasing SNPs co-occurring on the same (or paired) sequence reads into the same haplotype. We refrained from haplotype calling in the regions within 3 kb of the breakpoints. Inversion haplotypes were, on average, longest for faeders (14.1 kb), followed by satellites (3.4 kb) and then independents (1.0 kb). Deletions longer than 100 bp were identified on the basis of a morph-specific drop in read coverage (with read depth reduced to approximately 50% in inversion carriers; Supplementary Fig. 5).
After phasing, we used the GATK FastaAlternateReferenceMaker tool to generate haplotype-specific fasta files for each contig. Using these haplotypic sequences, we predicted genes and established amino acid sequences using Augustus (see “Annotation”). Of the 125 predicted genes in the inversion, 100 (94 with known homologs) were identified by the trained Augustus algorithm established for gene prediction in the reference (Supplementary Table 6). We then aligned the amino acid sequences encoded by these genes for all six haplotypes using the ClustalW algorithm (MEGA6, 6140220; ref. 74) and identified consistent morph-specific amino acid changes in 78 genes. Candidate gene predictions (Supplementary Table 7) were verified by comparing mRNA and BLAST evidence to available gene models suggested by Maker and Augustus. Where predictions conflicted, we chose the best-supported model. We resolved the complex copy number variation, rearrangement and deletion in the Faeder haplotype of the PLCG2 locus by combining coverage maps with rearrangement-spanning read pairs.
URLs.
Assemblage gene prediction pipeline, https://github.com/sujaikumar/assemblage/blob/master/README-annotation.md; CLC Bio assembler, http://www.clcbio.com/products/clc-assembly-cell/; Slide, https://github.com/mspopgen/slide; Evolib, https://github.com/mspopgen/Evolib; GWAS and population genetic pipelines, https://github.com/mspopgen/kuepper2015; ea-utils, http://code.google.com/p/ea-utils/; RepeatMasker, http://www.repeatmasker.org/; CRI-MAP, http://www.animalgenome.org/tools/share/crimap/.
Accession codes.
The sequences reported in this paper have been deposited in the European Nucleotide Archive (ENA) database (parent project, PRJEB11172; PacBio whole-genome sequencing, PRJEB11127; RNA sequencing, PRJEB10873; low-density RAD, PRJEB10868; high-density RAD, PRJEB10855; Illumina whole-genome sequencing, PRJEB10770; whole-genome resequencing, PRJEB10677).
Accession codes
Primary accessions
European Nucleotide Archive
References
van Rhijn, J.G. The Ruff (T. & A.D. Poyser, London, 1991).
Widemo, F. Alternative reproductive strategies in the ruff, Philomachus pugnax: a mixed ESS? Anim. Behav. 56, 329–336 (1998).
Jukema, J. & Piersma, T. Permanent female mimics in a lekking shorebird. Biol. Lett. 2, 161–164 (2006).
Lank, D.B., Smith, C.M., Hanotte, O., Burke, T. & Cooke, F. Genetic polymorphism for alternative mating behaviour in lekking male ruff, Philomachus pugnax. Nature 378, 59–62 (1995).
Lank, D.B., Farrell, L.L., Burke, T., Piersma, T. & McRae, S.B. A dominant allele controls development into female mimic male and diminutive female ruffs. Biol. Lett. 9, 20130653 (2013).
Hoffmann, A.A. & Rieseberg, L.H. Revisiting the impact of inversions in evolution: from population genetic markers to drivers of adaptive shifts and speciation? Annu. Rev. Ecol. Evol. Syst. 39, 21–42 (2008).
Roberts, R.B., Ser, J.R. & Kocher, T.D. Sexual conflict resolved by invasion of a novel sex determiner in Lake Malawi cichlid fishes. Science 326, 998–1001 (2009).
Wang, J. et al. A Y-like social chromosome causes alternative colony organization in fire ants. Nature 493, 664–668 (2013).
Shuster, S.M. & Wade,, M.J. Mating Systems and Strategies (Princeton Univ. Press, 2003).
Oliveira, R.F., Taborsky, M. & Brockman, J. Alternative Reproductive Tactics: An Integrative Approach (Cambridge Univ. Press, 2008).
Hugie, D.M. & Lank, D.B. The resident's dilemma: a female-choice model for the evolution of alternative male reproductive strategies in lekking male ruffs (Philomachus pugnax). Behav. Ecol. 8, 218–225 (1997).
Lank, D.B. et al. High frequency of polyandry in a lek mating system. Behav. Ecol. 13, 209–215 (2002).
Lank, D.B., Coupe, M. & Wynne-Edwards, K.E. Testosterone-induced male traits in female ruffs (Philomachus pugnax): autosomal inheritance and gender differentiation. Proc. R. Soc. Lond. B 266, 2323–2330 (1999).
Knapp, R. & Neff, B.D. Steroid hormones in bluegill, a species with male alternative reproductive tactics including female mimicry. Biol. Lett. 3, 628–631 (2007).
Horton, B.M. et al. Estrogen receptor α polymorphism in a species with alternative behavioral phenotypes. Proc. Natl. Acad. Sci. USA 111, 1443–1448 (2014).
Baird, N.A. et al. Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3, e3376 (2008).
Farrell, L.L., Burke, T., Slate, J., McRae, S.B. & Lank, D.B. Genetic mapping of the female mimic morph locus in the ruff. BMC Genet. 14, 109 (2013).
Carroll, C.W., Silva, M.C.C., Godek, K.M., Jansen, L.E.T. & Straight, A.F. Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat. Cell Biol. 11, 896–902 (2009).
Kirkpatrick, M. How and why chromosome inversions evolve. PLoS Biol. 8, e1000501 (2010).
Lamichhaney, S. et al. Structural genomic changes underlie alternative reproductive strategies in the ruff (Philomachus pugnax). Nat. Genet. 10.1038/ng.3430 (16 November 2015).
Mindnich, R., Möller, G. & Adamski, J. The role of 17 β-hydroxysteroid dehydrogenases. Mol. Cell. Endocrinol. 218, 7–20 (2004).
Pedram, A., Razandi, M., Deschenes, R.J. & Levina, E.R. DHHC-7 and -21 are palmitoylacyltransferases for sex steroid receptors. Mol. Biol. Cell 23, 188–199 (2012).
Maney, D.L. Endocrine and genomic architecture of life history trade-offs in an avian model of social behavior. Gen. Comp. Endocrinol. 157, 275–282 (2008).
Mundy, N.I. A window on the genetics of evolution: MC1R and plumage colouration in birds. Proc. Biol. Sci. 272, 1633–1640 (2005).
Bunney, T.D. & Katan, M. PLC regulation: emerging pictures for molecular mechanisms. Trends Biochem. Sci. 36, 88–96 (2011).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Atit, R., Ronald, A.C. & Niswander, L. EGF signaling patterns the feather array by promoting the interbud fate. Dev. Cell 4, 231–240 (2003).
Joron, M. et al. Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry. Nature 477, 203–206 (2011).
Shuster, S.M. & Wade, M.J. Equal mating success among male reproductive strategies in a marine isopod. Nature 350, 608–610 (1991).
Sinervo, B. & Lively, C.M. The rock-paper-scissors game and the evolution of alternative male strategies. Nature 380, 240–243 (1996).
Bruford, M.W., Hanotte, O., Brookfield, J.F.Y. & Burke, T. in Molecular Genetic Analysis of Populations: A Practical Approach 2nd edn (ed. Hoelzel, A.R.) 287–336 (IRL Press, 1998).
Hogan-Warburg, A.J. Social behaviour of the ruff, Philomachus pugnax (L.). Ardea 54, 109–229 (1966).
Lank, D.B. & Dale, J. Visual signals for individual identification: the silent “song” of ruffs. Auk 118, 759–765 (2001).
Dale, J., Lank, D.B. & Reeve, H.K. Signaling individual identity versus quality: a model and case studies with ruffs, queleas, and house finches. Am. Nat. 158, 75–86 (2001).
van Rhijn, J., Jukema, J. & Piersma, T. Diversity of nuptial plumages in male ruffs Philomachus pugnax. Ardea 102, 5–20 (2014).
Höglund, J. & Lundberg, A. Plumage color correlates with body size in the ruff (Philomachus pugnax). Auk 106, 336–338 (1989).
Stonor, C.R. On a case of a male ruff (Philomachus pugnax) in the plumage of an adult female. Proc. Zool. Soc. Lond. A 107, 85–88 (1937).
Piersma, T., Rogers, K.G., Boyd, H., Bunskoeke, E.J. & Jukema, J. Demography of Eurasian golden plovers Pluvialis apricaria staging in The Netherlands, 1949–2000. Ardea 93, 49–64 (2005).
Morgan, T. Hormonal Regulation of Alternative Reproductive Strategies M.S. thesis, Univ. Alaska, Fairbanks (2006).
Lank, D.B. & Smith, C.M. Conditional lekking in ruff (Philomachus pugnax). Behav. Ecol. Sociobiol. 20, 137–145 (1987).
Goymann, W. & Wingfield, J.C. Competing females and caring males. Sex steroids in African black coucals, Centropus grillii. Anim. Behav. 68, 733–740 (2004).
Wingfield, J.C. & Farner, D.S. The determination of five steroids in avian plasma by radioimmunoassay and competitive protein-binding. Steroids 26, 311–321 (1975).
Farrell, L.L., Dawson, D.A., Horsburgh, G.J., Burke, T. & Lank, D.B. Isolation, characterization and predicted genome locations of ruff (Philomachus pugnax, AVES) microsatellite loci. Cons. Genet. Resource 4, 763–771 (2012).
Kalinowski, S.T., Taper, M.L. & Marshall, T.C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 16, 1099–1106 (2007).
Wang, J. An improvement on the maximum likelihood reconstruction of pedigrees from marker data. Heredity 111, 165–174 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Simpson, J.T. Exploring genome characteristics and sequence quality without a reference. arXiv http://arxiv.org/abs/1307.8026 (2013).
Chikhi, R. & Medvedev, P. Informed and automated k-mer size selection for genome assembly. Bioinformatics 30, 31–37 (2014).
Simpson, J.T. et al. ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123 (2009).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv http://arxiv.org/abs/1303.3997 (2013).
Boetzer, M., Henkel, C.V., Jansen, H.J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
English, A.C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS ONE 7, e47768 (2012).
Ekblom, R., Farrell, L.L., Lank, D.B. & Burke, T. Gene expression divergence and nucleotide differentiation between males of different colour morphs and mating strategies in the ruff. Ecol. Evol. 2, 2485–2505 (2012).
Haas, B.J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Li, B. & Dewey, C. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Campbell, M.S., Holt, C., Moore, B. & Yandell, M. Genome annotation and curation using MAKER and MAKER-P. Curr. Protoc. Bioinformatics 48, 4.11.1–4.11.39 (2002).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Conesa, A. & Götz, S. Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genomics 2008, 619832 (2008).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Groenen, M.A.M. et al. A consensus linkage map of the chicken genome. Genome Res. 10, 137–147 (2000).
Aslam, M.L. et al. A SNP based linkage map of the turkey genome reveals multiple intrachromosomal rearrangements between the turkey and chicken genomes. BMC Genomics 11, 647 (2010).
Kawakami, T. et al. A high-density linkage map enables a second-generation collared flycatcher genome assembly and reveals the patterns of avian recombination rate variation and chromosomal evolution. Mol. Ecol. 23, 4035–4058 (2014).
van Oers, K. et al. Replicated high-density genetic maps of two great tit populations reveal fine-scale genomic departures from sex-equal recombination rates. Heredity 112, 307–316 (2014).
Aulchenko, Y.S., Ripke, S., Isaacs, A. & van Duijin, C.M. GenABEL: an R library for genome-wide association analysis. Bioinformatics 23, 1294–1296 (2007).
Corbett-Detig, R.B., Cardeno, C. & Langley, C.H. Sequence-based detection and breakpoint assembly of polymorphic inversions. Genetics 192, 131–137 (2012).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Jarvis, E.D. et al. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014).
Tamura, K. et al. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Acknowledgements
We thank the laboratory and bioinformatics staff at Edinburgh Genomics at the University of Edinburgh for Illumina sequencing and the Lausanne Genomic Technology Facility for PacBio sequencing. This work was supported by a grant to T.B., J.S. and M.B. from the UK Biotechnology and Biological Sciences Research Council (BBSRC; BB/J0189371), by grants to D.B.L. from the Natural Sciences and Engineering Research Council of Canada (NSERC), by the H.F. Guggenheim Foundation, the National Geographic Society and the US National Science Foundation (NSF), and by a European Union (EU) Marie Curie fellowship to C.K. The Simon Fraser University Work-Study program facilitated the maintenance of the captive population since 1993. Hormone work was supported by the NSF Experimental Program to Stimulate Competitive Research (EPSCoR) program at the University of Alaska, Fairbanks.
Author information
Authors and Affiliations
Contributions
T.B., D.B.L., J.S., M.B. and C.K. designed the study. M.S. and J.E.R. analyzed the genomic and transcriptomic data and assembled the genome. N.d.R., L.L.F., S.B.M. and C.K. undertook the laboratory work. D.B.L. bred, raised and measured captive birds and obtained tissue samples for genetic and hormone analyses. S.B.M. and D.B.L. made behavioral observations. T.C.M., A.S.K. and J.C.W. measured hormones. T.P. and Y.I.V. obtained faeder samples, donated faeder founders to the captive population and obtained testis measurements. D.B.L., T.P., Y.I.V., N.K., P.P. and C.K. obtained tissue samples from wild birds. M.B. supervised the sequencing and genome assembly. C.K. and J.S. undertook the linkage analysis. C.K., L.L.F., N.d.R., S.B.M. and D.B.L. constructed the pedigree using microsatellite and SNP data. M.S., C.K., J.S., K.Z. and T.B. analyzed the SNP data and performed molecular evolutionary analyses. T.B., D.B.L., C.K., M.S., J.E.R., M.B. and J.S. wrote the manuscript with input from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Testis size index during late winter and spring.
(a) The shaded area delineates the sampling window used to calculate log-transformed testis size residuals during the period of gonadal recrudescence (Fig. 1d). (b) Modeled relationship (solid line) between log testes index and date from a regression of log-transformed testis size index against date and date2 during the sampling window (F2,36 = 7.62, P = 0.0002). Points, residuals; shaded area, 95% confidence limits; dashed lines, 95% prediction limits.
Supplementary Figure 2 Schematic of the genome assembly strategy.
Summary of the genome assembly strategy showing raw data (orange), programs used (white) and output generated (blue).
Supplementary Figure 3 Alignment of the genome assembly to the chicken genome.
(a) Synteny of the ruff genome assembly (y axis) with the chicken genome (galgal4; x axis) from nucmer alignment. (b) Nucmer alignment of the inversion region contigs (y axis) to chromosome 11 of chicken (x axis); contigs denoted by an asterisk are reversed.
Supplementary Figure 4 Association of SNPs with morph across the inversion in the linkage mapping population.
Numerous SNPs within the chromosome 11 inversion are perfectly associated with faeder and satellite. Differences in LOD scores reflect different morph numbers in the pedigree rather than strength of association. Note that the y axes for the recombination fraction panels are shown on a decreasing scale, with 0 representing complete linkage between the SNP and a morph-determining locus. Alternating colors indicate different contigs.
Supplementary Figure 5 Morph-specific variation in read depth throughout the inversion contigs.
Read depth of reads mapping to contigs across the inversion for one independent (yellow), two satellite (blue) and two faeder (red) individuals. Arrows indicate regions showing morph-specific differences in read depth that suggest the presence of morph-specific deletions or copy numbers variants.
Supplementary Figure 6 Complex, Faeder-specific deletions and rearrangements in the genomic region encoding the transmembrane signaling receptor PLCG2.
(a) Read mapping from a representative faeder individual sequenced at 80× (viewed in IGV) to a 100-kb region of the independent reference genome showing blocks with haploid (40×), diploid, triploid and tetraploid coverage. The predicted intron-exon structures of PLCG2 and three downstream genes are shown, and the predicted effects of deletions (see panel c) on exon retention are indicated. (b) Cartoon of the 100-kb region illustrating coverage, SNP patterns in each block and read-pair mappings that define rearrangements in the region. (c) Model of the Faeder PLCG2 region, consistent with the genomic evidence, and its origin through deletion and tandem duplication. Deletion of block B deletes exon 17, encoding the SH3 domain of PLCG2, and deletion of blocks D3 and E deletes exons 24–26 of PLCG2. Deletion of block Y6 deletes the first exon of gene 18877. Deletion of block Y4 (shared with Satellite) and of block Y2 has no effect on the predicted coding sequences. In the model, the relative timing of the Y2, Y4 and Y6 deletions is not resolved.
Supplementary Figure 7 Patterns of divergence and read depth at inversion breakpoints in the assembly.
(a,b) Divergence (Dxy) for the contigs containing the inversion breakpoints. (c,d) Read depth at and around the inversion breakpoints for the faeder and independent morphs. Black lines close to the x axis indicate the location of reads mapping in parallel to the reference. The red and black, solid and dashed lines indicate the read depth of forward (fwd) and reverse (rev) reads mapping in the correct orientation to the reference. Data for one faeder and one independent individual only are shown for clarity.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–5 and 7. (PDF 1296 kb)
Supplementary Table 6: Annotated genes within the inverted region.
A list of annotated genes within the inverted region, including start and stop positions, gene names and descriptions from BLAST searches, and the presence or absence of morph-specific variants. (XLSX 24 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the articles Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Küpper, C., Stocks, M., Risse, J. et al. A supergene determines highly divergent male reproductive morphs in the ruff. Nat Genet 48, 79–83 (2016). https://doi.org/10.1038/ng.3443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3443
This article is cited by
-
Sex-specific morphs: the genetics and evolution of intra-sexual variation
Nature Reviews Genetics (2023)
-
Cellular profiling of a recently-evolved social behavior in cichlid fishes
Nature Communications (2023)
-
The roles of sexual selection and sexual conflict in shaping patterns of genome and transcriptome variation
Nature Ecology & Evolution (2023)
-
A novel distribution of supergene genotypes is present in the socially polymorphic ant Formica neoclara
BMC Ecology and Evolution (2022)
-
The significant role of post-pairing male behavior on the evolution of male preferences and female traits
Communications Biology (2022)