Abstract
Bacteria that cause chronic and/or recurrent diseases often rely on a biofilm lifestyle. The foundation of the biofilm structure is the extracellular polymeric substance (EPS) that acts as a barrier to both effectors of the immune system and antimicrobial agents. Recent work has highlighted extracellular DNA (eDNA) as a key component common to many pathogenic biofilms. Here, we show that the DNABII family of proteins, well known for their strong structural influences on intracellular DNA, was also critical for the integrity of the EPS matrix of biofilms that contain eDNA. In fact, antisera derived against a purified Escherichia coli DNABII family member rapidly disrupts the biofilm EPS formed by multiple human pathogens in vitro. In addition, when a member of this family of proteins was used as an immunogen in an animal model in which the bacteria had already formed a robust biofilm at the site of infection, the resultant targeted immune response strongly ameliorated this biofilm disease in vivo. Finally, this methodology to debulk the biofilm of EPS was shown to work synergistically with otherwise ineffective traditional anti-microbial approaches in vitro. We discuss the prospects for targeting DNABII family members as a potential universal strategy for treating biofilm diseases.
Similar content being viewed by others
INTRODUCTION
The National Institutes of Health and Center for Disease Control and Prevention estimate that >80% of all bacterial infections include a necessary biofilm phase within the disease course (http://grants.nih.gov/grants/guide/pa-files/PA-03-047.html). Biofilms are not mere accretions of bacteria formed on a surface, but rather are active communities replete with organized functions, such as division of labor, interbacterial communication, and transportation networks. Operationally, the most remarkable difference between biofilm bacteria and their non-adhered planktonic counterparts is the presence of an extensive extracellular polymeric substance (EPS) that enshrouds and protects the resident bacteria of a biofilm community.
The EPS serves many functional roles during the biofilm phase. First, EPS can account for 90% of the dry mass of the biofilm.1 The EPS also creates a semi-permeable barrier that not only keeps critical nutrients and metabolites “in” by lessening diffusion and maintaining proximity but is also essential in terms of keeping dangers to the bacteria “out.” Moreover, in addition to the EPS limiting access by effectors of both the innate and the adaptive immune systems to bacteria within the biofilm, the EPS also protects the resident bacteria against therapeutic modalities (e.g., antibiotics that often are required at a dose 1,000-fold greater than would be required to eliminate acute planktonic infections).2
Although constituents of the EPS vary among bacteria, one common component is extracellular DNA (eDNA). Whitchurch et al.3 showed that treatment of pre-formed Pseudomonas biofilms with DNase sufficiently undermined the EPS to cause dispersal of the resident bacteria. Subsequent studies have yielded similar results.4, 5, 6, 7, 8, 9, 10, 11, 12 The DNA within these biofilms appears to be primarily prokaryotic in origin, resulting from lysed or autolysed resident bacterial cells. However, eDNA can also originate from polymorphonuclear neutrophils, which release DNA in the form of “nets” that can trap and kill microbes and is an area of tremendous current interest.13 In diseases with a biofilm component, biofilms formed in vivo are likely to be composed of eDNA of both host and bacterial origins. In terms of how this eDNA is integrated into the biofilm at the site of infection, Jurcisek and Bakaletz8 have shown that in biofilms formed in vivo by non-typeable Haemophilus influenzae (NTHI), the eDNA is observed to be organized within a heavily interwoven mesh-like structure. Although eDNA arrangement of this nature has not been reported for all bacteria, this observation does suggest that eDNA is not randomly arranged, and further that the infrastructure of the eDNA is likely important to the structural integrity of the biofilm. Indeed, recently, Lappann et al.12 showed that in Neisseria meningitidis, although crude chromosomal DNA readily promoted wild-type biofilm formation and structure, purified DNA could not. This latter observation led the authors to suggest that there might be critical proteins required to facilitate the structure and function of the eDNA in situ.
Whereas bacteria produce many extracellular proteins, few if any DNA-binding proteins have been identified outside the bacterial cell. One exception is the DNABII family of proteins. These proteins exist in two subtypes, HU and IHF (histone-like protein from Escherichia coli strain U93 and integration host factor, respectively). HU is ubiquitous in Eubacteria. Although HU binds (weakly) and bends double-stranded DNA in a non-sequence-specific manner,14 HU demonstrates an up to 1,000-fold increase in affinity for highly structured/pre-bent (e.g., Holliday Junctions or cruciform structures)15, 16, 17 double-stranded DNA. Conversely, IHF is expressed only by bacteria within the α- and γ-proteobacteria genera.18 Although IHF also forms a heterodimer, binds, and compacts double-stranded DNA and has a preference for structured/pre-bent DNA,18 this latter DNABII family member is unique in that it demonstrates high affinity for double-stranded DNA that shares the consensus sequence WATCAANNNNTTR (where W is A or T, N is any base, and R a purine18). All known DNABII family members are ∼10 kDa, in which the functional protomer is a dimer of identical or similar subunits.
The fact that the DNABII family of proteins also exists extracellularly has been known for ∼30 years.19 At the time this observation was reported, Murray Stinson et al. were examining pathogenic Streptococcus, searching for soluble factors responsible for the development of post-infection sequelae. They found that Streptococcal HU not only bound to host cell surfaces but also that the purified protein elicited a strong innate immune response. Recently, others have found that HU from additional genera are also present in the extracellular milieu.20, 21, 22, 23, 24
Given that the biofilm matrix formed by NTHI possesses a highly interwoven structure of eDNA, we decided to investigate whether the DNABII family had a role in the structural integrity of biofilm EPS by its interaction with eDNA. In vitro immunofluorescence studies of pre-formed NTHI biofilms that used antibody directed against E. coli IHF showed strategic positioning of a cross-reacting protein on each vertex of crossed strands of eDNA. Furthermore, biofilms formed by either NTHI or all of eight additional pathogenic bacterial species showed notable “debulking” of the EPS upon incubation with an anti-IHF antibody. We further show that when purified E. coli IHF was used as an immunogen in an established animal model of experimental otitis media, in which NTHI has been allowed to first form a robust biofilm within the middle ear cavities, the resultant immune response significantly augmented eradication of the biomass and clearance of bacteria from this anatomical niche. Combined, these outcomes resulted in significantly greater and more rapid resolution of disease when compared with controls. Finally, we show that debulking the biofilm of EPS rendered NTHI markedly more susceptible to other traditional therapeutic modalities. Implications for harnessing the power of the immune system to induce debulking of biofilms formed in vivo as a more universal strategy to resolve multiple chronic biofilm diseases are discussed.
RESULTS
DNABII proteins were located at the vertices of bent and crossed strands of eDNA within an NTHI biofilm formed in vivo
In consideration of the interwoven arrangement of the eDNA present within an NTHI biofilm that we know from our previous work bears a striking resemblance to cruciform DNA (compare Figure 1a with b, lower section) and further, of the precedent that DNABII proteins exist extracellularly, we hypothesized that a DNABII family member might be responsible for the architectural character of NTHI EPS and thus could perhaps be localized by immunofluorescence microscopy. To this end, we chose to examine whether rabbit polyclonal antiserum directed against purified E. coli IHF (henceforth referred to as anti-IHF) could label frozen sections of an NTHI-formed biofilm recovered from the chinchilla middle ear during experimental otitis media. This antiserum has been shown to cross-react with all DNABII family members from various genera including HU isolated from a Gram-positive bacterium (although at lower avidity; Steven Goodman, unpublished).
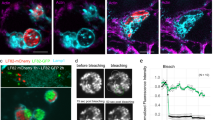
Localization of a DNABII protein at vertices of bent eDNA within an in situ biofilm. (a) Immunofluorescent image of a biofilm formed in the middle ear of the chinchilla (21 days after challenge with non-typeable Haemophilus influenzae). Fine, widely spaced dsDNA strands are labeled with DAPI and appear blue in this image, whereas non-typeable H. influenzae are labeled with FITC-conjugated antiserum directed at a surface-exposed protein and appear green. Marker bar=5 μm. (b) DNABII family members have a stronger preference for binding to bent DNA—a generalized scheme. (c) Immunohistochemical labeling of IHF within an NTHI biofilm formed in vivo. The strong labeling of each vertice where individual strands of DNA cross (arrows) must be noted. FITC, fluorescein isothiocyanate; IHF, integration host factor; NTHI, non-typeable Haemophilus influenzae.
As shown in Figure 1c, eDNA can be seen as strands of DNA (white), appearing in the form of an interwoven lattice or mesh. Remarkably, at virtually all visualized vertices of eDNA, we observed positive immunolabeling with anti-IHF antibody as shown by punctate fluorescence signals (pseudo-colored red). The use of naive rabbit serum in combination with Alexa Fluor-conjugated goat anti-rabbit IgG or with Alexa Fluor-conjugated goat anti-rabbit IgG alone did not result in labeling of vertices. Mean distance between vertices was ∼6 μm or ∼18 kb between each vertex if one assumes 0.34 nm per base of DNA for the B-form DNA. To the best of our knowledge, the only proteins that possess epitopes that cross-react with anti-IHF are HU and IHF. Therefore, based on these observations, it seemed that not only were there extracellular DNABII proteins within the NTHI biofilm matrix but more importantly also that these proteins seemed to be exclusively positioned on eDNA strands that resembled cruciform structures in conformation (see Figure 1b, bottom section), thus strongly suggesting their role in mediating the resulting bent conformation of the eDNA.
In vitro pre-formed NTHI biofilms were destabilized by, and resident bacteria were dispersed, upon treatment with anti-IHF
The notable labeling of what seemed to be virtually all visualized vertices of the eDNA lattice within the NTHI biofilm when incubated with anti-IHF suggested that a DNABII family member was involved in the structural integrity of this abundant eDNA. To determine whether incubation of these biofilms with anti-IHF might result in a loss of structural integrity of the biofilm matrix, we treated pre-formed NTHI biofilms with either anti-IHF or naive serum at a 1:50 dilution for 16 h. When compared with a biofilm incubated with naive serum (Figure 2a), as shown in Figure 2b, the biofilm showed a dramatic loss of structure when incubated with antiserum directed against IHF. Through COMSTAT analysis of multiple replicate assays, the measured parameters of biofilm height, biomass, and biofilm thickness were all diminished by a mean of >80% upon incubation with anti-IHF.
Effect of incubation of biofilms formed in vitro by NTHI with either naive rabbit serum (panel a) or rabbit anti-IHF (panel b). Naive serum had no measurable effect on biofilm height, architecture or viability (panel a) however anti-IHF notably reduced biofilm height and architecture (panel b). Use of live-dead staining and the absence of red fluorescence in panel b suggests that despite reduction of the biomass by incubation with antiserum directed against IHF, remaining bacteria were viable. IHF, integration host factor; NTHI, non-typeable Haemophilus influenzae.
As live-dead staining showed that the bacteria remaining in the biofilm after incubation with anti-IHF were viable (the absence of red fluorescence in Figure 2b must be noted), we reasoned that exposure to anti-IHF was inducing release of NTHI from the biofilm rather than (or perhaps in addition to) death. To determine whether there was an increase in released (e.g., planktonic) bacteria over time upon treatment with anti-IHF, we thereby repeated this experiment with additional titration of the antisera, as well as enumeration of viable planktonic bacteria recovered from the culture medium present within the chamber slide but above the anchored biofilm. As shown in Figure 3, relatively few bacteria were released into the culture medium of the chamber slide upon incubation of a pre-formed biofilm with either sterile culture medium (sterile brain heart infusion (sBHI)) or naive rabbit serum over time (compare red and blue lines at 1 vs. 10 h of incubation). However, in contrast, incubation with anti-IHF resulted in a marked increase in planktonic bacteria available for culture from the medium within the chamber slide within ∼6 h, and increasing notably at 10 h of incubation (see green line in Figure 3). These results suggested the release of bacteria from the biofilm matrix. When considered in combination with confocal images shown in Figure 2, these data further suggested that incubation with anti-IHF perhaps induced a loss of structural integrity of the biofilm, which resulted in a physical release of bacteria from the EPS matrix.
Viable planktonic bacteria released from an existing biofilm formed in vitro by NTHI after exposure to sterile medium, naive rabbit serum, or rabbit anti-IHF serum. Whereas increased release of bacteria into the planktonic phase upon incubation with anti-IHF serum, but not with either naive serum or sBHI, was detected within the first 8 h of treatment, this result was markedly increased at 10 h of incubation. IHF, integration host factor; NTHI, non-typeable Haemophilus influenzae, sBHI, sterile brain heart infusion.
Biofilms formed in vitro by various pathogenic bacteria of multiple genera show loss of structure upon treatment with anti-IHF
As mentioned earlier, it is well known that other bacterial genera also release DNABII family members into the extracellular milieu.20, 21, 22, 23, 24 To determine whether the loss of structural integrity of the biofilm formed by NTHI upon incubation with anti-E. coli IHF was a general phenomenon, as opposed to one restricted to this microbe, we examined biofilms formed by eight additional bacteria of importance in other chronic and/or recurrent biofilm diseases of humans. As summarized in Table 1, incubation with anti-IHF with each of the five human pathogens resulted in a marked decrease in biofilm height, thickness, and overall biomass for all microbes tested compared with naive serum. Incubation of three additional important human pathogens (Staphylococcus aureus, Neisseria gonorrhoeae, and Psedomonas aeruginosa) with anti-IHF similarly resulted in a notable reduction in mean thickness and biomass (see Supplementary Figure S1 online). Thus, DNABII family members were likely critical components in the maintenance of the structural integrity of each biofilm. This result also strongly suggested that there were likely conserved epitopes within members of the DNABII family that could be targeted for the development of a novel, and perhaps also universal immune therapy approach for diseases caused by these organisms, as well as possibly others.
IHF and HU were critical to the formation of an E. coli biofilm
Members of the DNABII family are highly pleiotropic for multiple nucleoprotein systems including gene transcription, and moreover, are thus often essential. In contrast, both HU and IHF mutants have been generated in laboratory strains of E. coli and studied extensively. To determine what role IHF and HU have in eDNA formation, we incubated biofilms formed by E. coli strain MG1655 or its HU- and IHF-deficient derivatives with anti-IHF. As shown in Figure 4, whereas the parental isolate, strain MG1655 produced a robust biofilm in vitro (Figure 4a), both HU and IHF mutant strains were less robust (Figure 4c and e, respectively). An HU-deficient mutant yielded a biofilm that was ∼½ the height of the wild-type biofilm, whereas IHF-deficient strains produced a biofilm that was ∼2/3 the height of the parental isolate. This result suggested that both proteins were involved in either the production and/or the integrity of the E. coli biofilm EPS. After treatment with anti-IHF, the biofilm formed by the wild-type isolate showed notable debulking in terms of height, biomass, and mean thickness (Figure 4b), whereas that formed by the HU mutant showed no reduction in height and a lesser effect on both biomass and mean thickness (Figure 4d). In contrast, no effect on the biofilm formed by the IHF mutant was observed in terms of loss of height, biomass, or mean thickness (Figure 4f). Collectively, these data indicated that for this strain of E. coli, IHF was likely the only structural element that could be targeted by use of the anti-IHF antibody. To date, we have only seen the highly structured interwoven eDNA in biofilms that have been formed in vivo, confirmation of the expression of the remaining DNABII family member by each respective mutant, and any change in the resultant DNA structure, awaits an in vivo immunofluorescent analysis.
Effect of incubation of biofilms formed in vitro by E. coli with either naive serum or with anti-IHF serum. Representative images of biofilms are shown in panels a–f with height of individual biofilms shown to the right of each image, whereas mean values in terms of percentage reduction in biofilm height, biomass, and mean thickness as mediated by incubation with anti-IHF are shown in the tables at the end of each row. (Panels a and b) Parental strain MG1655; (panels c and d) HU-deficient hupA, hupB double mutant; (panels e and f) IHF-deficient himD, himA double mutant. The ability of anti-IHF serum to reduce biomass induced by either the parental isolate or the HU-deficient mutant, but not the IHF-deficient strain, as expected must be noted. (Negative values listed in the table that accompanies panels e and f indicate that no relative reduction in biomass was observed upon incubation with anti-IHF). E. coli, Escherichia coli; HU, histone-like protein; IHF, integration host factor; NTHI, non-typeable Haemophilus influenzae.
Active immunization of chinchillas that have pre-formed NTHI-induced biofilms within their middle ears with purified E. coli IHF resulted in rapid disease resolution
In vitro debulking of biofilms formed by multiple genera of bacteria known to be causative agents of human disease by anti-IHF provided compelling evidence as to the role of DNABII family members in the maintenance of EPS integrity. However, it remained to be seen whether using a DNABII family member as an immunogen could induce the formation of antibodies in a mammalian host that were then similarly capable of both eradicating a pre-formed biofilm and also inducing more rapid resolution of an experimental disease state compared with animals that were immunized with adjuvant only as a control.
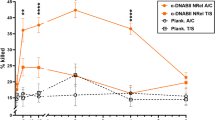
To this end, chinchillas that had been previously challenged with NTHI directly into the middle ear space to induce the formation of a robust biofilm were immunized to determine whether induction of an anti-IHF immune response would alter the course and/or severity of experimental disease. The chinchilla model of biofilms formed in situ by NTHI has been shown by several laboratories to recapitulate the disease state in humans.8, 25, 26, 27, 28, 29, 30, 31, 32, 33 Similarly, upon direct challenge of the middle ear as performed here, chinchillas present with extensive fluid and a biomass that typically fills both the superior and the inferior spaces of the tympanic bullae (or middle ear). To determine whether we could induce a local and systemic acquired immune response that was sufficient to resolve these pre-formed biofilms, we began to immunize chinchillas 4 days after they had been directly challenged into the middle ear with NTHI. Chinchillas received a primary immunizing dose on day 4 after bacterial challenge and a boosting dose 7 days later (on day 11 after the NTHI challenge). Immunogens were delivered transcutaneously using either purified E. coli IHF that had been admixed with the adjuvant dmLT or adjuvant alone. On day 18, after bacterial challenge, chinchillas were killed and their middle ears were tapped of fluids for bacterial culture, followed by macroscopic imaging to allow blinded scoring of any remaining bacterial biomass resident within the middle ear space. Images were scored by 9 blinded evaluators using a 0 to 4+ scoring system (Table 2) to gauge the severity of the remaining disease state. As shown in Figure 5a, the mean score for the remaining biomass within the middle ears of chinchillas immunized with adjuvant only was 2.8, which indicated significant remaining disease and a lack of resolution of pre-formed biomass in most animals by day 18 after bacterial challenge of the middle ear. In contrast, the mean score for an E. coli IHF+ adjuvant-immunized animal was 1.5. Representative images of a residual biomass that received a mean score of +2.8 and one that received a mean score of +1.5 are shown in Figure 5b. Moreover, as shown in Supplementary Figure S2 online, disease resolution was additionally measured by both histological evidence of altered biomass architecture within the middle ear (see Figure 5a) and a statistically significant reduction of bacterial load present within the remaining biomass as measured by homogenization of the biomass and culture on chocolate agar (see Figure 5b). Furthermore, all animals immunized with isolated E. coli IHF mounted a strong local and systemic immune response and no animal presented with obvious secondary sequelae as the result of immunization as noted upon necropsy (data not shown).
Active immunization with native IHF from E. coli induced rapid resolution of experimental otitis media due to NTHI. (a) Immunization with IHF through a transcutaneous delivery route induced the formation of antibodies that significantly reduced the biomass of an NTHI-induced biofilm resident within the middle ears of chinchillas (P<0.001). (b, first column) Representative images of biomasses that remained in the ears of animals immunized with adjuvant alone vs. those immunized with IHF+adjuvant. It must be noted that the majority of the middle ear was filled with a biomass in the animal immunized with adjuvant only (see area circled in red), vs. minimal biomass present within the middle ear cavity of the animal immunized with IHF+adjuvant (see green arrow). The last column in panel b shows images of biomasses at the extremes of the scoring system used here. The top image is that of a middle ear that contains a biomass that would receive a score of 4+, whereas lower image is a healthy middle ear that would receive a score of 0, indicating no biomass. IHF, integration host factor; NTHI, non-typeable Haemophilus influenzae.
The antigenicity of IHF is occluded by bound DNA
The notable observed efficacy when we used anti-IHF in vitro to debulk biofilms and also of purified E. coli IHF, when used as an immunogen in vivo to induce the formation of polyclonal antibodies that could resolve an ongoing biofilm disease, created a conundrum as to why mammalian hosts do not naturally mount an effective immune response to DNABII proteins that are associated with eDNA within the bacterial biofilms of recurrent and/or chronic disease states. In review of the solved structure of IHF when it is bound to DNA,34 it is clear that a significant portion of the protein structure is occluded by bound DNA, which suggested to us the potential for occlusion of protective epitopes or domains of IHF and/or HU when bound to eDNA within a bacterial biofilm. Thus, we hypothesized that the use of native IHF, to which no DNA was bound, as the immunogen might provide a mechanism to overcome such occlusion and thereby foster production of protective antibodies.
To determine whether eDNA could indeed prevent the development of protective antibodies directed against a DNABII family member upon immunization, whereas the use of native protein was effective, we performed a second larger cohort study in which we essentially repeated the chinchilla study as detailed in the previous section. In brief, we allowed NTHI to first form robust biofilms within the middle ears of directly challenged chinchillas before transcutaneous immunization with isolated E. coli IHF, except that we now added two additional cohorts of animals that were immunized either with IHF to which an excess of DNA had been bound or with DNA alone as additional controls. Middle ears were again subsequently scored from 0 to 4+ for disease severity upon completion of the study. To better assure that the IHF and DNA remained in complex for immunization, we used synthetic oligonucleotides identical to those used in the published co-crystalization study34 of a high-affinity IHF-binding site from the bacteriophage λ-recombination site attP at a molar ratio of 2:1 (DNA (10 μm) to IHF (5 μm)), which was at least three orders of magnitude over the Kd of IHF bound to this DNA target. As shown in Supplementary Figure S3 online, IHF indeed bound dsDNA as demonstrated by its reduced mobility in an electrophoretic mobility shift assay.
As shown in Figure 6, animals immunized with isolated E. coli IHF showed a dramatic reduction in disease state with an mean residual middle ear biomass score of 0.9 as compared with controls that had been immunized with adjuvant alone (mean biomass score=2.2) or with those that had been immunized with DNA that had been admixed with adjuvant (mean biomass score=2.8). Interestingly, those animals that had been immunized with the IHF–DNA complex also demonstrated middle ears with significant remaining bacterial biomass, yielding a mean biomass score of 2.5, which was not statistically significantly different from cohorts that received either the adjuvant alone or the DNA that had been admixed with adjuvant. This outcome strongly suggested that if sufficient eDNA fragments were present, as one could hypothesize would be the case during natural disease, this situation could result in occlusion of critical IHF epitopes necessary for the generation of neutralizing antibodies.
Transcutaneous immunization with native IHF+adjuvant induced the formation of antibodies that significantly reduced the biomass resident within the middle ears of chinchillas compared with receipt of either adjuvant alone (P< 0.017), dsDNA alone (P<0.003), or IHF to which dsDNA was already bound+adjuvant (P<0.001). This outcome suggested that binding of dsDNA to native IHF masked protective epitopes of this DNABII family member. IHF, integration host factor.
To assure ourselves that the observed DNA occlusion result was not specific to the use of a transcutaneous immunization route, we repeated these immunizations using a subcutaneous immunization route to ensure delivery of antigens to antigen-presenting cells within the chinchilla host. As shown in Supplementary Figure S4 online, whereas subcutaneous immunization with isolated E. coli IHF yielded the generation of a strong immune response to isolated IHF, immunization with E. coli IHF that had been pre-bound to an excess of DNA induced antibodies that resulted in only very faint recognition of IHF when assayed by western blot. Similarly, when assayed by enzyme-linked immunosorbent assay, reciprocal titer vs. isolated IHF was 1,000 for the animal immunized with IHF that had been pre-bound to oligonucleotides, whereas that for the animal immunized with native IHF was 8,000 (both animal's pre-immune reciprocal titers against IHF were 100). Collectively, these results are consistent with our hypothesis that the binding of IHF to eDNA, as would occur during a natural disease state, has the potential to block epitopes or domains of IHF necessary for generation of a protective acquired immune response. Furthermore, immunization with native IHF (to which no DNA is bound) appeared to allow us to effectively direct the immune response toward the generation of protective or neutralizing antibodies, as demonstrated in both pre-clinical chinchilla studies described here.
Debulking of pre-formed NTHI biofilms with anti-IHF in vitro allowed demonstration of a synergistic effect when used in combination with therapeutic modalities
Whereas we have demonstrated in vitro and in vivo that both anti-IHF and the use of IHF as an immunogen show utility in debulking biofilms and/or resolving biofilm disease respectively, we next speculated whether debulking of NTHI biofilms with anti-IHF might also allow synergism when used in conjunction with other therapeutic modalities. To this end, we assessed the ability to induce augmented structural destabilization of pre-formed NTHI biofilms when a suboptimal concentration of anti-IHF was used along with one of each of three unique reagents, namely a DNA-degrading enzyme (DNaseI) known to be able to degrade an NTHI biofilm in vitro (but used here at a suboptimal concentration), antisera to an outer membrane protein of NTHI that did not destabilize a pre-formed NTHI biofilm when used alone (anti-P5) (data not shown), or an antibiotic typically used as a first-line choice in children with chronic and/or recurrent OM (amoxicillin),35 but which has limited efficacy against bacteria resident within a biofilm community.
Figure 7a shows that treatment of an NTHI biofilm with a concentration of DNase shown to be suboptimal has a marginal effect (see Figure 7a II). Similarly, a 1:200 dilution of anti-IHF had limited effect on the pre-formed NTHI biofilm (see Figure 7a III). However, in contrast, when these two reagents were used in concert, the biofilm was notably diminished (see Figure 7a IV). Upon repetition of this experiment three times (see Supplementary Table S1 online), we found that the most marked synergistic effect of debulking of the biofilm was measured as a diminution in height. One simple explanation for this outcome was that as the DNABII protein was being titrated away from the periphery of the biofilm, the eDNA became more accessible to the action of the DNase.
Synergistic effect of rabbit anti-IHF when used in combination with other therapeutic modalities. Demonstration of synergism between a suboptimal concentration of anti-IHF serum (1:200) and DNAseI individually, then mixed (a); a suboptimal concentration of anti-IHF (1:100) and anti-outer membrane protein P5 serum (OMP P5) individually, then mixed (b); or that of an effective dilution of anti-IHF (in terms of debulking a biofilm but not inducing bacterial cell death) and amoxicillin individually, then mixed (c). In each of these situations, when any agent was combined with anti-IHF, the biofilm debulking and/or killing effect observed was greater than that noted when any single agent was used alone. Biofilm height (in microns) is indicated under each image. IHF, integration host factor.
Figure 7b shows the results of treatment with anti-P5 on pre-formed NTHI biofilms. Although this antisera is strongly reactive with isolated NTHI P5 (data not shown) and further, active immunization with isolated P5 is effective in mediating significant protection against experimental OM in chinchilla models,36, 37, 38, 39, 40, 41, 42 this antiserum does not induce a change in the structural integrity of an NTHI biofilm that has been formed in vitro when used at a dilution of 1:50 (see Figure 7b II) (when replicate assays were analyzed, the mean reduction in maximum height was 4%). Similarly, we observed a marginal effect when these biofilms were incubated with a suboptimal dilution of anti-IHF (used at a 1:100 dilution here) (see Figure 7b I). When used at this dilution, anti-IHF induced a 38% mean reduction in maximum height. However, when combined, the use of anti-P5 plus anti-IHF resulted in a mean reduction in the height of the biofilm that exceeded the sum of the two antisera when used singly (50%) (see Figure 7b III), thus indicating a synergistic outcome. Hence, we concluded that the use of anti-IHF to destabilize the NTHI EPS matrix resulted in the exposure of the targeted bacterial cell surface protein (e.g., OMP P5) that would otherwise be obscured by eDNA and perhaps other components of the EPS, thus allowing immune-mediated bacterial clearance by as yet unknown mechanisms.
Finally, Figure 7c shows the results of treating pre-formed NTHI biofilms with amoxicillin. When used at a concentration of 64 μg ml−1, amoxicillin had no measurable effect on the architecture of pre-formed NTHI biofilms (see Figure 7c II), despite evidence of limited bacterial cell death. When replicate runs were analyzed, treatment with amoxicillin alone resulted in an 8% mean reduction in maximum height. Treatment with IHF antisera at a 1:50 dilution substantially reduced the height of the biofilm (mean reduction 61%) as we have shown previously (see Figure 7c III). However, interestingly, when the two reagents were used simultaneously, not only was there a dramatic reduction in the height of the biofilm from 17 μm (as induced by treatment with anti-IHF alone) to 10 μm (as induced by anti-IHF used in combination with amoxicillin) (see Figure 7c IV) but the use of a vital dye also indicated that the majority of the bacteria remaining in the biofilm were now dead as noted by the predominant fluorescence in the red channel within the imaged biofilm. This result showed that debulking of the biofilm with anti-IHF likely exposed the bacteria sufficiently so as to create conditions more akin to susceptibility to amoxicillin concentrations known to be effective when assayed against planktonic NTHI. This outcome may have been mediated by either increased physical exposure of bacteria within the remaining biofilm matrix to the action of amoxicillin and/or through increased release of bacteria into the planktonic phase as we showed earlier can occur during biofilm debulking by exposure to anti-IHF antibodies (see Figure 3).
DISCUSSION
We have demonstrated both in vitro and in vivo that members of the DNABII family are intimately involved in the maintenance of the structural integrity of eDNA present within biofilms formed by NTHI, the most prevalent pathogen of chronic and recurrent otitis media. Furthermore, in all of the genera of bacteria tested in vitro to date, the DNABII family of proteins seems to have a similar pivotal role. For otitis media due to NTHI, the failure of the immune system to mount a significant counter-response during experimental disease seemed to be the result of DNA occlusion of critical epitopes of the native IHF protein. Finally, the general debulking of eDNA within the biofilm EPS, through the action of anti-IHF, seemed to allow sufficient access to NTHI residing within the biofilm community so as to create the opportunity to invigorate otherwise ineffective therapeutic modalities and to facilitate immune-mediated clearance.
DNA structure, other nucleoid-associated proteins, cruciform binding
The involvement of the DNABII family in maintaining the structural integrity of eDNA is consistent with their ubiquitous presence in Eubacteria, their complete absence in Eukarya, their apparent presence extracellularly, and their preference for association with pre-bent structural motifs. The fact that the interwoven mesh of eDNA present within an NTHI biofilm that was formed in situ within the middle ear cavity so remarkably resembles cruciform structures only adds to the premise that these proteins find the bent portions of these structures as high-affinity targets. Kamashev and Rouviere-Yaniv17 have shown that E. coli HU binds with high affinity to a multitude of DNAs with bent architectures and to branched-like DNA structures. Although the exact nature of these DNA structures remains to be determined, we suggest that the DNABII family could either stabilize DNA conformations by binding to them and/or recruit other proteins that stabilize the structure.
Could other proteins be involved in contributing to and/or stabilizing the structure of the eDNA within a biofilm matrix? The DNABII family is known to function as an accessory protein in a multitude of nucleoprotein systems. Although these proteins can show strong site preference in these systems, there has been no compelling evidence to demonstrate that they readily form any specific protein–protein interactions with heterologous proteins; their function seems to be strictly architectural.43, 44 However, there are a plethora of DNA-binding proteins expressed by bacteria. Furthermore, the DNABII family is a member of the nucleoid-associated protein superfamily. In E. coli, there are at least 11 nucleoid-associated proteins in addition to HU and IHF,45, 46 and thus, at this juncture, it would be impossible to completely rule out their involvement (cursory or otherwise) in contributing to the DNA structure described herein. Two of the most abundant proteins, H-NS and DPS, are known to have a predominant role affecting DNA structure intracellularly. However, antisera directed against these proteins failed to alter the structure of either an NTHI (data not shown) or an E. coli biofilm (see Supplementary Figure S5 online). This latter observation suggested that these proteins do not have a dominant affect on biofilm structure in and of themselves. It remains to be seen whether any additional nucleoid-associated proteins, either individually or en masse, affect the EPS structure. Finally, Bianchi and co-workers15 showed that, of all of the DNA-binding proteins known to be expressed by E. coli, only HU bound to cruciform structures with high affinity. If the eDNA lattice that we along with others have observed within bacterial biofilms is truly a sufficiently close derivative of the cruciform structure, this finding may indicate that only the DNABII family of proteins contribute significantly to the structural integrity of the eDNA.
How does debulking occur?
It is difficult to imagine a means by which the antibody per se could effectively penetrate the EPS to gain access to the resident bacteria, otherwise antisera directed against other surface antigens (e.g., OMP P5 of NTHI) could be effective therapeutic reagents as well. Our parsimonious model is that exposure to anti-IHF antibodies induces an equilibrium shift. The equilibrium dissociation constant for the DNABII family to specific targets is in the nanomolar range. We argue that when these proteins are in the “off” state they are titrated away from the biofilm by anti-IHF antibody. As the equilibrium shifts to the off state, the eDNA structures within the biofilm are destabilized and collapse.
Universality of the DNABII family and universality of eDNA, a common matrix with exceptions
The significance of the results presented here lies in the ubiquity of the DNABII family, the apparent conservation of epitopes, and their capacity to bind DNA-secondary structures. To the best of our knowledge, all Eubacteria express at least one member of the DNABII family and to date this is the protein HU. Ironically, all of our in vitro experiments were performed with polyclonal antiserum directed against IHF, isolated from E. coli. Importantly, all of the biofilms that we prepared were susceptible to the action of this antibody, regardless of whether that strain expressed a specific IHF; this is particularly notable for the Gram-positive bacteria tested as they do not express IHF. All members of the DNABII family share substantial homology which can vary significantly between genera.47 Collectively, our observations would suggest that DNABII proteins expressed by various bacteria share one or more highly conserved epitopes, thereby signifying the presence of a potentially more universal therapeutic target than a protein expressed by one or a limited number of bacteria.
Most intriguing is the apparent conservation of eDNA structural integrity (i.e., that it is reliant on both eDNA and the presence of at least one DNABII member for robustness). We have already speculated as to the mechanisms that underlie initiation of a common structure, and despite the fact that others have found that even in biofilms formed by representative pathogenic bacteria as described here, an obvious motif (as judged by confocal microscopy) is lacking.1 We nonetheless postulate that this three-dimensional architectural arrangement of eDNA and associated proteins has evolved to accommodate the intermingling of heterogeneous bacterial genera. It is evident that in most cases in various ecological niches, biofilms are non-homogeneous. Hence, for diverse bacteria to form productive consortia, it would be necessary to have a compatible EPS, i.e., an EPS that is sufficiently universal such that all constituent bacterial species within the consortia would have a common functional medium. We argue that eDNA is such a material, and the implied universality of the presence of DNABII family proteins within an eDNA-abundant biofilm matrix, and its apparent important structural function, strongly suggests that the resulting structures also need to be compatible.
An exception to the above-stated hypothesis would be, e.g., a species that would need to remain microbiologically homogeneous in terms of its biofilm state. In this case, we would argue that eDNA and/or its three-dimensional EPS structure must remain unique to prevent other species or genera from infiltrating the biofilm. Case in point, Berne et al.48 recently showed that eDNA actually prevents Caulobacter crescentus from transitioning into a sessile stalked-cell state, a clear situation in which these bacteria need to remain homogenous to form an appropriate aggregate structure.
How does the DNABII protein get outside the cell?
These proteins do not have any known signal peptide sequence and hence are not likely to use the canonical secretory pathways to exit bacterial cells. However, it is interesting that in Gram-positive bacteria, HU is essential49 for the signal-recognition particle complex. It is conceivable that HU mediates exit from the cell through this pathway. In contrast, there is no other obvious novel mode that would explain the presence of the DNABII family outside the cell. In the absence of any other explanation, we imagine that the exiting mechanism for the DNABII family members by which they gain access to the extracellular environment is by the same mechanism as that for eDNA, by lysis, autolysis, and/or other programmed means.50, 51
Occlusion of epitopes, conserved epitopes, protein–protein vs. protein–DNA interaction
Our studies showed that the presence of DNA bound to IHF sufficiently prevented the immune system from mounting a protective adaptive immune response. The co-crystal structure of IHF bound to the same DNA sequence used in our study shows three regions wherein there was extensive binding of DNA,34 namely specific regions of the N terminus, β-ribbons that comprise the initial DNA contact points, and specific regions of the C terminus. Interestingly, all three regions are conserved throughout the DNABII family. The simplest explanation is that the DNA itself occludes critical epitopes from presentation to the immune system. Alternatively, as the active protomer of this family is a dimer, it is possible that true epitopes are actually occluded through protein–protein interactions of individual monomers and that the role of the bound DNA is to stabilize the dimer structure. Another formal possibility is that the presence of DNA alters the processing of the antigen and the complex is not presented properly as the case for the native protein. In the absence of any evidence to the contrary, we prefer the theory that it is the DNA itself that occludes the critical protective epitopes, although all explanations are not mutually exclusive.
Synergism and future directions for debulking
Our most intriguing results were: (i) the ability to use IHF as an immunogen to mediate the formation of antibodies that were able to resolve an existing biofilm infection and (ii) the ability to debulk the biofilm EPS of eDNA by exposure to anti-IHF antibodies as a means to make chronic reservoirs of bacteria more susceptible for other therapeutic modalities. Although both of these outcomes were exciting and highly encouraging in terms of the potential to develop a more universal immunization and/or therapeutic strategy as has been a desired goal for remaining diseases of recurrent and chronic nature,52 the power of this approach would need to be appropriately controlled. Although we observed no remote site sequelae in any of our animal studies, it is conceivable that a general debulking of biofilms throughout the body, and including biofilms composed of commensal or normal flora, could facilitate the formation of opportunistic infections of both bacterial and non-bacterial origins, as we do not yet know whether all resident bacteria in a human host would be equally affected by a general vaccination strategy wherein IHF or HU was used as the immunogen.
To harness the power of this strategy, yet also to ameliorate the possibility of inducing collateral damage, we imagine three general non-exclusive approaches. First, we would propose to use a highly directed immunization strategy, attempting to induce the most robust immune response directly at the disease site,53, 54 thereby lessening the likelihood of undesired remote site effectiveness. We have recently shown that the same transcutaneous approach used in the studies presented here have been highly effective in stimulating the most robust response by those dendritic cells in closest proximity to the immunization site,55 as has been shown by others.53 Second, we believe that it will likely be possible, through epitope-mapping efforts, to determine the limited portion of IHF or HU required to induce a highly effective and broadly protective immune response, thereby making an induced immune response more species or even perhaps, strain specific. Third, as we have shown in vitro by combining antisera or other antimicrobial therapies with antibodies directed against a DNABII family member, it should similarly be possible to debulk any biofilm that meets these specifications (e.g., contains both eDNA and an integrated DNABII family member protein) in general using anti-DNABII protein antibodies, followed by or in concert with, a highly microbe- and/or disease-specific agent used locally to thereby mediate an anatomic site-directed “surgical” approach to biofilm removal and/or disease resolution.
METHODS
Antisera and reagents. Purified E. coli IHF and polyclonal antisera from rabbits directed against E. coli IHF were gifts from Howard Nash.34, 56 Purified antibody directed against E. coli DPS and antisera to E. coli DPS were gifts from Emilia Chiancone; the DPS protein was used to verify the reactivity of the antisera by western analysis (data not shown). Monoclonal antibody H113 directed against E. coli H-NS was the gift from Jay Hinton and its reactivity was verified through western analysis of purified H-NS protein, the gift from Linda Kenney (data not shown).
Bacterial strains and growth conditions. Conditions for growing NTHI (strain 86-028NP),36, 37, 38, 57, 58, 59 E. coli (strain MG1655), uropathogenic E. coli (strain UTT89), Streptococcus mutans (strain UA159), Streptococcus pneumoniae (strain 1121), S. aureus (strain 29123), N. gonorrhoeae (strain 1291), P. aeruginosa (strain 27853), and Staphylococcus epidermidis for use in in vitro biofilm assays were as follows. NTHI was cultured on chocolate agar for 18 h at 37°C, in a humidified atmosphere containing 5% CO2, then suspended in BHI broth that had been supplemented with 2 μg each of heme (Sigma-Aldrich, St Louis, MO) and β-NAD (Fisher Scientific, Pittsburgh, PA) per ml (sBHI). P. aeruginosa, S. aureus, and S. epidermidis were cultured on chocolate agar, then suspended in tryptic soy broth. Moraxella catarrhalis was grown on chocolate agar before suspension in Todd Hewitt broth. S. pneumoniae and S. pyogenes were cultured on blood agar. S. pneumoniae was then suspended in Todd Hewitt broth, whereas S. pyogenes was suspended in tryptic soy broth. S. mutans was grown on BHI agar and then suspended in Todd Hewitt broth, whereas N. gonorrheae was grown on gonococcal agar and then suspended in Todd Hewitt broth. Finally, E. coli was cultured on Luria-Bertani agar and suspended in the Luria-Bertani broth.
The E. coli himD∷cam, himAΔ82∷Tn10 mutations were independently and sequentially introduced into MG1655. The himD∷cam insertional mutation was transduced from E. coli HN106960 into MG165561 using standard P1 transduction,62 and transductants were selected on Luria-Bertani agar (Fisher Scientific) containing 25 μg chloramphenicol per ml (Fisher Scientific). The himAΔ82∷Tn10 mutation was then introduced into MG1655 or MG1655 himD∷cam from strain HN107763 into MG1655 himD∷cam using P1 transduction with selection on Luria-Bertani agar containing 25 μg chloramphenicol (MG1655 himAΔ82∷Tn10) or 25 μg chloramphenicol per ml and 25 μg tetracycline hydrochloride per ml (MG1655 himD∷cam, himAΔ82∷Tn10) (Fisher Scientific). The MG1655 hupA∷cam, hupB∷kan double mutant was constructed by P1 transduction from SG732,64 and transductants were selected on Luria-Bertani Broth agar (Fisher Scientific) containing 25 μg chloramphenicol per ml and 25 μg kanamycin sulfate per ml (Fisher Scientific).
Given that the binding site IHF 1 is required for the induction of the fim operon, as described previously,65 the production of type 1 pili by each of the mutants was determined by the ability to agglutinate yeast.66 Mannose-sensitive agglutination was observed for MG1655 himAΔ82∷Tn10, indicating the presence of type 1 pili. Consistent with other reports,65 no agglutination was observed for either MG1655 himD∷cam or MG1655 himD∷cam, himAΔ82∷Tn10, indicating the absence of type 1 piliation.
In vitro biofilm analysis. For in vitro biofilm analysis of NTHI, after suspending bacteria in sBHI as described above, the suspension was adjusted to an optical density of 0.65 at 490 nm. Cultures were then incubated statically for 3 h at 37°C, 5% CO2. We next diluted cultures 1:2,500 with sBHI, and 200 μl of this bacterial suspension was inoculated into each well of an eight-well chamber slide (Fisher Scientific). After 16 h incubation at 37°C, 5% CO2, the medium in each well was replaced with fresh medium. After an additional 8 h incubation period, the medium was removed and replaced with either pre-warmed medium, medium that contained a 1:50 dilution of naive rabbit serum, or medium that contained a 1:50 dilution of rabbit anti-IHF serum. Chamber slides were then incubated an additional 16 h as before.
For in vitro biofilm analysis of all other bacteria strains, bacteria were cultured on their respective agar for 18 h at 37°C, 5% CO2, suspended in the appropriate medium, adjusted to an optical density of 0.65 at 490 nm, and diluted 1:6 in medium. Cultures were then incubated statically for 3 h at 37°C, 5% CO2. The cultures were next diluted 1:2,500 in their appropriate medium and 200 μl inoculated into each well of an eight-well chamber slide (Fisher Scientific). After 16 h incubation at 37°C, 5% CO2, the medium in each well was changed. After an additional 8 h incubation, the medium was removed and replaced with either pre-warmed medium, medium with naive rabbit serum diluted 1:50, or medium with rabbit anti-IHF diluted 1:50 and incubated an additional 16 h as before.
For all bacterial isolates, after the final incubation with immune serum, the medium was removed, the biofilms washed with saline and stained with Live/Dead Stain (Molecular Probes, Eugene, OR) for 15 min. Biofilms were then fixed with 2% paraformaldehyde in 0.1 m phosphate buffer, pH 7.2, for a minimum of 1 h. After washing twice with sterile saline, plastic wells were removed from the slide and a coverslip was placed over the slide, which was then sealed to prevent dehydration. Biofilms were imaged using a × 63 objective on a Zeiss 510 Meta-laser scanning confocal microscope (Carl Zeiss, Thornwood, NY). All in vitro biofilm assays were repeated a minimum of three times, on separate days, and all individual biofilm assays included replicates of three chambers per assay condition on each assay day. Data are presented as mean values ± s.d.
Pre-clinical immunization and challenge studies to determine the therapeutic vaccine potential of IHF. To determine whether antibodies directed against IHF could debulk a biofilm formed in vivo as we had observed in vitro, we first performed a small proof-of-concept study wherein we challenged 8 adult chinchillas with ∼1,000 c.f.u. (colony-forming units) NTHI per bulla and allowed a robust biofilm to develop as we have described previously.55 Four days after bacterial challenge, chinchillas received the first of two transcutaneous immunizations with either 10 μg LT (R192G-L211A) (dmLT; a generous gift from Dr. John Clements, Tulane University) admixed with 10 μg IHF or with 10 μg dmLT alone. In brief, transcutaneous immunizations were performed as follows: both pinnae of each alert animal were hydrated for 5 min by placement of a gauze soaked in sterile, pyrogen-free saline (Hospira, Lake Forest, IL) on the skin that lines the inner surface of the pinnae. Pinnae were then blotted with a dry gauze, and 50 μl of each vaccine formulation was applied using a sterile pipette. The pinnae were then folded over so that the opposing surfaces could be gently rubbed together. Chinchillas received a total of two immunizations delivered at a 1-week interval. One week after receipt of the second dose, animals were killed and images of middle ears captured. Images of the left and right middle ear cavities, with resident biofilms, were scrambled and all 16 images (2 per animal) were compiled into a single file for ranking by blinded evaluators. The relative amount of biomass remaining within the middle ear of each animal was ranked on a 0 to 4+ scale by blinded reviewers using the scale shown in Table 2.
For the second animal immunization study, 20 adult chinchillas (body mass between 500 and 700 g), shown to have no evidence of middle ear disease by tympanometry and video otoscopy,38 were enrolled and divided into 4 cohorts of 5 animals each. All animals were again challenged transbullarly with ∼1,000 c.f.u. NTHI strain 86-028NP per bulla. Four days later, after a biofilm formed in the middle ear cavities of these animals, they were immunized by a transcutaneous route (transcutaneous immunization)55 as described above. Formulations consisted of 10 μg IHF admixed with 10 μg dmLT, 10 μg IHF that had been pre-bound to DNA+dmLT, DNA+dmLT, or 10 μg dmLT alone.
To determine whether transcutaneous immunizations with IHF pre-bound to DNA resulted in a similar immune response to that induced when the same nucleoprotein complex was delivered subcutaneously, and compared with that induced by IHF alone, two adult chinchillas were immunized to generate antisera. Each animal received a priming dose followed by two identical boosts delivered at 30-day intervals. Immunogens were admixed with the adjuvant monophosphoryl lipid A (10 μg per dose) (Sigma-Aldrich) because of its demonstrated strong adjuvant properties in the chinchilla host.37, 38 One chinchilla was immunized with 10 μg of IHF+monophosphoryl lipid A and the other received 10 μg IHF bound to DNA+monophosphoryl lipid A. All doses were delivered subcutaneously in a total volume of 200 μl. Fifteen days after receiving the final boost, animals were bled to procure serum and sera were assayed by western blot and enzyme-linked immunosorbent assay to determine both reciprocal titer and specificity of antibody reactivity to IHF.
Immunofluorescence imaging of frozen sections of biofilms formed in vivo. After dissection, the middle ear of each chinchilla was filled with optimal cutting temperature embedding compound (Fisher Scientific) and flash frozen over liquid nitrogen. The bone that forms the inferior bulla was removed to leave the middle ear mucosa and existing biofilm intact. The resulting block was then bisected to reveal a cross-section of the biofilm and re-embedded in optimal cutting temperature . Serial sections of 10-μm thickness were cut using a Microm rotary cryotome, adhered to glass slides (Mercedes Medical, Sarasota, FL), and stored at −80°C. Sections were later stained to determine the relative incorporation of IHF within biofilms that had formed in vivo. In brief, slides were air-dried, fixed in cold acetone, and then equilibrated in buffer (0.05 m Tris-HCl, 0.15 m NaCl, and 0.05% Tween 20, pH 7.4). Sections were blocked with image-iT FX signal enhancer (Molecular Probes) and with Background Sniper (BioCare Medical, Concord, CA) as per the manufacturer's instructions. Sections were then incubated with a 1:200 dilution of polyclonal rabbit anti-IHF overnight at 4°C, in a humidified chamber. Slides were further rinsed and incubated with goat anti-rabbit IgG conjugated to Alexa Fluor 594 (Invitrogen, Carlsbad, CA) Concord, CA for 30 min at room temperature. As a counterstain, sections were incubated with DAPI and coverslipped using ProLong Gold antifade reagent (Molecular Probes). The use of naive rabbit serum in place of immune serum and the use of secondary antibody alone served as negative control preparations for immunolabeling studies. Sections were viewed using a Zeiss LSM 510 Meta confocal system attached to a Zeiss Axiovert 200 inverted microscope (Carl Zeiss).
Statistical evaluation. Significant differences in mean c.f.u. per mg tissue and mean c.f.u. NTHI per ml supernatant were determined by paired t-test. Significance in relative biomass among cohorts was assessed by unpaired t-test. A P-value ≤0.05 was considered significant.
References
Flemming, H.C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Nickel, J.C., Ruseska, I., Wright, J.B. & Costerton, J.W. Tobramycin resistance of Pseudomonas aeruginosa cells growing as a biofilm on urinary catheter material. Antimicrob. Agents Chemother. 27, 619–624 (1985).
Whitchurch, C.B., Tolker-Nielsen, T., Ragas, P.C. & Mattick, J.S. Extracellular DNA required for bacterial biofilm formation. Science 295, 1487 (2002).
Steinberger, R.E. & Holden, P.A. Extracellular DNA in single- and multiple-species unsaturated biofilms. Appl. Environ. Microbiol. 71, 5404–5410 (2005).
Moscoso, M., Garcia, E. & Lopez, R. Biofilm formation by Streptococcus pneumoniae: role of choline, extracellular DNA, and capsular polysaccharide in microbial accretion. J. Bacteriol. 188, 7785–7795 (2006).
Vlassov, V.V., Laktionov, P.P. & Rykova, E.Y. Extracellular nucleic acids. Bioessays 29, 654–667 (2007).
Qin, Z. et al. Role of autolysin-mediated DNA release in biofilm formation of Staphylococcus epidermidis. Microbiology 153 (Part 7), 2083–2092 (2007).
Jurcisek, J.A. & Bakaletz, L.O. Biofilms formed by nontypeable Haemophilus influenzae in vivo contain both double-stranded DNA and type IV pilin protein. J. Bacteriol. 189, 3868–3875 (2007).
Izano, E.A., Amarante, M.A., Kher, W.B. & Kaplan, J.B. Differential roles of poly-N-acetylglucosamine surface polysaccharide and extracellular DNA in Staphylococcus aureus and Staphylococcus epidermidis biofilms. Appl. Environ. Microbiol. 74, 470–476 (2008).
Thomas, V.C., Thurlow, L.R., Boyle, D. & Hancock, L.E. Regulation of autolysis-dependent extracellular DNA release by Enterococcus faecalis extracellular proteases influences biofilm development. J. Bacteriol. 190, 5690–5698 (2008).
Vilain, S., Pretorius, J.M., Theron, J. & Brozel, V.S. DNA as an adhesin: Bacillus cereus requires extracellular DNA to form biofilms. Appl. Environ. Microbiol. 75, 2861–2868 (2009).
Lappann, M. et al. A dual role of extracellular DNA during biofilm formation of Neisseria meningitidis. Mol. Microbiol. 75, 1355–1371 (2010).
Brinkmann, V. et al. Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535 (2004).
Rouviere-Yaniv, J. & Gros, F. Characterization of a novel, low-molecular-weight DNA-binding protein from Escherichia coli. Proc. Natl Acad. Sci. USA 72, 3428–3432 (1975).
Pontiggia, A., Negri, A., Beltrame, M. & Bianchi, M.E. Protein HU binds specifically to kinked DNA. Mol. Microbiol. 7, 343–350 (1993).
Bonnefoy, E., Takahashi, M. & Yaniv, J.R. DNA-binding parameters of the HU protein of Escherichia coli to cruciform DNA. J. Mol. Biol. 242, 116–129 (1994).
Kamashev, D. & Rouviere-Yaniv, J. The histone-like protein HU binds specifically to DNA recombination and repair intermediates. EMBO J. 19, 6527–6535 (2000).
Swinger, K.K. & Rice, P.A. IHF and HU: flexible architects of bent DNA. Curr. Opin. Struct. Biol. 14, 28–35 (2004).
Stinson, M.W. & Bergey, E.J. Isolation of heart- and kidney-binding protein from group A streptococci. Infect. Immun. 35, 335–342 (1982).
Lunsford, R.D., Nguyen, N. & London, J. DNA-binding activities in Streptococcus gordonii: identification of a receptor-nickase and a histonelike protein. Curr. Microbiol. 32, 95–100 (1996).
Kim, N., Weeks, D.L., Shin, J.M., Scott, D.R., Young, M.K. & Sachs, G. Proteins released by Helicobacter pylori in vitro. J. Bacteriol. 184, 6155–6162 (2002).
Tjalsma, H., Scholler-Guinard, M., Lasonder, E., Ruers, T.J., Willems, H.L. & Swinkels, D.W. Profiling the humoral immune response in colon cancer patients: diagnostic antigens from Streptococcus bovis. Int. J. Cancer 119, 2127–2135 (2006).
Liu, D. et al. Histone-like DNA binding protein of Streptococcus intermedius induces the expression of pro-inflammatory cytokines in human monocytes via activation of ERK1/2 and JNK pathways. Cell Microbiol. 10, 262–276 (2008).
Gao, Q. S. Mutans GtfB and GtfC Gene Expression. [Ph.D. thesis].Univeristy of Southern California, Los Angeles, (2000).
Ehrlich, G.D. et al. Mucosal biofilm formation on middle-ear mucosa in the chinchilla model of otitis media. JAMA 287, 1710–1715 (2002).
Greiner, L.L. et al. Nontypeable Haemophilus influenzae strain 2019 produces a biofilm containing N-acetylneuraminic acid that may mimic sialylated O-linked glycans. Infect. Immun. 72, 4249–4260 (2004).
Bouchet, V. et al. Host-derived sialic acid is incorporated into Haemophilus influenzae lipopolysaccharide and is a major virulence factor in experimental otitis media. Proc. Natl Acad. Sci. USA 100, 8898–8903 (2003).
Swords, W.E., Moore, M.L., Godzicki, L., Bukofzer, G., Mitten, M.J. & VonCannon, J. Sialylation of lipooligosaccharides promotes biofilm formation by nontypeable Haemophilus influenzae. Infect. Immun. 72, 106–113 (2004).
West-Barnette, S., Rockel, A. & Swords, W.E. Biofilm growth increases phosphorylcholine content and decreases potency of nontypeable Haemophilus influenzae endotoxins. Infect. Immun. 74, 1828–1836 (2006).
Daines, D.A. et al. Haemophilus influenzae luxS mutants form a biofilm and have increased virulence. Microb. Pathog. 39, 87–96 (2005).
Hong, W., Mason, K., Jurcisek, J., Novotny, L., Bakaletz, L.O. & Swords, W.E. Phosphorylcholine decreases early inflammation and promotes the establishment of stable biofilm communities of nontypeable Haemophilus influenzae strain 86-028NP in a chinchilla model of otitis media. Infect. Immun. 75, 958–965 (2007).
Jurcisek, J., Greiner, L., Watanabe, H., Zaleski, A., Apicella, M.A. & Bakaletz, L.O. Role of sialic acid and complex carbohydrate biosynthesis in biofilm formation by nontypeable Haemophilus influenzae in the chinchilla middle ear. Infect. Immun. 73, 3210–3218 (2005).
Hong, W., Pang, B., West-Barnette, S. & Swords, W.E. Phosphorylcholine expression by nontypeable Haemophilus influenzae correlates with maturation of biofilm communities in vitro and in vivo. J. Bacteriol. 189, 8300–8307 (2007).
Rice, P.A., Yang, S., Mizuuchi, K. & Nash, H.A. Crystal structure of an IHF-DNA complex: a protein-induced DNA U-turn. Cell 87, 1295–1306 (1996).
American Academy of Pediatrics Subcommittee on Management of Acute Otitis Media. Diagnosis and management of acute otitis media. Pediatrics 113, 1451–1465 (2004).
Bakaletz, L.O., Leake, E.R., Billy, J.M. & Kaumaya, P.T. Relative immunogenicity and efficacy of two synthetic chimeric peptides of fimbrin as vaccinogens against nasopharyngeal colonization by nontypeable Haemophilus influenzae in the chinchilla. Vaccine 15, 955–961 (1997).
Bakaletz, L.O., Kennedy, B.J., Novotny, L.A., Duquesne, G., Cohen, J. & Lobet, Y. Protection against development of otitis media induced by nontypeable Haemophilus influenzae by both active and passive immunization in a chinchilla model of virus-bacterium superinfection. Infect. Immun. 67, 2746–2762 (1999).
Kennedy, B.J., Novotny, L.A., Jurcisek, J.A., Lobet, Y. & Bakaletz, L.O. Passive transfer of antiserum specific for immunogens derived from a nontypeable Haemophilus influenzae adhesin and lipoprotein D prevents otitis media after heterologous challenge. Infect. Immun. 68, 2756–2765 (2000).
Kyd, J.M., Cripps, A.W., Novotny, L.A. & Bakaletz, L.O. Efficacy of the 26-kilodalton outer membrane protein and two P5 fimbrin-derived immunogens to induce clearance of nontypeable Haemophilus influenzae from the rat middle ear and lungs as well as from the chinchilla middle ear and nasopharynx. Infect. Immun. 71, 4691–4699 (2003).
Novotny, L.A., Jurcisek, J.A., Pichichero, M.E. & Bakaletz, L.O. Epitope mapping of the outer membrane protein P5-homologous fimbrin adhesin of nontypeable Haemophilus influenzae. Infect. Immun. 68, 2119–2128 (2000).
Novotny, L.A. et al. Detection and characterization of pediatric serum antibody to the OMP P5-homologous adhesin of nontypeable Haemophilus influenzae during acute otitis media. Vaccine 20, 3590–3597 (2002).
Novotny, L.A. & Bakaletz, L.O. The fourth surface-exposed region of the outer membrane protein P5-homologous adhesin of nontypable Haemophilus influenzae is an immunodominant but nonprotective decoying epitope. J. Immunol. 171, 1978–1983 (2003).
Goodman, S.D. & Nash, H.A. Functional replacement of a protein-induced bend in a DNA recombination site. Nature 341, 251–254 (1989).
Goodman, S.D., Nicholson, S.C. & Nash, H.A. Deformation of DNA during site-specific recombination of bacteriophage lambda: replacement of IHF protein by HU protein or sequence-directed bends. Proc. Natl Acad. Sci. USA 89, 11910–11914 (1992).
Azam, T.A. & Ishihama, A. Twelve species of the nucleoid-associated protein from Escherichia coli. Sequence recognition specificity and DNA binding affinity. J. Biol. Chem. 274, 33105–33113 (1999).
Cooley, A.E. et al. DNA-binding by Haemophilus influenzae and Escherichia coli YbaB, members of a widely-distributed bacterial protein family. BMC Microbiol. 9, 137 (2009).
Oberto, J., Drlica, K. & Rouviere-Yaniv, J. Histones HMG HU IHF: meme combat. Biochimie 76, 901–908 (1994).
Berne, C., Kysela, D.T. & Brun, Y.V. A bacterial extracellular DNA inhibits settling of motile progeny cells within a biofilm. Mol. Microbiol. 77, 815–829 (2010).
Nakamura, K., Yahagi, S., Yamazaki, T. & Yamane, K. Bacillus subtilis histone-like protein HBsu, is an integral component of a SRP-like particle that can bind the Alu domain of small cytoplasmic RNA. J. Biol. Chem. 274, 13569–13576 (1999).
Rice, K.C. & Bayles, K.W. Molecular control of bacterial death and lysis. Microbiol. Mol. Biol. Rev. 72, 85–109, table of contents (2008).
Claverys, J.P. & Havarstein, L.S. Cannibalism and fratricide: mechanisms and raisons d’etre. Nat. Rev. Microbiol. 5, 219–229 (2007).
Cassone, A. & Rappuoli, R. Universal vaccines: shifting to one for many. MBio 1. pii: e00042-10 (2010).
Czerkinsky, C. & Holmgren, J. Mucosal delivery routes for optimal immunization: targeting immunity to the right tissues. Curr. Top. Microbiol. Immunol. PMID 21053117 (2010).
Otczyk, D. & Cripps, A.W. Mucosal immunization. Human Vaccines 6, 978–1006 (2010).
Novotny, L.A., Clements, J.D. & Bakaletz, L.O. Transcutaneous immunization as preventative and therapeutic regimens to protect against experimental otitis media due to nontypeable Haemophilus influenzae. Mucosal. Immunol. 4, 456–467 (2011).
Granston, A.E. & Nash, H.A. Characterization of a set of integration host factor mutants deficient for DNA binding. J. Mol. Biol. 234, 45–59 (1993).
Novotny, L.A., Jurcisek, J.A., Godfroid, F., Poolman, J.T., Denoel, P.A. & Bakaletz, L.O. Passive immunization with human anti-protein D antibodies induced by polysaccharide protein D conjugates protects chinchillas against otitis media after intranasal challenge with Haemophilus influenzae. Vaccine 24, 4804–4811 (2006).
Miyamoto, N. & Bakaletz, L.O. Kinetics of the ascension of NTHi from the nasopharynx to the middle ear coincident with adenovirus-induced compromise in the chinchilla. Microb. Pathog. 23, 119–126 (1997).
Harrison, A et al. Genomic sequence of an otitis media isolate of nontypeable Haemophilus influenzae: comparative study with H. influenzae serotype d, strain KW20. J. Bacteriol. 187, 4627–4636 (2005).
Flamm, E.L. & Weisberg, R.A. Primary structure of the hip gene of Escherichia coli and of its product, the beta subunit of integration host factor. J. Mol. Biol. 183, 117–128 (1985).
Blattner, F.R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Silhavy, T.J., Berman, M.L. & Engquist, L.W. Experiments with Gene Fusions ( Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, 1984 ).
Miller, H.I. Primary structure of the himA gene of Escherichia coli: homology with DNA-binding protein HU and association with the phenylalanyl-tRNA synthetase operon. Cold. Spring Harb. Symp. Quant. Biol 49, 691–698 (1984).
Oberto, J., Nabti, S., Jooste, V., Mignot, H. & Rouviere-Yaniv, J. The HU regulon is composed of genes responding to anaerobiosis, acid stress, high osmolarity and SOS induction. PLoS One 4, e4367 (2009).
Corcoran, C.P. & Dorman, C.J. DNA relaxation-dependent phase biasing of the fim genetic switch in Escherichia coli depends on the interplay of H-NS IHF and LRP. Mol. Microbiol. 74, 1071–1082 (2009).
Li, B., Smith, P., Horvath, D.J. Jr, Romesberg, F.E. & Justice, S.S. SOS regulatory elements are essential for UPEC pathogenesis. Microbes Infect. 12, 662–668 (2010).
Acknowledgements
We would like to dedicate this work in honor of Howard Nash, discoverer of IHF; he was an inspiration to us all. In addition, we also thank Dr Nash for his generous gifts of all his stocks of purified E. coli IHF and polyclonal antisera to E. coli IHF. We also thank the following individuals for their generous gifts: Emilia Chiancone, for both isolated E. coli DPS protein and antiserum directed against DPS; Jay Hinton, for monoclonal antibody (H113) directed against isolated E. coli H-NS; and Linda Kenney, for purified E. coli H-NS protein. Finally, we are grateful to both Peter Droge, for critical evaluation of our manuscript, and Jennifer Neelans for helping us prepare this manuscript. This work was supported by discretionary funds to both LOB and SDG.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Goodman, S., Obergfell, K., Jurcisek, J. et al. Biofilms can be dispersed by focusing the immune system on a common family of bacterial nucleoid-associated proteins. Mucosal Immunol 4, 625–637 (2011). https://doi.org/10.1038/mi.2011.27
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2011.27
This article is cited by
-
The morphology and metabolic changes of Actinobacillus pleuropneumoniae during its growth as a biofilm
Veterinary Research (2023)
-
Redirecting the immune response towards immunoprotective domains of a DNABII protein resolves experimental otitis media
npj Vaccines (2019)
-
Bio-enzymes for inhibition and elimination of Escherichia coli O157:H7 biofilm and their synergistic effect with sodium hypochlorite
Scientific Reports (2019)
-
Biofilm Formations in Pediatric Respiratory Tract Infection
Current Infectious Disease Reports (2019)
-
Skin Microbiota in Obese Women at Risk for Surgical Site Infection After Cesarean Delivery
Scientific Reports (2018)