Abstract
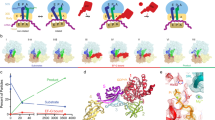
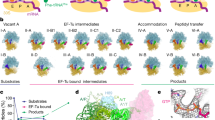
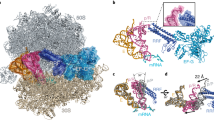
The elongation cycle of protein synthesis involves the delivery of aminoacyl-transfer RNAs to the aminoacyl-tRNA-binding site (A site) of the ribosome, followed by peptide-bond formation and translocation of the tRNAs through the ribosome to reopen the A site1,2. The translocation reaction is catalysed by elongation factor G (EF-G) in a GTP-dependent manner3. Despite the availability of structures of various EF-G–ribosome complexes, the precise mechanism by which tRNAs move through the ribosome still remains unclear. Here we use multiparticle cryoelectron microscopy analysis to resolve two previously unseen subpopulations within Thermus thermophilus EF-G–ribosome complexes at subnanometre resolution, one of them with a partly translocated tRNA. Comparison of these substates reveals that translocation of tRNA on the 30S subunit parallels the swivelling of the 30S head and is coupled to unratcheting of the 30S body. Because the tRNA maintains contact with the peptidyl-tRNA-binding site (P site) on the 30S head and simultaneously establishes interaction with the exit site (E site) on the 30S platform, a novel intra-subunit ‘pe/E’ hybrid state is formed. This state is stabilized by domain IV of EF-G, which interacts with the swivelled 30S-head conformation. These findings provide direct structural and mechanistic insight into the ‘missing link’ in terms of tRNA intermediates involved in the universally conserved translocation process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
EMBL/GenBank/DDBJ
Protein Data Bank
Data deposits
The electron density maps and models of the TIPRE and the TIPOST complexes have been deposited in the 3D-EM database with accession numbers EMD-1798 and EMD-1799, and in the Protein Data Bank database with PDB IDs 2xsy, 2xtg, 2xux and 2xuy.
References
Frank, J. & Spahn, C. M. The ribosome and the mechanism of protein synthesis. Rep. Prog. Phys. 69, 1383–1417 (2006)
Schmeing, T. M. & Ramakrishnan, V. What recent ribosome structures have revealed about the mechanism of translation. Nature 461, 1234–1242 (2009)
Shoji, S., Walker, S. E. & Fredrick, K. Ribosomal translocation: one step closer to the molecular mechanism. ACS Chem. Biol. 4, 93–107 (2009)
Ramakrishnan, V. Ribosome structure and the mechanism of translation. Cell 108, 557–572 (2002)
Moazed, D. & Noller, H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989)
Munro, J. B., Altman, R. B., O’Connor, N. & Blanchard, S. C. Identification of two distinct hybrid state intermediates on the ribosome. Mol. Cell 25, 505–517 (2007)
Blanchard, S. C. et al. tRNA dynamics on the ribosome during translation. Proc. Natl Acad. Sci. USA 101, 12893–12898 (2004)
Munro, J. B., Sanbonmatsu, K. Y., Spahn, C. M. & Blanchard, S. C. Navigating the ribosome’s metastable energy landscape. Trends Biochem. Sci. 34, 390–400 (2009)
Agirrezabala, X. et al. Visualization of the hybrid state of tRNA binding promoted by spontaneous ratcheting of the ribosome. Mol. Cell 32, 190–197 (2008)
Julian, P. et al. Structure of ratcheted ribosomes with tRNAs in hybrid states. Proc. Natl Acad. Sci. USA 105, 16924–16927 (2008)
Fischer, N. et al. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010)
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003)
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000)
Connell, S. R. et al. Structural basis for interaction of the ribosome with the switch regions of GTP-bound elongation factors. Mol. Cell 25, 751–764 (2007)
Spahn, C. M. et al. Domain movements of elongation factor eEF2 and the eukaryotic 80S ribosome facilitate tRNA translocation. EMBO J. 23, 1008–1019 (2004)
Zhang, W., Dunkle, J. A. & Cate, J. H. Structures of the ribosome in intermediate states of ratcheting. Science 325, 1014–1017 (2009)
Rodnina, M., Savelsbergh, A., Katunin, V. I. & Wintermeyer, W. Hydrolysis of GTP by elongation factor G drives tRNA movement on the ribosome. Nature 385, 37–41 (1997)
Savelsbergh, A. et al. An elongation factor G-induced ribosome rearrangement precedes tRNA-mRNA translocation. Mol. Cell 11, 1517–1523 (2003)
Gao, Y. G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009)
Penczek, P. A., Frank, J. & Spahn, C. M. A method of focused classification, based on the bootstrap 3D variance analysis, and its application to EF-G-dependent translocation. J. Struct. Biol. 154, 184–194 (2006)
Agrawal, R. K. et al. EF-G-dependent GTP hydrolysis induces translocation accompanied by large conformational changes in the 70S ribosome. Nature Struct. Biol. 6, 643–647 (1999)
Scheres, S. H. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nature Methods 4, 27–29 (2007)
Matassova, A. B., Rodnina, M. V. & Wintermeyer, W. Elongation factor G-induced structural change in helix 34 of 16S rRNA related to translocation on the ribosome. RNA 7, 1879–1885 (2001)
Ticu, C. et al. Conformational changes in switch I of EF-G drive its directional cycling on and off the ribosome. EMBO J. 28, 2053–2065 (2009)
Spirin, A. S. The ribosome as a conveying thermal ratchet machine. J. Biol. Chem. 284, 21103–21119 (2009)
Schuette, J. C. et al. GTPase activation of elongation factor EF-Tu by the ribosome during decoding. EMBO J. 28, 755–765 (2009)
Whitford, P. C. et al. Accommodation of aminoacyl-tRNA into the ribosome involves reversible excursions along multiple pathways. RNA 16, 1196–1204 (2010)
Sharma, M. R. et al. Interaction of Era with the 30S ribosomal subunit implications for 30S subunit assembly. Mol. Cell 18, 319–329 (2005)
Yusupov, M. M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Chen, J. Z. & Grigorieff, N. SIGNATURE: a single-particle selection system for molecular electron microscopy. J. Struct. Biol. 157, 168–173 (2006)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
Spahn, C. M. & Penczek, P. A. Exploring conformational modes of macromolecular assemblies by multiparticle cryo-EM. Curr. Opin. Struct. Biol. 19, 623–631 (2009)
Connell, S. R. et al. A new tRNA intermediate revealed on the ribosome during EF4-mediated back-translocation. Nature Struct. Mol. Biol. 15, 910–915 (2008)
Whitford, P. C. et al. An all-atom structure-based potential for proteins: bridging minimal models with all-atom empirical forcefields. Proteins 75, 430–441 (2009)
Whitford, P. C. et al. Nonlocal helix formation is key to understanding S-adenosylmethionine-1 riboswitch function. Biophys. J. 96, L7–L9 (2009)
Orzechowski, M. & Tama, F. Flexible fitting of high-resolution x-ray structures into cryoelectron microscopy maps using biased molecular dynamics simulations. Biophys. J. 95, 5692–5705 (2008)
Lindahl, E., Hess, B. & van der Spoel, D. J. GROMACS 3.0: a package for molecular simulation and trajectory analysis. J. Mol. Model. 7, 306–317 (2001)
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995)
Bocharov, E. V. et al. From structure and dynamics of protein L7/L12 to molecular switching in ribosome. J. Biol. Chem. 279, 17697–17706 (2004)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996)
Acknowledgements
The present work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG; SFB 740 TP A3 and TP Z1, SP 1130/2-1 to C.M.T.S., FU579 1-3 to P.F., HA 1672/7-5 to R.K.H. and WI3285/1-1 to D.N.W.), the European Union 3D-EM Network of Excellence (to C.M.T.S.), the European Union and Senatsverwaltung für Wissenschaft, Forschung und Kultur Berlin (UltraStructureNetwork, Anwenderzentrum) and US National Institutes of Health (NIH; grant GM 60635 to P.A.P.), the Cluster of Excellence ‘Macromolecular complexes’ at the Goethe University Frankfurt (DFG Project EXC 115 to P.F. and S.C.), and the Human Frontiers of Science Program Young Investigators Award HFSP67/07 (to P.F.). We thank the New Mexico Computing Application Center for generous time on the Encanto Supercomputer. P.C.W. is currently funded by a LANL Director’s Fellowship. This work was also supported by the Center for Theoretical Biological Physics sponsored by the National Science Foundation (NSF; grant PHY-0822283) with additional support from NSF-MCB-0543906, the LANL LDRD program and NIH grant R01-GM072686.
Author information
Authors and Affiliations
Contributions
A.M., A.L.S. and A.D. prepared the complexes. A.H.R. and T.M. collected the cryoelectron microscopy data. A.H.R, J.L., M.B., S.R.C. and C.M.T.S. did the image processing. P.C.W., Y.Y., J.O. and K.Y.S. developed and employed the MDFit method. P.W.H. participated in docking and analysed the FA-binding site. A.H.R., R.K.H., S.R.C., P.F., P.A.P., D.N.W. and C.M.T.S. discussed the results and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Results and Discussion, Supplementary Table 1, Supplementary Figures 1-7 with legends and Supplementary References. (PDF 1041 kb)
Supplementary Movie 1
The animation compares the cryo-EM maps of sub-state I (TIPRE) and sub-state II (TIPOST) of the 70S•EF-G•GDP•FA complex, which are shown as mesh superposed with docked molecular models in ribbons representation (see also legend to Fig. 1): EF-G (red), the tRNA (green), the 16S rRNA (yellow), and the 30S ribosomal proteins (pink). The maps are shown from the 50S side with the 50S subunit computationally removed. Alignment was based on the respective 50S subunits. (GIF 2823 kb)
Supplementary Movie 2
The animation compares the cryo-EM maps of sub-state I (TIPRE) and sub-state II (TIPOST) of the 70S•EF-G•GDP•FA complex, which are shown as mesh superposed with docked molecular models in ribbons representation (see also legend to Fig. 1): EF-G (red), the tRNA (green), the 16S rRNA (yellow), and the 30S ribosomal proteins (pink). The maps are shown as a top view onto the 30S subunit with the 50S subunit computationally removed. Alignment was based on the respective 50S subunits. (GIF 1272 kb)
Supplementary Movie 3
The animation compares the molecular models of sub-state I (TIPRE) and sub-state II (TIPOST) of the 70S•EF-G•GDP•FA complex in a close-up of the tRNA binding. The 30S subunit is shown with yellow ribbons, the tRNAs as blue ribbons and ribosomal residues that contact tRNAs in A-, P- and E-sites as spheres coloured magenta, green and orange, respectively (see also Fig. 2a, b). (GIF 1415 kb)
Supplementary Movie 4
In a common 50S alignment, the P/E-tRNA (green ribbon) of TIPRE (sub-state I) and the pe/EtRNA (magenta ribbon) of TIPOST (sub-state II) together with their respective mRNA codons11are compared to the positions of the tRNAs in classical A-, P- and E-sites and the mRNA (grey ribbons). See also Fig. 3d. (GIF 145 kb)
Rights and permissions
About this article
Cite this article
Ratje, A., Loerke, J., Mikolajka, A. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010). https://doi.org/10.1038/nature09547
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09547
This article is cited by
-
Visualizing translation dynamics at atomic detail inside a bacterial cell
Nature (2022)
-
Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
Nature Communications (2021)
-
Insights into genome recoding from the mechanism of a classic +1-frameshifting tRNA
Nature Communications (2021)
-
Distinct mechanisms of the human mitoribosome recycling and antibiotic resistance
Nature Communications (2021)
-
Structural basis for +1 ribosomal frameshifting during EF-G-catalyzed translocation
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.