Abstract
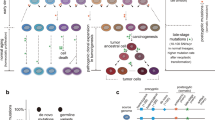
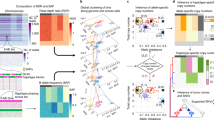
Prolonged culture of human embryonic stem cells (hESCs) can lead to adaptation and the acquisition of chromosomal abnormalities, underscoring the need for rigorous genetic analysis of these cells. Here we report the highest-resolution study of hESCs to date using an Affymetrix SNP 6.0 array containing 906,600 probes for single nucleotide polymorphisms (SNPs) and 946,000 probes for copy number variations (CNVs). Analysis of 17 different hESC lines maintained in different laboratories identified 843 CNVs of 50 kb–3 Mb in size. We identified, on average, 24% of the loss of heterozygosity (LOH) sites and 66% of the CNVs changed in culture between early and late passages of the same lines. Thirty percent of the genes detected within CNV sites had altered expression compared to samples with normal copy number states, of which >44% were functionally linked to cancer. Furthermore, LOH of the q arm of chromosome 16, which has not been observed previously in hESCs, was detected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Draper, J.S., Moore, H.D., Ruban, L.N., Gokhale, P.J. & Andrews, P.W. Culture and characterization of human embryonic stem cells. Stem Cells Dev. 13, 325–336 (2004).
Draper, J.S. et al. Recurrent gain of chromosomes 17q and 12 in cultured human embryonic stem cells. Nat. Biotechnol. 22, 53–54 (2004).
Hanson, C. & Caisander, G. Human embryonic stem cells and chromosome stability. APMIS 113, 751–755 (2005).
Enver, T. et al. Cellular differentiation hierarchies in normal and culture-adapted human embryonic stem cells. Hum. Mol. Genet. 14, 3129–3140 (2005).
Baker, D.E. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nat. Biotechnol. 25, 207–215 (2007).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Feuk, L., Carson, A.R. & Scherer, S.W. Structural variation in the human genome. Nat. Rev. Genet. 7, 85–97 (2006).
Iafrate, A.J. et al. Detection of large-scale variation in the human genome. Nat. Genet. 36, 949–951 (2004).
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Kallioniemi, A. CGH microarrays and cancer. Curr. Opin. Biotechnol. 19, 36–40 (2008).
Jong, K. et al. Cross-platform array comparative genomic hybridization meta-analysis separates hematopoietic and mesenchymal from epithelial tumors. Oncogene 26, 1499–1506 (2007).
Zheng, H.T., Peng, Z.H., Li, S. & He, L. Loss of heterozygosity analyzed by single nucleotide polymorphism array in cancer. World J. Gastroenterol. 11, 6740–6744 (2005).
Cervantes, R.B., Stringer, J.R., Shao, C., Tischfield, J.A. & Stambrook, P.J. Embryonic stem cells and somatic cells differ in mutation frequency and type. Proc. Natl. Acad. Sci. USA 99, 3586–3590 (2002).
Donahue, S.L., Lin, Q., Cao, S. & Ruley, H.E. Carcinogens induce genome-wide loss of heterozygosity in normal stem cells without persistent chromosomal instability. Proc. Natl. Acad. Sci. USA 103, 11642–11646 (2006).
Inzunza, J. et al. Comparative genomic hybridization and karyotyping of human embryonic stem cells reveals the occurrence of an isodicentric X chromosome after long-term cultivation. Mol. Hum. Reprod. 10, 461–466 (2004).
Maitra, A. et al. Genomic alterations in cultured human embryonic stem cells. Nat. Genet. 37, 1099–1103 (2005).
Caisander, G. et al. Chromosomal integrity maintained in five human embryonic stem cell lines after prolonged in vitro culture. Chromosome Res. 14, 131–137 (2006).
Wu, H. et al. Copy number variant analysis of human embryonic stem cells. Stem Cells 26, 1484–1489 (2008).
Spits, C. et al. Recurrent chromosomal abnormalities in human embryonic stem cells. Nat. Biotechnol. 12, 1361–1363 (2008).
Hubbard, T.J. et al. Ensembl 2007. Nucleic Acids Res. 35, D610–D617 (2007).
Monk, M., Hitchins, M. & Hawes, S. Differential expression of the embryo/cancer gene ECSA(DPPA2), the cancer/testis gene BORIS and the pluripotency structural gene OCT4, in human preimplantation development. Mol. Hum. Reprod. 14, 347–355 (2008).
Lindblom, A., Rotstein, S., Skoog, L., Nordenskjold, M. & Larsson, C. Deletions on chromosome 16 in primary familial breast carcinomas are associated with development of distant metastases. Cancer Res. 53, 3707–3711 (1993).
Cleton-Jansen, A.M. et al. Different mechanisms of chromosome 16 loss of heterozygosity in well- versus poorly differentiated ductal breast cancer. Genes Chromosom. Cancer 41, 109–116 (2004).
Carter, B.S. et al. Allelic loss of chromosomes 16q and 10q in human prostate cancer. Proc. Natl. Acad. Sci. USA 87, 8751–8755 (1990).
Jenner, M.W. et al. Gene mapping and expression analysis of 16q loss of heterozygosity identifies WWOX and CYLD as being important in determining clinical outcome in multiple myeloma. Blood 110, 3291–3300 (2007).
Mortensen, R.M., Conner, D.A., Chao, S., Geisterfer-Lowrance, A.A. & Seidman, J.G. Production of homozygous mutant ES cells with a single targeting construct. Mol. Cell. Biol. 12, 2391–2395 (1992).
Lefort, N. et al. Human embryonic stem cells reveal recurrent genomic instability at 20q11.21. Nat. Biotechnol. 26, 1364–1366 (2008).
Mantel, C. et al. Checkpoint-apoptosis uncoupling in human and mouse embryonic stem cells: a source of karyotpic instability. Blood 109, 4518–4527 (2007).
Rodriguez-Jimenez, F.J., Moreno-Manzano, V., Lucas-Dominguez, R. & Sanchez-Puelles, J.M. Hypoxia causes downregulation of mismatch repair system and genomic instability in stem cells. Stem Cells 26, 2052–2062 (2008).
Garcia-Perez, J.L. et al. LINE-1 retrotransposition in human embryonic stem cells. Hum. Mol. Genet. 16, 1569–1577 (2007).
Hastings, P.J. Adaptive amplification. Crit. Rev. Biochem. Mol. Biol. 42, 271–283 (2007).
Osafune, K. et al. Marked differences in differentiation propensity among human embryonic stem cell lines. Nat. Biotechnol. 26, 313–315 (2008).
Andrews, P.W. et al. Embryonic stem (ES) cells and embryonal carcinoma (EC) cells: opposite sides of the same coin. Biochem. Soc. Trans. 33, 1526–1530 (2005).
The International HapMap Consortium The international HapMap project. Nature 426, 789–796 (2003).
Eyre, T.A. et al. The HUGO gene nomenclature database, 2006 updates. Nucleic Acids Res. 34, D319–D321 (2006).
Dai, M. et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res. 33, e175 (2005).
Bengtsson, H., Simpson, K., Bullard, J. & Hansen, K. . Aroma.Affymetrix: A Generic Framework In R For Analyzing Small To Very Large Affymetrix Data Sets In Bounded Memory. Technical report 745. (Department of Statistics, University of California, Berkeley, 2008).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Hautaniemi, S. et al. A strategy for identifying putative causes of gene expression variation in human cancers. J. Franklin Inst. 341, 77–88 (2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., B 57, 289–300 (1995).
Jarvinen, A.K. et al. Identification of target genes in laryngeal squamous cell carcinoma by high-resolution copy number and gene expression microarray analyses. Oncogene 25, 6997–7008 (2006).
Acknowledgements
We are grateful to everyone who has taken care of sample collection and handling: T. Golan-Lev, A. Urrutikoetxea-Uriguen, S. Haupt, P. Koch, I. Laufenberg, B. Ley, A. Hampl, M. Vodinska, K. Koudelkova, S. Ström, F. Holm, A.-M. Strömberg, C. Olsson, M. Mikkola, S. Vuoristo, P. Junni and M. Hakkarainen. We especially acknowledge M. Linja, T. Heinonen and the Finnish DNA Microarray Centre for their excellent technical assistance. We acknowledge the Turku Graduate School of Biomedical Sciences. This study is supported by funding for the ESTOOLS consortium under the Sixth Research Framework Programme of the European Union, Juvenile Diabetes Research Foundation, The Academy of Finland and the Finnish Cancer Organizations, The Improving Outcomes Guidance Trust, The Ministry of Education, Youth, and Sport of the Czech Republic, Ida Montin Foundation, The Academy of Finland, projects no. 129657 (Finnish Centre of Excellence program 2006-11) and no. 134117 and the Medical Research Council, UK.
Author information
Authors and Affiliations
Contributions
E.N., R.A., N.B., P.W.A., O.Y.-H. and R.L. designed the experiments, E.N. and R.L. were responsible for the coordination of the project and microarray experiments. R.A., E.N. and O.Y.-H. were responsible for data analysis, integration and statistical analysis. N.R. performed RNA extractions. L.K. built the gene annotation list of genes overlapping CNVs. D.B. performed conventional karyotyping. E.N. and N.R. performed copy-number state validations with RT-PCR. J.I.-E. provided I3 and I6 lines for the study. P.D., O.H., T.O., T.T., N.B., W.C., O.B., E.M., H.D.M., P.W.A., O.Y.-H. and R.L. provided the samples and coordinated the project in their groups. E.N., R.A., N.R., L.K., N.H., D.K., L.B., J.I.-E., O.R., P.D., O.H., T.O., T.T., N.B., W.C., O.B., D.B., E.M., H.D.M., P.W.A., O.Y.-H. and R.L. contributed to writing the paper.
Corresponding authors
Ethics declarations
Competing interests
D.K. is affiliated with Stem Cell Technologies, Ltd. (However, the study was not supported by the company.)
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–4 and Supplementary Tables 4,5,6,12 (PDF 332 kb)
Supplementary Table 1.
SNP profiles and Hapmap codes.xls (XLS 89 kb)
Supplementary Table 2.
CNV region list (XLS 952 kb)
Supplementary Table 3.
HapMap CNV region list (XLS 2410 kb)
Supplementary Table 7.
Genes affected by CNVs HapMap (XLS 1037 kb)
Supplementary Table 8.
Genes affected by CNVs (XLS 236 kb)
Supplementary Table 9.
genes changed by adaptation (XLS 57 kb)
Supplementary Table 10a.
integrated analysis, losses (XLS 169 kb)
Supplementary Table 10b.
integrated analysis, gains (XLS 4038 kb)
Supplementary Table 11.
Culture conditions (XLS 20 kb)
Rights and permissions
About this article
Cite this article
Närvä, E., Autio, R., Rahkonen, N. et al. High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity. Nat Biotechnol 28, 371–377 (2010). https://doi.org/10.1038/nbt.1615
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1615
This article is cited by
-
The Regenerative Microenvironment of the Tissue Engineering for Urethral Strictures
Stem Cell Reviews and Reports (2024)
-
Xeno-free generation of human induced pluripotent stem cells from donor-matched fibroblasts isolated from dermal and oral tissues
Stem Cell Research & Therapy (2023)
-
TPX2 Amplification-Driven Aberrant Mitosis in Culture Adapted Human Embryonic Stem Cells with gain of 20q11.21
Stem Cell Reviews and Reports (2023)
-
Passage number affects differentiation of sensory neurons from human induced pluripotent stem cells
Scientific Reports (2022)
-
Functional in vivo and in vitro effects of 20q11.21 genetic aberrations on hPSC differentiation
Scientific Reports (2020)