Abstract
Many genetic liver diseases in newborns cause repeated, often lethal, metabolic crises. Gene therapy using nonintegrating viruses such as adeno-associated virus (AAV) is not optimal in this setting because the nonintegrating genome is lost as developing hepatocytes proliferate1,2. We reasoned that newborn liver may be an ideal setting for AAV-mediated gene correction using CRISPR-Cas9. Here we intravenously infuse two AAVs, one expressing Cas9 and the other expressing a guide RNA and the donor DNA, into newborn mice with a partial deficiency in the urea cycle disorder enzyme, ornithine transcarbamylase (OTC). This resulted in reversion of the mutation in 10% (6.7–20.1%) of hepatocytes and increased survival in mice challenged with a high-protein diet, which exacerbates disease. Gene correction in adult OTC-deficient mice was lower and accompanied by larger deletions that ablated residual expression from the endogenous OTC gene, leading to diminished protein tolerance and lethal hyperammonemia on a chow diet.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cunningham, S.C., Dane, A.P., Spinoulas, A., Logan, G.J. & Alexander, I.E. Gene delivery to the juvenile mouse liver using AAV2/8 vectors. Mol. Ther. 16, 1081–1088 (2008).
Wang, L., Wang, H., Bell, P., McMenamin, D. & Wilson, J.M. Hepatic gene transfer in neonatal mice by adeno-associated virus serotype 8 vector. Hum. Gene Ther. 23, 533–539 (2012).
Batshaw, M.L., Tuchman, M., Summar, M., Seminara, J. & Members of the Urea Cycle Disorders Consortium. A longitudinal study of urea cycle disorders. Mol. Genet. Metab. 113, 127–130 (2014).
Lichter-Konecki, U., Caldovic, L., Morizono, H. & Simpson, K. Ornithine transcarbamylase deficiency (1993–2013). in GeneReviews (eds. Pagon, R.A. et al.) http://www.ncbi.nlm.nih.gov/books/NBK154378/ (29 August 2013).
Ah Mew, N., Krivitzky, L., McCarter, R., Batshaw, M., Tuchman, M. & Urea Cycle Disorders Consortium of the Rare Diseases Clinical Research Network. Clinical outcomes of neonatal onset proximal versus distal urea cycle disorders do not differ. J. Pediatr. 162, 324–9.e1 (2013).
Hodges, P.E. & Rosenberg, L.E. The spfash mouse: a missense mutation in the ornithine transcarbamylase gene also causes aberrant mRNA splicing. Proc. Natl. Acad. Sci. USA 86, 4142–4146 (1989).
Sharma, R. et al. In vivo genome editing of the albumin locus as a platform for protein replacement therapy. Blood 126, 1777–1784 (2015).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Anguela, X.M. et al. Robust ZFN-mediated genome editing in adult hemophilic mice. Blood 122, 3283–3287 (2013).
Barzel, A. et al. Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature 517, 360–364 (2015).
Ran, F.A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Friedland, A.E. et al. Characterization of Staphylococcus aureus Cas9: a smaller Cas9 for all-in-one adeno-associated virus delivery and paired nickase applications. Genome Biol. 16, 257 (2015).
Kleinstiver, B.P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Doudna, J.A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Wang, D. et al. Adenovirus-mediated somatic genome editing of Pten by CRISPR/Cas9 in mouse liver in spite of Cas9-specific immune responses. Hum. Gene Ther. 26, 432–442 (2015).
Cheng, R. et al. Efficient gene editing in adult mouse livers via adenoviral delivery of CRISPR/Cas9. FEBS Lett. 588, 3954–3958 (2014).
Dingemanse, M.A. et al. Development of the ornithine cycle in rat liver: zonation of a metabolic pathway. Hepatology 24, 407–411 (1996).
Schmidt, F. & Grimm, D. CRISPR genome engineering and viral gene delivery: a case of mutual attraction. Biotechnol. J. 10, 258–272 (2015).
Vasileva, A. & Jessberger, R. Precise hit: adeno-associated virus in gene targeting. Nat. Rev. Microbiol. 3, 837–847 (2005).
Mahiny, A.J. et al. In vivo genome editing using nuclease-encoding mRNA corrects SP-B deficiency. Nat. Biotechnol. 33, 584–586 (2015).
Gao, G.P. et al. Novel adeno-associated viruses from rhesus monkeys as vectors for human gene therapy. Proc. Natl. Acad. Sci. USA 99, 11854–11859 (2002).
Wang, L. et al. AAV8-mediated hepatic gene transfer in infant rhesus monkeys (Macaca mulatta). Mol. Ther. 19, 2012–2020 (2011).
Yang, L. et al. Targeted and genome-wide sequencing reveal single nucleotide variations impacting specificity of Cas9 in human stem cells. Nat. Commun. 5, 5507 (2014).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Shi, Y., Falahati, R., Zhang, J., Flebbe-Rehwaldt, L. & Gaensler, K.M. Role of antigen-specific regulatory CD4+CD25+ T cells in tolerance induction after neonatal IP administration of AAV-hF.IX. Gene Ther. 20, 987–996 (2013).
Nivsarkar, M.S. et al. Evidence for contribution of CD4+ CD25+ regulatory T cells in maintaining immune tolerance to human factor IX following perinatal adenovirus vector delivery. J. Immunol. Res. 2015, 397879 (2015).
LoDuca, P.A., Hoffman, B.E. & Herzog, R.W. Hepatic gene transfer as a means of tolerance induction to transgene products. Curr. Gene Ther. 9, 104–114 (2009).
Hinderer, C. et al. Neonatal systemic AAV induces tolerance to CNS gene therapy in MPS I dogs and nonhuman primates. Mol. Ther. 23, 1298–1307 (2015).
Lock, M. et al. Rapid, simple, and versatile manufacturing of recombinant adeno-associated viral vectors at scale. Hum. Gene Ther. 21, 1259–1271 (2010).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Moscioni, D. et al. Long-term correction of ammonia metabolism and prolonged survival in ornithine transcarbamylase-deficient mice following liver-directed treatment with adeno-associated viral vectors. Mol. Ther. 14, 25–33 (2006).
Daly, T.M. AAV-mediated gene transfer to the liver. Methods Mol. Biol. 246, 195–199 (2004).
Augustin, L., Mavinakere, M., Morizono, H. & Tuchman, M. Expression of wild-type and mutant human ornithine transcarbamylase genes in Chinese hamster ovary cells and lack of dominant negative effect of R141Q and R40H mutants. Pediatr. Res. 48, 842–846 (2000).
Morizono, H. et al. Expression, purification and kinetic characterization of wild-type human ornithine transcarbamylase and a recurrent mutant that produces 'late onset' hyperammonaemia. Biochem. J. 322, 625–631 (1997).
Wang, L. et al. Sustained correction of OTC deficiency in spf(ash) mice using optimized self-complementary AAV2/8 vectors. Gene Ther. 19, 404–410 (2012).
Acknowledgements
We thank Penn Vector Core for supplying vectors, Penn Bioinformatics Core for assistance on deep sequencing data analysis, Y. Zhu and M. Nayalat for help on immunohistochemistry analysis, and L. Mays for assistance on manuscript preparation. This work was supported by National Institute of Child Health and Human Development P01-HD057247 (J.M.W.) and the Kettering Family Foundation (M.L.B.).
Author information
Authors and Affiliations
Contributions
L.W. and J.M.W. conceived this study. L.W., Y.Y. and J.M.W. designed the experiments. Y.Y., P.B., D.M., Z.H., J.W., H.Y. and C.X. performed the experiments. K.M. conducted the bioinformatics analysis of the deep sequencing data. J.M.W., L.W., Y.Y., P.B., H.M., K.M. and M.L.B. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.M. Wilson is an advisor to REGENXBIO, Dimension Therapeutics and Solid Gene Therapy, and is a founder of, holds equity in, and has a sponsored research agreement with REGENXBIO and Dimension Therapeutics; in addition, he is a consultant to several biopharmaceutical companies and is an inventor on patents licensed to various biopharmaceutical companies.
Integrated supplementary information
Supplementary Figure 1 In vitro validation of OTC sgRNAs and donor template.
(a) In vitro validation of sgRNAs targeted to OTC in the MC57G mouse cell line by transient transfection followed by 4-day puromycin enrichment and SURVEYOR nuclease assays. sgRNA1 showed the highest efficiency in inducing indels in the targeted loci and was therefore chosen for subsequent studies. Arrows denote SURVEYOR nuclease cleaved fragments of the OTC PCR products. Results were replicated in 2 independent experiments. (b) In vitro validation of OTC donor template. MC57G cells were transiently transfected with a plasmid co-expressing OTC sgRNA1, SaCas9, and an AgeI restriction site tagged OTC donor plasmid followed by 4-day puromycin enrichment. RFLP analysis was performed following AgeI digestion to detect HDR in vitro. Co-transfection of the AgeI-tagged OTC donor template with an SaCas9 plasmid without OTC sgRNA1 did not result in detectable HDR. Arrows denote AgeI-sensitive cleavage products resulting from HDR. Results were replicated in 2 independent experiments. Indel and HDR frequency were calculated based on band intensities31.
Supplementary Figure 2 Vector dose optimization to improve in vivo gene correction.
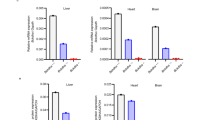
Postnatal day 2 spfash pups received temporal vein injection of 5x1010 GC AAV8.SaCas9 and either 5x1010 (n=5), 1x1011 (n=3), or 5x1011 (n=5) GC of AAV8.sgRNA1.donor vector. Liver samples were collected 3 weeks post vector treatment for analysis. (a) Quantification of gene correction based on the percentage of area on liver sections expressing OTC by immunostaining. (b) Quantification of OTC mRNA levels in the liver by RT-qPCR using primers spanning exons 4–5 to amplify wild-type OTC. Mean ± SEM are shown. ** P<0.01, Dunnett’s test.
Supplementary Figure 3 Time course of gene expression by Western analysis and HDR analysis by RFLP.
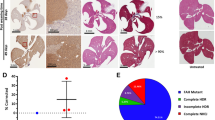
(a) HDR analysis by RFLP. OTC target region was PCR amplified from the liver genomic DNA isolated from untreated spfash mice or spfash mice treated with the dual AAV vectors. Untreated spfash control samples were collected at 8 weeks of age. Samples from the treated spfash mice were collected at 1, 3, and 8 weeks (n=3 animals per time point) following neonatal injection of the dual AAV8 vectors. Targeted animals received AAV8.SaCas9 (5x1010 GC/pup) and AAV8.sgRNA1.donor (5x1011 GC/pup). Untargeted animals received AAV8.SaCas9 (5x1010 GC/pup) and AAV8.control.donor (5x1011 GC/pup). AgeI digestion was performed and estimated HDR percentages are shown. (b) Western blot analysis. Liver lysates were prepared from untreated WT and spfash mice or spfash mice treated with the dual AAV vectors for detection of FLAG-SaCas9 and OTC protein.
Supplementary Figure 4 Examination of liver toxicity in animals treated with AAV8.CRISPR-SaCas9 dual vectors.
(a) Histological analysis on livers harvested 3 and 8 weeks following the dual vector treatment. Scale bar, 100 µm. (b) Liver transaminase levels in untreated spfash mice (n=9) or 8 weeks following dual vector treatment. Untargeted mice received 5x1010 GC AAV8.SaCas9 and 5x1011 of AAV8.control.donor vectors (n=8), while gene-targeted mice received 5x1010 GC AAV8.SaCas9 and 5x1011 GC of AAV8.sgRNA1.donor (n=7). Mean ± SEM are shown. There were no statistically significant differences between groups, Dunnett’s test.
Supplementary Figure 5 Examination of liver toxicity in adult animals treated with AAV8.CRISPR-SaCas9 dual vectors.
(a) Histological analysis on livers harvested 3 weeks (low-dose) or 2 weeks (high-dose) following dual vector treatment. Scale bar, 100 µm. (b) Liver transaminase levels in untreated spfash mice, or 3 weeks following low-dose dual vector treatment, or 2 weeks following high-dose dual vector treatment (n=3 for each group). Low-dose, untargeted mice received 1x1011 GC AAV8.SaCas9 and 1x1012 GC of AAV8.control.donor vectors, while low-dose, gene-targeted mice received 1x1011 GC AAV8.SaCas9 and 1x1012 GC of AAV8.sgRNA1.donor. High-dose, untargeted mice received 1x1012 GC AAV8.SaCas9 and 5x1012 GC of AAV8.control.donor vectors, while high-dose, gene-targeted mice received 1x1012 GC AAV8.SaCas9 and 5x1012 GC of AAV8.sgRNA1.donor. Mean ± SEM are shown. Adult animals received high-dose, gene-targeted vectors showed a trend of elevated ALT and AST levels, although not statistically different when compared with other groups (Dunnett’s test).
Supplementary Figure 6 Comparison of SaCas9 vector DNA and mRNA levels in the livers of neonatal treated and adult treated mice.
Neonatal spfash mice received 5x1010 GC AAV8.SaCas9 and 5x1011 GC of AAV8.sgRNA1.donor vectors and were sacrificed at 1 (n=5), 3 (n=6), and 8 weeks (n=7) after injection. Low-dose adult spfash mice received 1x1011 GC AAV8.SaCas9 and 1x1012 GC of AAV8.sgRNA1.donor vectors and were sacrificed at 3 weeks (n=3) after injection. High-dose adult spfash mice received 1x1012 GC AAV8.SaCas9 and 5x1012 GC of AAV8.sgRNA1.donor vectors and were sacrificed at 2 weeks (n=3) after injection. (a) Quantification of SaCas9 vector DNA in the liver by qPCR. (b) Quantification of SaCas9 mRNA in the liver by RT-qPCR. Mean ± SEM are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1 and 3–6 (PDF 916 kb)
Rights and permissions
About this article
Cite this article
Yang, Y., Wang, L., Bell, P. et al. A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice. Nat Biotechnol 34, 334–338 (2016). https://doi.org/10.1038/nbt.3469
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3469
This article is cited by
-
Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos
Nature Biotechnology (2024)
-
Utilizing AAV-mediated LEAPER 2.0 for programmable RNA editing in non-human primates and nonsense mutation correction in humanized Hurler syndrome mice
Genome Biology (2023)
-
In vivo adenine base editing corrects newborn murine model of Hurler syndrome
Molecular Biomedicine (2023)
-
Homology-directed repair of an MYBPC3 gene mutation in a rat model of hypertrophic cardiomyopathy
Gene Therapy (2023)
-
A positive feedback between cholesterol synthesis and the pentose phosphate pathway rather than glycolysis promotes hepatocellular carcinoma
Oncogene (2023)