Abstract
Molecular shape recognition for larger guest molecules (typically over 1 nm) is a difficult task because it requires cooperativity within a wide three-dimensional nanospace coincidentally probing every molecular aspect (size, outline shape, flexibility and specific groups). Although the intelligent functions of proteins have fascinated many researchers, the reproduction by artificial molecules remains a significant challenge. Here we report the construction of large, well-defined cavities in macromolecular hosts. Through the use of semi-rigid dendritic phenylazomethine backbones, even subtle differences in the shapes of large guest molecules (up to ~2 nm) may be discriminated by the cooperative mechanism. A conformationally fixed complex with the best-fitting guest is supported by a three-dimensional model based on a molecular simulation. Interestingly, the simulated cavity structure also predicts catalytic selectivity by a ruthenium porphyrin centre, demonstrating the high shape persistence and wide applicability of the cavity.
Similar content being viewed by others
Introduction
The ultimate challenge in host–guest chemistry is the design of synthetic hosts that can universally recognize guest molecules with various structural characteristics including size, shape and details such as the positions of functional groups. Of particular scientific interest is the concept of the artificial enzyme, which can act as a catalyst for a specific substrate via cooperative recognition1. The importance of practical materials can be seen in solution-phase recognition, which allows efficient chemical separation despite very small structural differences2. The essential requirement for recognition is a shape-persistent cavitand generally made of covalent macrocycles3,4,5 or supramolecular cages6,7,8,9. However, recognition has been limited to relatively small guest molecules of subnanometre size (up to two aromatic rings). Apart from particular examples10,11, attempts at simple enlargement of the cavitands without a strategy are generally fruitless because the larger host molecules are no longer shape persistent. One typical example is the case of dendrimers. The molecular weight and chemical structure of the dendrimers are unity. In addition, the core-shell architecture generally provides a unique encapsulation effect12. At the dawn of dendrimer chemistry, it was believed that the giant inner cavity would be applicable to various types of molecular recognition, similar to natural enzymes13, and that dendrimers were promising candidates for large-scale recognition14. The idea was that they would act as large containers, allowing encapsulation of many guest molecules simultaneously15,16,17. However, the previous dendrimers were found to be suitable only for identification of simple and small guests18,19,20,21,22. Although dendrimers are potential containers for a larger guest or many guest molecules, they can become compact through back-folding of their terminal monomers towards the inside of the cavity, leading to loss of the cavity23,24,25. Meanwhile, dendritic structures with extra-rigid backbones do not participate to a significant extent in host–guest binding26,27. Except for some intriguing approaches such as molecular imprinting28, molecules >1 nm are still difficult to recognize. Even now, it is impossible achieving this only by the molecular shape because this requires detailed design of the nanospace through a semi-rigid architecture.
In this study, we revisit host–guest chemistry using a modern dendrimer design for the host molecules. Phenylazomethine dendrimers composed of rigid π-conjugated backbones exhibit exceptionally fine coordination processes, such as step-by-step assembly29,30,31, allowing one-atom controlled synthesis of metal clusters32. In addition, previous studies have shown that the stable quadrupole character of such dendrimers allows an efficient photoelectric process based on their rectification properties33. By using this unique dendrimer, we successfully achieve very fine recognition of large guest molecules.
Results
Screening and refinement of guests
The host molecules employed in this study are dendrimers with a metalloporphyrin core as a binding centre for guest molecules such as pyridine derivatives. Four dendron units were horizontally spread to the outside from the porphyrin core, whereas the two axial spaces were retained (Fig. 1). For example, the experimental hydrodynamic diameter of ZnG4a is over 4 nm. The percentage of free volume in the dendrimer, defined as vacant space of the atomically occupied van der Waals volume in the molecular hydrodynamic space, is ca. 80% (ref. 34). To study the characteristics of these synthetic cavities, we initially investigated binding between a guest molecule and the zinc porphyrin core buried in the cavity. A pyridine derivative was used as a guest molecule, because 1:1 complex formation between zinc porphyrin and this derivative has been established and is widely accepted35,36. ZnG4a was synthesized according to a literature method37. By comparing the binding constants of ZnG4a and the corresponding first-generation model (ZnG1a), we were able to estimate the steric contribution of the dendron groups to host–guest complexation. Thereafter, the ratio KG4a/KG1a was estimated for each guest molecule (pyridine derivatives, Fig. 2) as an index parameter for adaptability with the synthetic cavity.
The dendrimers are composed of three different parts. The core unit is a zinc or ruthenium carbonyl porphyrin, which can bind one pyridine derivative at the axial position. The bridges (B) are linear or bent phenylene groups, adjusting the position and direction of the dendron subunits. The dendron (D), which consists of dendritic phenylazomethine or benzylether groups, creates a nanospace through the formation of a rigid or flexible shell, respectively.
Shape-persistent pyridine derivatives were selected as molecular ‘keys’ for the host molecules based on the dendrimer architecture. Each key was composed of a pyridine and several phenyl groups with different connections. The pyridine group acted as the connecting tip to the axial coordination site of the zinc porphyrin core fixed in the dendrimer host. The phenyl group acted as an identifier for the molecules, recognized by the cavity around the axial ligation site of the host.
Initial screening was carried out using pyridine derivatives with several aromatic components with various connections. An ultraviolet–visible titration analysis was carried out in a toluene/acetonitrile (v/v=1:1) solvent, followed by theoretical curve fitting of the experimental results to obtain binding constants with the synthetic hosts at 20 °C (Supplementary Table S1). The binding stoichiometry of the pyridine to the zinc porphyrin core (axial position) is 1:1, as determined by ultraviolet–visible absorption, isothermal titration calorimetry (ITC) and NMR spectroscopy (a typical example is shown in Fig. 3). The binding constants were independently obtained using all of these measurements, and each result was found to be in good agreement with the others. It is notable that the binding affinity with the dendrimer host ZnG4a varied in a sensitive manner depending on the shape of the guest molecule. Pyridine derivatives containing two phenyl groups at different positions (5, 6, 7, 8, 9 and 10) gave very different binding constants (KG4a), although their formulas and molecular weights were identical. One of the derivatives, which was Y-shaped (4-(3,5-diaryl)phenylpyridine, 11) was found to coordinate to the dendrimer host with a larger binding constant, whereas guests with other shapes (15, 16, 17 and 18) did not coordinate so strongly (Fig. 3f). Even the shapes of very large molecules (ca. 19 Å width) with the same chemical formula (12, 13 and 14) were correctly recognized. In contrast, among the selected guest molecules, there was little variation in binding affinity with the non-dendritic host (ZnG1a). These observations are evidence of shape recognition, but not of an electronic effect caused by the electron-donating/withdrawing character of the substituted groups. The dendrimer shell exhibited a positive effect on the binding of Y-shaped guests, as can be observed from the value of the binding constant ratio KG4a/KG1a (up to 4.2 for 11). Previous reports on host–guest chemistry involving dendrimer architecture have shown that an increase in the generation number results in a severe decrease in binding affinity because of the shell effect18,19,20,21,22. In fact, the dendrimer host under discussion exhibits unusual enhancement on host–guest complexation.
(a) Schematic representation of cooperative binding. Docking tests for guest molecules with various shapes allowed probing of the shape of the cavity. (b) Ultraviolet–visible absorption spectra (20 °C) of ZnG4a (2.5 μM) on addition of the guest molecule 20 in toluene/acetonitrile (1:1). (c) Plots of the change in absorbance at two different wavelengths for various concentrations of 20. A theoretical fitting curve (solid line) was applied to the absorbance data. (d) Isothermal titration calorimetric data for the addition of 20 to a solution of ZnG4a (51 μM) in toluene/acetonitrile (1:1). (e) A plot of the molar reaction heat for each titration point. Theoretical fitting of the experimental data gave the thermodynamic parameters for host–guest binding. (f) Host–guest-binding constants (K) between zinc porphyrin-containing dendrimers (ZnG1a and ZnG4a) and pyridine derivatives in toluene/acetonitrile (1:1). Overall, Y-shaped guest molecules showed high binding constants.
To seek the best-fitting molecule, we carried out further tuning of the Y-shaped guests. Chemical modification of the tips of 11, giving 19, 20 and 21, increased the binding constant (KG4a). Of particular interest was the finding that chemical modification of the ‘dip’ of the Y-shaped pyridine 25 significantly enhanced the binding strength, resulting in a high binding constant ratio (KG4a/KG1a) of up to 19 for 25. In sharp contrast, such an enhancement was not observed for an I-shaped derivative (24). The dendrimer shell did not act as a positive factor for other guests such as unsubstituted pyridine (1) and rigid I-shaped pyridines (4 and 10). Further, the kinked molecules 5, 6, 7 and 8, and oversized derivatives that deviated from the Y-shape (12, 22 and 23), were excluded from the host. Thus, the common Y-shaped structure acted as an anchor that specifically bound to the dendrimer host.
Modulation of interactions
The dependence of complementary host–guest binding on the host structure was studied by testing zinc-porphyrin-cored phenylazomethine dendrimers (ZnG n a: n=1~4) with generation numbers of 1–4. This examination revealed that a dendritic structure with at least four generations is essential for the formation of a cavity that is capable of molecular discrimination. Indeed, no shape-selective binding to the host was observed when the generation number was <4 (Fig. 4a). A rough estimate of the binding structure using a molecular space-filling model suggested that the cavities in dendrimers of generation 3 or less were too shallow to tightly anchor the guest.
(a) Dependence on the generation number (n) of the dendrimer ZnG4 n. The fourth-generation dendrimer (ZnG4a) showed a strong affinity for Y-shaped guest molecules. No significant dependence was found for dendrimers with a generation number <3. (b) Dependence on cavity size and flexibility of the dendrimers. The specific recognition observed for ZnG4a was lost when the cavity size was extended or shrunk. The flexible backbone also excluded the larger guest molecules.
The structure of the dendrimer significantly affected host–guest complexation through shape modification of the cavity. We synthesized dendrimers with alternative structures: one had four dendrons at the meta-positions on the phenyl groups of the zinc porphyrin core (ZnG4b), whereas the other had four dendrons at the para-positions with 1,4-phenylene spacers (ZnG4c). On the basis of space-filling models obtained by molecular modelling, it became apparent that ZnG4c had an extended cavity, whereas that of ZnG4b was shrunken. These molecular images were confirmed by exploring the free volume percentages within the hydrodynamic volume (ZnG4a, 78%; ZnG4b, 71%; ZnG4c, 77%)34. The ZnG4b host, which had a narrow cavity, did not allow binding of the guest molecule 20, but the ZnG4c host, which had an extended cavity, also showed a lower binding affinity with 20. ZnG4c preferred larger guest molecules such as 14 and 22. We also examined a flexible dendrimer composed of a benzylether structure (ZnG4d) with the same topological architecture around the zinc porphyrin core, but with a very small free volume percentage (45%)38. As shown in previous studies, flexible dendrons exhibit exclusive host–guest binding18,20,21,22. The host ZnG4d allowed binding only of small guest molecules (1 and 4) with one or no phenyl group (Fig. 4b). This observation was consistent with previous findings that flexible dendrimers simply decrease guest molecular binding (the well-recognized ‘shell effect’).
Conversely, exchanging the metal complex did not affect the selectivity of the host molecule to the same extent. The ruthenium-porphyrin-cored phenylazomethine dendrimer RuG4a, a carbonyl-capped complex, showed almost the same trend in its shape selectivity as ZnG4a (Supplementary Fig. S1). This fact indicated that metal complexation in the porphyrin core did not affect the shape of the cavity.
Structural analysis
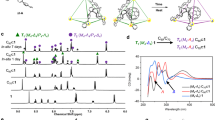
To determine the binding state, a 1H NMR analysis was carried out using the Y-shaped guest molecule 20 with tert-butyl (tBu) groups. The chemical shift corresponding to the protons of the tBu unit in 20 was shifted upfield on addition of 20 to a ZnG4a solution at 20 °C (Fig. 5a). This observation implied that chemical exchange between the trapped and free-guest molecules was faster than the NMR timescale (a few milliseconds). At a low temperature (−20 °C), we could observe the proton signals corresponding to the trapped and free-guest molecules. When two equivalents of 20 were added, the signal integration values of these trapped and free-guest molecules were almost equivalent, supporting 1:1 host–guest complexation. One of these signals appearing downfield had a chemical shift (δ) of 1.31 p.p.m. and a spin-lattice relaxation time (T1) of 684 ms, which were identical to those of the free-guest molecule (δ=1.31 p.p.m., T1=665 ms for 20), whereas the other signal exhibited a very different chemical shift (δ=1.02 p.p.m.) and relaxation time (T1=874 ms). The longer T1 for the signal at 1.02 p.p.m. can be understood based on effective encapsulation by the dendrimer shell, as previously reported16,18,37,39,40. This behaviour is typical for the environment in rigid phenylazomethine dendrimers, where T1 and T1/T2 increase were observed for the encapsulated protons as the generation number increases37,41. This increase is interrelated as a slower dynamic motion of the dendrimer (longer rotational correlation time)42.
(a) Variable-temperature 1H NMR spectra of a ZnG4a with 20 at a mixing ratio of 1:2 (H:G) in toluene-d8/acetonitrile-d3 (2:1). The signals of tBu groups in the guest molecule (20) are shown for bound and free states. (b) A computer-generated space-filling model of ZnG4a and a cross-sectional image shown as a solvent-accessible surface (probe radius: 1.4 Å) with 20. The porphyrin core is highlighted in magenta. This 3D coordinate was obtained from semi-empirical molecular orbital calculations (MOPAC-AM1). (c) 1H DOSY-NMR spectrum of a RuG4a with 20 at a mixing ratio of 1:2 (H:G) in dichloromethane-d2. (d) 1H nuclear Överhauser enhancement spectroscopy-NMR spectrum of RuG4a with 20 at a mixing ratio of 1:1 (H:G) in dichloromethane-d2. The inset figure is a 3D model of the host–guest complex.
1H diffusion-ordered spectroscopy (DOSY)-NMR measurements supported our contention that the signals at 1.31 and 1.02 p.p.m. corresponded to the free and binding guest, respectively. The diffusion coefficient (D) of ZnG4a (2.3 × 1010 m2 s−1) determined by DOSY-NMR was much smaller than that of zinc tetraphenylporphyrin (9.2 × 1010 m2 s−1) because of the larger hydrodynamic radius (Supplementary Fig. S2). These values are comparable to the diffusion coefficients of ZnG4a (1.1 × 1010 m2 s−1) and zinc tetraphenylporphyrin (8.0 × 1010 m2 s−1) in tetrahydrofuran, calculated from the hydrodynamic radii using the Stokes–Einstein equation34. If the guest molecule binds to ZnG4a, the D-value corresponding to the binding guest molecule should be identical to that of the dendrimer host molecule because they would diffuse together in the solution. However, D of the tBu unit corresponding to ZnG4a–20 was slightly different on the diffusion axis even at a low temperature (−20 °C). This result indicated that the timescale of chemical exchange between the complex and the free base was close to the diffusion delay (40 ms). For quantitative analysis, we next employed RuG4a, in which binding was stronger than that in ZnG4a. It was found that the value of D for the tBu unit corresponding to RuG4a–20 agreed exactly with the value of D for the dendron moieties (Fig. 5c).
1H nuclear Överhauser enhancement spectroscopy-NMR measurement of the RuG4a–20 system was carried out for the stable complex over a 100-ms timescale. This technique has been extensively used to elucidate dendrimer-based host–guest systems43. In the two-dimensional spectrum, many cross-peaks were observed among the aromatic protons of the dendron subunits as intramolecular nuclear Överhauser effect (NOE) couplings. In addition, a clear cross-peak between the tBu protons of the binding guest molecule (20) and a specific aromatic proton signal of the dendron subunit was also observed as intermolecular NOE coupling (Fig. 5d). As the phenylazomethine monomer had two non-equivalent aromatic rings at syn- and anti-positions, the NMR proton signals for every aromatic ring in one dendron subunit could not be reconciled. On the basis of this, the tBu protons of the binding guest molecule (20) were thought to be close to a specific arene in the dendrimer cavity without any tilt or rotation. It is noteworthy that the generation-3 dendrimer RuG3a did not show any NOE cross-peak with 20, suggesting that the fourth layer of the dendrimer contributed to guest stabilization, as noted above. This result feasibly explains the molecular picture of the complex in a three-dimensional (3D) space (Fig. 5b) in which the monomer unit on the fourth layer of a dendron approaches the tBu unit of the guest molecule (20), probably because of weak CH/π interactions. The upfield shift of the aliphatic proton signal observed in the 1H NMR spectrum of the ZnG4a–20 system (Fig. 5a) is a typical behaviour for a CH/π interaction44,45.
Thermodynamic analysis
Thermodynamic data from ITC and ultraviolet–visible analysis (van′t Hoff plot) provided information about the mechanism of molecular recognition. The values of ΔH° (enthalpy change) and ΔS° (entropy change) obtained from ITC analysis were in good agreement with those from the van′t Hoff analysis and were very different for each host–guest pair. In the case of 20 (a good fit), ΔH° had a very large negative value, whereas −TΔS° (T=293 K) had a large positive value (Supplementary Table S1). In contrast, the absolute values of ΔH° and −TΔS° for the more badly fitting molecules were smaller. These facts can be interpreted to mean that maximal contact between the host and guest result in greater stabilization in terms of free energy (ΔG°=ΔH°−TΔS°). This stabilization is mainly due to the large intermolecular force associated with the enthalpy factor (ΔH°). Surprisingly, the value of ΔH° for 20 was almost twice that for 4, which implies that the contributions from the upper ‘anchor’ structure and from pyridine coordination were nearly equivalent. This increase in ΔH° can be understood as a summation of weak intermolecular interactions including π/π, CH/N, CH/π, van der Waals attraction and hydrophobic interaction. The non-dendritic host–guest system gave a much smaller ΔH° variation for guest molecules with different shapes. Therefore, the primary factor in the high molecular recognition ability of the dendrimer ZnG4a is thought to be the enthalpy contribution, based on the large contact area between the host and guest molecules. An impressive difference was found between guest molecules 24 and 25, although the only structural difference was the presence of two phenyl rings in 25. Although the alkyl chain in 25 contributes the tight binding very much, the same structure in 24 did not contribute any improvement in host–guest binding. A similar difference was found between 26 and 27.
The titration experiments demonstrated that binding between the host and the Y-shaped pyridine in pure toluene was not as strong as in a mixed solvent of toluene/acetonitrile (1:1). A higher mixing ratio of a less solubilizing medium (acetonitrile) allowed stronger binding based on an enthalpic driving force (Supplementary Table S2). In particular, the use of a specifically polar aromatic solvent (for example, dimethyl phthalate) as a good medium significantly enhanced the binding affinity (KG4=5.4 × 105 M–1) and selectivity (KG4a/KG1a=142) of the reaction, as shown in Supplementary Table S3. This result indicates the dominance of a non-classical hydrophobic effect as discussed by Diederich and co-workers46, which enables stronger host–guest binding in a dendrimer system. The thermodynamic behaviour is different from that of the classical hydrophobic interaction, in which entropic favourability due to desolvation is a principal driving force47. In general, hydrophobic interactions do not have a crucial role in molecular recognition. However, it is interesting to note that the solvent effect enhances selectivity in the present system. This enthalpic favourability was found only in the shape-matching guest molecules and not in the other pyridine derivatives.
Selectivity in catalysis
The synthetic nanocavity can select favourable guest molecules based on their shapes. In a natural enzyme, this principle is utilized to achieve excellent selectivity between substrates and products. On the basis of a Michaelis–Menten model, the stability of an intermediate enzyme–substrate complex is important in the determination of the entire kinetics. Catalytic epoxidation of aromatic olefins by a ruthenium porphyrin (RuPor) was demonstrated48,49,50. The mechanism of catalysis, using a 2,6-dichloropyridine-N-oxide as the oxidative reagent, is well established, as follows: (1) the reagent oxidizes the RuPor to afford an oxo-complex (Ru=OPor)49. (2) An olefin derivative approaches the oxygen atom on Ru and forms an adduct as the intermediate state51. (3) The Ru=OPor reverts to RuPor by desorption of the epoxide product. We reasoned that if a RuPor in a dendrimer-based deep cavity was used as a catalyst, molecular recognition of the intermediate state might significantly affect the kinetics as predicted by the Michaelis–Menten model.
Before the experiment, we simulated the structure of intermediate species with various olefins. For example, methoxystyrene has three structural isomers (o-, m- and pMeO-Sty). These substrates were tested with the 3D structural model of the dendrimer verified from the above-mentioned host–guest chemistry. In this prediction analysis, mMeO-Sty fit the cavity very well, whereas the pMeO-Sty conflicted with the sidewall, as shown in Fig. 6a,b.
(a) Reaction scheme. (b) 3D simulation model of mMeO-Sty in the RuG4a cavity. This substrate fits in the cavity in a side-on position in relation to the oxygen atom on the RuPor core. (c) 3D simulation model of pMeO-Sty in the RuG4a cavity. The methoxy group conflicts with the sidewall. (d) Ratio of the reaction rate catalysed by RuG4a and RuG1a for each methoxystyrene substrate. (e) Ratio of the reaction rate catalysed by RuG4a and RuG1a for each phenyl styrene substrate.
The kinetics were determined (Supplementary Table S4) for each pair of two dendrimers (RuG1a and RuG4a) and two substrates (mMeO-Sty and pMeO-Sty). The reactions were carried out using 100 mmol l−1 of the substrates, an equimolar amount of oxidative reagent (2,6-dichloropyridine-N-oxide) and 0.05 mol% of the catalyst (RuG1a and RuG4a) at room temperature (25 °C) in chloroform. The reaction progress was monitored using 1H NMR and gas chromatography. Although the entire kinetics of the reaction using RuG4a (VG4) were slower than those for RuG1a (VG1) because of high steric hindrance, the ratio VG4/VG1 clearly demonstrated the effect of the cavity on substrate selectivity. As shown in Fig. 6c, the ratio VG4/VG1 for mMeO-Sty was twice that for pMeO-Sty. To achieve a greater understanding of the catalytic activity, the dependence on substrate concentration was examined based on a Michaelis–Menten model. In the case of a ruthenium tetraphenylporphyrin monocarbonyl catalyst, the apparent product formation kinetics showed almost linear dependence on the substrate concentration between 100 and 1,000 mmol l−1 (Supplementary Fig. S3). The Michaelis–Menten constant was determined to be >5 mol l−1 for mMeO-Sty. In contrast, a much smaller concentration dependence was observed when RuG4a was used as a catalyst, suggesting a higher affinity between mMeO-Sty and RuG4a. The estimated Michaelis–Menten constant for RuG4a was 90 mmol l−1. A similar kinetic investigation was also applied to mPh-Sty and pPh-Sty, which exhibited almost the same trend. This suggested that selectivity in the present experimental conditions ([S]=100 mM) was thermodynamically controlled by the enzyme–substrate complex equilibrium, but not kinetically by complex formation. Of particular importance is the agreement between the experimental results and the simple simulation based on a static molecular 3D model. These results are in contrast to previously reported selectivity by dendrimers in terms of predictability52.
Discussion
Although the entropy factor (−TΔS°) acts as a negative driving force for binding, this is smaller than the positive driving force because of enthalpy (ΔH°). Similar to the literature5,13,53,54, the host–guest system under discussion showed a good linear correlation (correlation coefficient: r=0.98) between ΔH° and TΔS° (Supplementary Fig. S4). This behaviour, which is called enthalpy-entropy compensation, has been widely observed in natural and synthetic host–guest systems. As it is known that an apparent correlation between ΔH° and TΔS° may also be observed because of a statistical artifact, we analysed these data with caution13. Indeed, we examined the reproducibility of host–guest binding between ZnG4a and 20 five times, and the resulting experimental values (ΔH° and TΔS° by ITC) were found to be within a typical experimental error of ±10%, stemming from the summation of measurements (weight or volume) and data processing (curve fitting). The experimental values of K (ΔG°) and ΔH° for all of the guest molecules were distributed over a substantial range exceeding the error range; therefore, the observed relationship may be regarded not as an apparent correlation based on statistical error but as a physical phenomenon. Here an unexpectedly low proportional coefficient α (−TΔΔS°/ΔΔH°=0.75 at 293 K) was observed for the ZnG4a system; this was substantially lower than that of the non-dendritic zinc porphyrin (zinc tetraphenylporphyrin: α=0.91). The observed relationship suggests that 25% (1–α) of the enthalpy gain can be used for free-energy gain (ΔG°) on molecular recognition. This slope is similar to that of α-cyclodextrins and rather smaller than that of γ-cyclodextrins5. The intercept of the enthalpy-entropy compensation plot gave a value of TΔS0°=13.9 kJ mol–1, close to the value for γ-cyclodextrins5. This intercept value is referred to as an entropy gain accompanying desolvation from the binding centre in the cavity of the free host molecule. The relatively small α (comparable to α-cyclodextrins) and large TΔS0° (comparable to γ-cyclodextrins) indicate good shape persistence of the large cavity in the dendrimer.
The significant contribution of the long alkyl chain on 25 can be interpreted as the hydrophobic effect or CH/π interaction, which enhances host–guest affinity. An upfield shift of the NMR proton signals corresponding to the alkyl chain was observed on complexation with ZnG4a. This result indicated a cooperative recognition mechanism55 with interlocking at each recognition site including the pyridine (tip), Y-shaped aromatic rings (anchor) and long alkyl chain moieties (thread) as shown in Fig. 7. In spite of the significant contribution of the anchor and thread in 25 (an additional 40 kJ mol−1 ΔH°), it is notable that 25 did not show any interaction with the free-base porphyrin form of the dendrimer (H2G4a), as monitored by ultravioilet–visible titration and ITC. This result demonstrates that prefixing of the guest molecules through pyridine coordination is the key to the subsequent molecular shape recognition. In addition, further interlocking, which can be seen as the second key, allows the long alkyl chain to function as a positive driving force. Significant amplification of recognition was achieved through sequential cooperativity between different parts of the guest molecule, which may be related to an induced-fit mechanism13,47. Such a mechanism may be the key to understanding protein-based recognition systems.
In conclusion, we have created a synthetic keyhole that finely discriminates the shape of a specific molecular key. The essential element in probing the multidimensional structure of guest molecules is the shape persistence of the macromolecular host in terms of conformation and cavity geometry. Whereas the shape of the cavity in conventional dendrimers is undefined, an appropriate design with sufficient rigidity allowed us to provide a large cavity with a programmed function for molecular recognition and catalysis. Further, the reasonably semi-rigid keyhole, in which shape persistence and adaptability go together, provides cooperative recognition, allowing amplification of recognition through complexation-induced fixation of the cavity structure, which may be related to allosteric nature of this kind of host molecule56.
Methods
Chemicals
Phenylazomethine dendrimers with a zinc porphyrin core (ZnG n a: n=1, 2, 3 and 4) used in this study were synthesized by a convergent method as described in the literature37. Other types of phenylazomethine dendrimer (RuG4a, ZnG4b and ZnG4c) were newly synthesized as described in the Supplementary Methods. A benzylether (Fréchet type) dendrimer with a zinc porphyrin core was synthesized as described in the literature57. All pyridine derivatives used as guest molecules were purchased or synthesized as described in the Supplementary Methods.
NMR
1H and 13C NMR spectra were obtained using a Bruker Avance III-400 spectrometer equipped with a 5-mm BBFO z-gradient probe (gradient strength: 57 G cm−1). The chemical shift values were calibrated to an internal tetramethoxysilane standard. The DOSY experiment was carried out with the following settings. The diffusion delay was in the range of 40–60 ms depending on the sample. The gradient pulse length and the gradient strength were varied from 1.5 to 3.0 ms and from 2 to 100%, respectively. The 1H–1H nuclear Överhauser enhancement spectroscopy experiment was carried out using standard pulse sequences on the same spectrometer with a 120-ms mixing time.
Determination of binding parameters
Coordination constants and thermodynamic parameters for all host–guest pairs were determined by titration experiments based on ultraviolet–visible absorption measurements and isothermal calorimetry at 20 °C (see Supplementary Figs S5–S30). Ultraviolet–visible absorption spectra were measured on a Shimadzu UV-3150PC spectrometer. Curve-fitting analyses based on a least-squares method at two independent wavelengths were done using homemade software to determine the binding constants. ITC was carried out using a Microcal VP-ITC instrument. Analysis was carried out on built-in software to determine the binding constants and enthalpy and entropy differences (ΔH and ΔS).
Determination of the kinetic parameters
Catalytic epoxidation of styrene derivatives by the RuPor catalyst was initially confirmed by 1H NMR measurements in CDCl3. Detailed kinetic data in CHCl3 were obtained by monitoring the epoxide product using a gas chromatograph (Shimadzu, GC-14B) with a TCD detector and He carrier gas. Turnover frequency values were determined from the time course of product formation. All the reactions were carried out under a N2 atmosphere at 25 °C.
Additional information
How to cite this article: Imaoka, T. et al. Macromolecular semi-rigid nanocavities for cooperative recognition of specific large molecular shapes. Nat. Commun. 4:2581 doi: 10.1038/ncomms3581 (2013).
References
Krishnamurthy, V. M. et al. Carbonic anhydrase as a model for biophysical and physical-organic studies of proteins and protein-ligand binding. Chem. Rev. 108, 946–1051 (2008).
Mitra, T. et al. Molecular shape sorting using molecular organic cages. Nat. Chem. 5, 276–281 (2013).
Harada, A. Cyclodextrin-based molecular machines. Acc. Chem. Res. 34, 456–464 (2001).
Izatt, R. M., Pawlak, K. & Bradshaw, J. S. Thermodynamic and kinetic data for macrocycle interaction with cations and anions. Chem. Rev. 91, 1721–2085 (1991).
Rekharsky, M. V. & Inoue, Y. Complexation thermodynamics of cyclodextrins. Chem. Rev. 98, 1875–1918 (1998).
Diederich, F. & Gómez-López, M. Supramolecular fullerene chemistry. Chem. Soc. Rev. 28, 263–277 (1999).
Yoshizawa, M., Tamura, M. & Fujita, M. Chirality enrichment through the heterorecognition of enantiomers in an achiral coordination host. Angew. Chem. Int. Ed. 46, 3874–3876 (2007).
Pluth, M. D. & Raymond, K. N. Reversible guest exchange mechanisms in supramolecular host-guest assemblies. Chem. Soc. Rev. 36, 161–171 (2007).
Rebek, J. Jr. Molecular behavior in small spaces. Acc. Chem. Res. 42, 1660–1668 (2009).
Ferrand, Y., Crump, M. P. & Davis, A. P. A Synthetic lectin analog for biomimetic disaccharide recognition. Science 318, 619–622 (2007).
Fujita, D. et al. Protein encapsulation within synthetic molecular hosts. Nat. Commun. 3, 1093 (2012).
Hecht, S. & Fréchet, J. M. J. Dendritic encapsulation of function: applying nature’s site isolation principle from biomimetics to materials science. Angew. Chem. Int. Ed. 40, 74–91 (2001).
Houk, K. N., Leach, A. G., Kim, S. P. & Zhang, X. Binding affinities of host-guest, protein-ligand, and protein-transition-state complexes. Angew. Chem. Int. Ed. 42, 4872–4897 (2003).
Zeng, F. & Zimmerman, S. C. Dendrimers in supramolecular chemistry: from molecular recognition to self-assembly. Chem. Rev. 97, 1681–1712 (1997).
Bauer, R. E., Clark, C. G. & Müllen, K. Precision host-guest chemistry of polyphenylene dendrimers. N. J. Chem. 31, 1275–1282 (2007).
Jansen, J. F. G. A., de Brabander-van den Berg, E. M. M. & Meijer, E. W. Encapsulation of guest molecules into a dendritic box. Science 266, 1226–1229 (1994).
Broeren, M. A. et al. Multicomponent host-guest chemistry of carboxylic acid and phosphonic acid based guests with dendritic hosts: an NMR study. J. Am. Chem. Soc. 127, 10334–10343 (2005).
Tomoyose, Y. et al. Aryl ether dendrimers with an interior metalloporphyrin functionality as a spectroscopic probe: Interpenetrating interaction with dendritic imidazoles. Macromolecules 29, 5236–5238 (1996).
Jiang, D.-L. & Aida, T. A dendritic iron porphyrin as a novel haemoprotein mimic: effects of the dendrimer cage on dioxygen-binding activity. Chem. Commun. 1523–1524 (1996).
Shinoda, S., Ohashi, M. & Tsukube, H. ‘Pocket dendrimers’ as nanoscale receptors for bimolecular guest accommodation. Chem. Eur. J. 13, 81–89 (2007).
Ong, W. et al. Electrochemical and guest binding properties of Fréchet- and Newkome-type dendrimers with a single viologen unit located at their apical positions. J. Am. Chem. Soc. 127, 3353–3361 (2005).
Balzani, V. et al. Host-guest complexes between an aromatic molecular tweezer and symmetric and unsymmetric dendrimers with a 4,4'-bipyridinium core. J. Am. Chem. Soc. 128, 637–648 (2006).
Ballauff, M. & Likos, C. N. Dendrimers in solution: insight from theory and simulation. Angew. Chem. Int. Ed. 43, 2998–3020 (2004).
Potschke, D., Ballauff, M., Lindner, P., Fischer, M. & Vögtle, F. Analysis of the structure of dendrimers in solution by small-angle neutron scattering including contrast variation. Macromolecules 32, 4079–4087 (1999).
Lim, J., Pavan, G. M., Annunziata, O. & Simanek, E. E. Experimental and computational evidence for an inversion in guest capacity in high-generation triazine dendrimer hosts. J. Am. Chem. Soc. 134, 1942–1945 (2012).
Posocco, P., Ferrone, M., Fermeglia, M. & Pricl, S. Binding at the core. Computational study of structural and ligand binding properties of naphthyridine-based dendrimers. Macromolecules 40, 2257–2266 (2007).
Zimmerman, S. C., Wang, Y., Bharathi, P. & Moore, J. S. Analysis of amidinium guest complexation by comparison of two classes of dendrimer hosts containing a hydrogen bonding unit at the core. J. Am. Chem. Soc. 120, 2172–2173 (1998).
Zimmerman, S. C., Wendland, M. S., Rakow, N. A., Zharov, I. & Suslick, K. S. Synthetic hosts by monomolecular imprinting inside dendrimers. Nature 418, 399–403 (2002).
Yamamoto, K. & Imaoka, T. Dendrimer complexes based on fine-controlled metal assembling. Bull. Chem. Soc. Jpn 79, 511–526 (2006).
Ochi, Y. et al. Controlled storage of ferrocene derivatives as redox-active molecules in dendrimers. J. Am. Chem. Soc. 132, 5061–5069 (2010).
Ochi, Y., Fujii, A., Nakajima, R. & Yamamoto, K. Stepwise radial complexation of triphenylmethyliums on a phenylazomethine dendrimer for organic−metal hybrid assembly. Macromolecules 43, 6570–6576 (2010).
Yamamoto, K. et al. Size-specific catalytic activity of platinum clusters enhances oxygen reduction reactions. Nat. Chem. 1, 397–402 (2009).
Imaoka, T., Ueda, H. & Yamamoto, K. Enhancing the photoelectric effect with a potential-programmed molecular rectifier. J. Am. Chem. Soc. 8412–8415 (2012).
Imaoka, T., Tanaka, R. & Yamamoto, K. Investigation of a molecular morphology effect on polyphenylazomethine dendrimers; physical properties and metal-assembling processes. Chem. Eur. J. 12, 7328–7336 (2006).
Fowler, C. J. et al. Metal complexes of porphycene, corrphycene, and hemiporphycene: stability and coordination chemistry. Chem. Eur. J. 8, 3485–3496 (2002).
Imai, H., Nakagawa, S. & Kyuno, E. Recognition of axial ligands by a zinc porphyrin host on the basis of nonpolar interligand interaction. J. Am. Chem. Soc. 114, 6719–6723 (1992).
Imaoka, T. et al. Probing stepwise complexation in phenylazomethine dendrimers by a metallo-porphyrin core. J. Am. Chem. Soc. 127, 13896–13905 (2005).
Matos, M. S. et al. Effect of core structure on photophysical and hydrodynamic properties of porphyrin dendrimers. Macromolecules 33, 2967–2973 (2000).
Hecht, S. & Fréchet, J. M. J. An alternative synthetic approach toward dendritic macromolecules: novel benzene-core dendrimers via alkyne cyclotrimerization. J. Am. Chem. Soc. 121, 4084–4085 (1999).
Uyemura, M. & Aida, T. Characteristics of organic transformations in a confined dendritic core: studies on the AIBN-initiated reaction of dendrimer cobalt(II) porphyrins with alkynes. Chem. Eur. J. 9, 3492–3500 (2003).
Higuchi, M., Shiki, S., Ariga, K. & Yamamoto, K. First synthesis of phenylazomethine dendrimer ligands and structural studies. J. Am. Chem. Soc. 123, 4414–4420 (2001).
Pinto, L. F., Correa, J., Martin-Pastor, M., Riguera, R. & Fernandez-Megia, E. The dynamics of dendrimers by NMR relaxation: interpretation pitfalls. J. Am. Chem. Soc. 135, 1972–1977 (2013).
Hu, J., Xu, T. & Cheng, Y. NMR insights into dendrimer-based host-guest systems. Chem. Rev. 112, 3856–3891 (2012).
Fukazawa, Y., Usui, S., Tanimoto, K. & Hirai, Y. Conformational-analysis by the ring current method—the structure of 2,2,13,13-tetramethyl[4.4]metacyclophane. J. Am. Chem. Soc. 116, 8169–8175 (1994).
Tárkányi, G., Kiráry, P., Varga, S., Vakulya, B. & Soós, T. Edge-to-face CH/pi aromatic interaction and molecular self-recognition in epi-cinchona-based bifunctional thiourea organocatalysis. Chem. Eur. J. 14, 6078–6086 (2008).
Meyer, E. A., Castellano, R. K. & Diederich, F. Interactions with aromatic rings in chemical and biological recognition. Angew. Chem. Int. Ed. 42, 1210–1250 (2003).
Iwamoto, H., Mizutani, T. & Kano, K. Thermodynamics of hydrophobic interactions: entropic recognition of a hydrophobic moiety by poly(ethylene oxide)-zinc porphyrin conjugates. Chem. Asian J. 2, 1267–1275 (2007).
Hu, W.-X., Li, P.-R., Jiang, G., Che, C. M. & Chen, J. A mild catalytic oxidation system: ruthenium porphyrin and 2,6-dichloropyridine N-oxide applied for alkene dihydroxylation. Adv. Synth. Catal. 352, 3190–3194 (2010).
Che, C.-M. & Huang, J.-S. Metalloporphyrin-based oxidation systems: from biomimetic reactions to application in organic synthesis. Chem. Commun. 3996–4015 (2009).
Groves, J. T. & Quinn, R. Aerobic epoxidation of olefins with ruthenium porphyrin catalysts. J. Am. Chem. Soc. 107, 5790–5792 (1985).
Ishikawa, A. & Sakaki, S. Theoretical study of photoinduced epoxidation of olefins catalyzed by ruthenium porphyrin. J. Phys. Chem. A 115, 4774–4785 (2011).
Zhang, J. L., Zhou, H. B., Huang, J.-S. & Che, C. M. Dendritic ruthenium porphyrins: a new class of highly selective catalysts for alkene epoxidation and cyclopropanation. Chem. Eur. J. 8, 1554–1562 (2002).
Frederick, K. K., Marlow, M. S., Valentine, K. G. & Wand, A. J. Conformational entropy in molecular recognition by proteins. Nature 448, 325–329 (2007).
Leung, D. H., Bergman, R. G. & Raymond, K. N. Enthalpy-entropy compensation reveals solvent reorganization as a driving force for supramolecular encapsulation in water. J. Am. Chem. Soc. 130, 2798–2805 (2008).
Williams, D. H., Calderone, C. T., O’Brien, D. P. & Zerella, R. Changes in motion versus bonding in positively versus negatively cooperative interactions. Chem. Commun. 1266–1267 (2002).
Albrecht, K., Kasai, Y., Kuramoto, Y. & Yamamoto, K. Dynamic control of dendrimer–fullerene association by axial coordination to the core. Chem. Commun. 49, 6861–6863 (2013).
Pollak, K. W., Leon, J. W., Fréchet, J. M. J., Maskus, M. & Abruña, H. D. Effects of dendrimer generation on site isolation of core moieties: electrochemical and fluorescence quenching studies with metalloporphyrin core dendrimers. Chem. Mater. 10, 30–38 (1998).
Acknowledgements
This work was supported in part by the CREST program of the Japan Science and Technology (JST) Agency and Grants-in-Aid for Scientific Research on Innovative Areas ‘Coordination Programming’ (area 2107, Number 21108009) from the Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Contributions
T.I., Y.K. and T.K. prepared the dendrimers and pyridine derivatives. T.I. and Y.K. carried out the titration experiments. T.K. carried out the catalytic epoxidation experiment. T.I. and K.Y. conceived experiments and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S30, Supplementary Tables S1-S4, Supplementary Methods and Supplementary References (PDF 2114 kb)
Rights and permissions
About this article
Cite this article
Imaoka, T., Kawana, Y., Kurokawa, T. et al. Macromolecular semi-rigid nanocavities for cooperative recognition of specific large molecular shapes. Nat Commun 4, 2581 (2013). https://doi.org/10.1038/ncomms3581
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms3581
This article is cited by
-
Functionalization of phenylazomethine dendrimers
Polymer Journal (2022)
-
Build-up enhancement of photoluminescence from phenylazomethine bismuth dendrimer using Bi(OTf)3
Journal of Nanoparticle Research (2018)
-
Metal Chelate Monomers as Precursors of Polymeric Materials
Journal of Inorganic and Organometallic Polymers and Materials (2016)
-
Self-organized nanotube materials and their application in bioengineering
Polymer Journal (2014)
-
The role of the host–guest interactions in the relative stability of compressed encapsulated homodimers and heterodimers of amides and carboxylic acids
Theoretical Chemistry Accounts (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.