Abstract
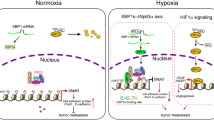
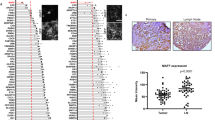
The molecular determinants of malignant cell behaviours in breast cancer remain only partially understood1. Here we show that SHARP1 (also known as BHLHE41 or DEC2) is a crucial regulator of the invasive and metastatic phenotype in triple-negative breast cancer (TNBC), one of the most aggressive types of breast cancer. SHARP1 is regulated by the p63 metastasis suppressor and inhibits TNBC aggressiveness through inhibition of hypoxia-inducible factor 1α (HIF-1α) and HIF-2α (HIFs). SHARP1 opposes HIF-dependent TNBC cell migration in vitro, and invasive or metastatic behaviours in vivo. SHARP1 is required, and sufficient, to limit expression of HIF-target genes. In primary TNBC, endogenous SHARP1 levels are inversely correlated with those of HIF targets. Mechanistically, SHARP1 binds to HIFs and promotes HIF proteasomal degradation by serving as the HIF-presenting factor to the proteasome. This process is independent of pVHL (von Hippel–Lindau tumour suppressor), hypoxia and the ubiquitination machinery. SHARP1 therefore determines the intrinsic instability of HIF proteins to act in parallel to, and cooperate with, oxygen levels. This work sheds light on the mechanisms and pathways by which TNBC acquires invasiveness and metastatic propensity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elias, A. D. Triple-negative breast cancer: a short review. Am. J. Clin. Oncol. 33, 637–645 (2010)
Di Cosimo, S. & Baselga, J. Management of breast cancer with targeted agents: importance of heterogeneity. Nature Rev. Clin. Oncol. 7, 139–147 (2010)
Adorno, M. et al. A Mutant-p53/Smad complex opposes p63 to empower TGFβ-induced metastasis. Cell 137, 87–98 (2009)
Su, X. et al. TAp63 suppresses metastasis through coordinate regulation of Dicer and miRNAs. Nature 467, 986–990 (2010)
Muller, P. A. et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 139, 1327–1341 (2009)
Gordan, J. D. & Simon, M. C. Hypoxia-inducible factors: central regulators of the tumor phenotype. Curr. Opin. Genet. Dev. 17, 71–77 (2007)
Kaelin, W. G., Jr & Ratcliffe, P. J. Oxygen sensing by metazoans: the central role of the HIF hydroxylase pathway. Mol. Cell 30, 393–402 (2008)
Semenza, G. L. Defining the role of hypoxia-inducible factor 1 in cancer biology and therapeutics. Oncogene 29, 625–634 (2010)
Helczynska, K. et al. Hypoxia-inducible factor-2α correlates to distant recurrence and poor outcome in invasive breast cancer. Cancer Res. 68, 9212–9220 (2008)
Zhang, H. et al. HIF-1-dependent expression of angiopoietin-like 4 and L1CAM mediates vascular metastasis of hypoxic breast cancer cells to the lungs. Oncogene 31, 1757–1770 (2011)
Lu, X. et al. In vivo dynamics and distinct functions of hypoxia in primary tumor growth and organotropic metastasis of breast cancer. Cancer Res. 70, 3905–3914 (2010)
Sato, F. et al. Basic-helix-loop-helix (bHLH) transcription factor DEC2 negatively regulates vascular endothelial growth factor expression. Genes Cells 13, 131–144 (2008)
Padua, D. et al. TGFβ primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 133, 66–77 (2008)
Wong, C. C. et al. Hypoxia-inducible factor 1 is a master regulator of breast cancer metastatic niche formation. Proc. Natl Acad. Sci. USA 108, 16369–16374 (2011)
Buffa, F. M. et al. microRNA-associated progression pathways and potential therapeutic targets identified by integrated mRNA and microRNA expression profiling in breast cancer. Cancer Res. 71, 5635–5645 (2011)
Vleugel, M. M. et al. Differential prognostic impact of hypoxia induced and diffuse HIF-1α expression in invasive breast cancer. J. Clin. Pathol. 58, 172–177 (2005)
Girardini, J. E. et al. A Pin1/mutant p53 axis promotes aggressiveness in breast cancer. Cancer Cell 20, 79–91 (2011)
Strano, S. et al. Physical interaction with human tumor-derived p53 mutants inhibits p63 activities. J. Biol. Chem. 277, 18817–18826 (2002)
Uchida, T. et al. Prolonged hypoxia differentially regulates hypoxia-inducible factor (HIF)-1α and HIF-2α expression in lung epithelial cells: implication of natural antisense HIF-1alpha. J. Biol. Chem. 279, 14871–14878 (2004)
Kong, X., Alvarez-Castelao, B., Lin, Z., Castano, J. G. & Caro, J. Constitutive/hypoxic degradation of HIF-α proteins by the proteasome is independent of von Hippel Lindau protein ubiquitylation and the transactivation activity of the protein. J. Biol. Chem. 282, 15498–15505 (2007)
Sdek, P. et al. MDM2 promotes proteasome-dependent ubiquitin-independent degradation of retinoblastoma protein. Mol. Cell 20, 699–708 (2005)
Asher, G., Tsvetkov, P., Kahana, C. & Shaul, Y. A mechanism of ubiquitin-independent proteasomal degradation of the tumor suppressors p53 and p73. Genes Dev. 19, 316–321 (2005)
Cho, S. et al. Binding and regulation of HIF-1α by a subunit of the proteasome complex, PSMA7. FEBS Lett. 498, 62–66 (2001)
Chowdary, D. R., Dermody, J. J., Jha, K. K. & Ozer, H. L. Accumulation of p53 in a mutant cell line defective in the ubiquitin pathway. Mol. Cell. Biol. 14, 1997–2003 (1994)
Acknowledgements
We thank O. Wessely, G. Del Sal, C. Oakman and A. Di Leo for comments; and W. Kaelin, C. Borner, S. Libutti, Y. Maeda and Y. Kato for gifts of reagents. M.M. is a recipient of a FIRC (Federazione Italiana Ricerca Cancro) fellowship. We are in debt to E. Tagliafico and the Modena Affimetrix platform team for help with microarrays. E.E. is a recipient of a Cariparo PhD fellowship. M.M. was a recipient of an AIRC (Italian Association for Cancer Research) fellowship. This work is supported by a Young Italian Researchers grant of the Italian Welfare Ministry and an AIRC (Associazione Italiana per la Ricerca sul Cancro) MFAG grant to M.C.; a Fondazione Città della Speranza Grant, and MIUR (Ministero dell'Istruzione dell'Università e della Ricerca Italia) and PRIN grants to G.B; and an AIRC Principal Investigator grant, an AIRC Special Program Molecular Clinical Oncology ‘5 per mille’ grant, an HSFP grant, a University of Padua Strategic grant, an IIT Excellence grant, a CNR–Miur Epigenetics Flagship project, and a Comitato Promotore Telethon grant to S.P.
Author information
Authors and Affiliations
Contributions
M.M., M.C. and S.P. designed research, and M.M., E.E, M.F., M.C., E.R., G.B., G.L. and F.Z. performed experiments. M.C., S.B. and M.F. performed bioinformatics analysis, and A.R. and A.P. helped with assays in mice and tumour pathology. M.M., M.C. and S.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information 1
This file contains Supplementary Figures 1-29, Supplementary Methods, Supplementary Tables 1-2, 4, and 6-11 (see separate files for Supplementary Tables 3 and 5) and Supplementary references. (PDF 15593 kb)
Supplementary Table 3
This file contains a table with the Characteristics of the 250 TNBC samples. (XLS 92 kb)
Supplementary Table 5
This file contains a table with the Genes identified by SAM as up-regulated in Sharp1/CyclinG2 Low TNBCs compared to Sharp1/CyclinG2 High TNBCs. (XLS 550 kb)
Supplementary Information 2
This file contains the Sweave document in .pdf format for the analysis of gene expression data from MDA-MB-231 cells depleted of HIF-1α and HIF-2α (MDA-shHIF) and MDA-MB-231 cells overexpressing Sharp1 (MDA-Sharp1). (PDF 272 kb)
Rights and permissions
About this article
Cite this article
Montagner, M., Enzo, E., Forcato, M. et al. SHARP1 suppresses breast cancer metastasis by promoting degradation of hypoxia-inducible factors. Nature 487, 380–384 (2012). https://doi.org/10.1038/nature11207
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11207
This article is cited by
-
PIN1 and CDK1 cooperatively govern pVHL stability and suppressive functions
Cell Death & Differentiation (2023)
-
SCFβTrCP-mediated degradation of SHARP1 in triple-negative breast cancer
Cell Death & Disease (2023)
-
Discovery of differentially expressed novel miRNAs in breast normal cells and their putative targets
Molecular and Cellular Biochemistry (2023)
-
Landscape of NcRNAs involved in drug resistance of breast cancer
Clinical and Translational Oncology (2023)
-
A signature of six-hypoxia-related genes to evaluate the tumor immune microenvironment and predict prognosis in gastric cancer
BMC Medical Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.