Abstract
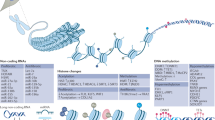
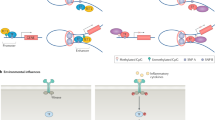
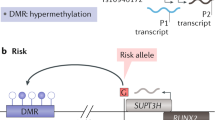
Genome-wide association studies have shown that genetic polymorphisms make a substantial but incomplete contribution to the risk of developing rheumatoid arthritis (RA). Efforts to understand the nongenetic contributions to RA disease susceptibility have recently focused on the study of epigenetic mechanisms, namely modifications of DNA and histones, which are subject to environmental influences and regulate gene expression. A surprising theme emerging from studies of the enzymes responsible for these epigenetic modifications, particularly histone deacetylases, is that they regulate inflammatory activation of cell populations relevant to RA through independent, direct, and dynamic interactions with nonhistone proteins. Herein, we highlight studies, the findings of which collectively suggest that revisiting the original definition of epigenetics, conceived some 70 years ago, might advance our interpretation of DNA and histone modifications with regard to gene expression and clinical outcome in RA. Such an approach could also facilitate the development of strategies to target these epigenetic modifications in the clinic.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bogdanos, D. P. et al. Twin studies in autoimmune disease: genetics, gender and environment. J. Autoimmun. 38, J156–J169 (2012).
Bax, M., van Heemst, J., Huizinga, T. W. & Toes, R. E. Genetics of rheumatoid arthritis: what have we learned? Immunogenetics 63, 459–466 (2011).
Knevel, R. et al. Genetic predisposition of the severity of joint destruction in rheumatoid arthritis: a population-based study. Ann. Rheum. Dis. 71, 707–709 (2012).
Ballestar, E. Epigenetic alterations in autoimmune rheumatic diseases. Nat. Rev. Rheumatol. 7, 263–271 (2011).
Huang, S. The molecular and mathematical basis of Waddington's epigenetic landscape: a framework for post-Darwinian biology? Bioessays 34, 149–157 (2012).
Waddington, C. H. The epigenotype. Endeavour 1, 18–20 (1942).
Russo V. E., Martienssen R. A. & Riggs A. D. (Eds) Epigenetic Mechanisms of Gene Regulation (Cold Spring Harbor Laboratory Press, Woodbury, 1996).
Bottini, N. & Firestein, G. S. Duality of fibroblast-like synoviocytes in RA: passive responders and imprinted aggressors. Nat. Rev. Rheumatol. 9, 24–33 (2013).
Ospelt, C., Reedquist, K. A., Gay, S. & Tak, P. P. Inflammatory memories: is epigenetics the missing link to persistent stromal cell activation in rheumatoid arthritis? Autoimmun. Rev. 10, 519–524 (2011).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Karouzakis, E. et al. DNA methylation regulates the expression of CXCL12 in rheumatoid arthritis synovial fibroblasts. Genes Immun. 12, 643–652 (2011).
Nakano, K., Whitaker, J. W., Boyle, D. L., Wang, W. & Firestein, G. S. DNA methylome signature in rheumatoid arthritis. Ann. Rheum. Dis. 72, 110–117 (2013).
Duroux-Richard, I., Jorgensen, C. & Apparailly, F. What do microRNAs mean for rheumatoid arthritis? Arthritis Rheum. 64, 11–20 (2012).
Gardner, K. E., Allis, C. D. & Strahl, B. D. Operating on chromatin, a colorful language where context matters. J. Mol. Biol. 409, 36–46 (2011).
Yun, M., Wu, J., Workman, J. L. & Li, B. Readers of histone modifications. Cell Res. 21, 564–578 (2011).
Horiuchi, M. et al. Expression and function of histone deacetylases in rheumatoid arthritis synovial fibroblasts. J. Rheumatol. 36, 1580–1589 (2009).
Huber, L. C. et al. Histone deacetylase/acetylase activity in total synovial tissue derived from rheumatoid arthritis and osteoarthritis patients. Arthritis Rheum. 56, 1087–1093 (2007).
Kawabata, T. et al. Increased activity and expression of histone deacetylase 1 in relation to tumor necrosis factor-α in synovial tissue of rheumatoid arthritis. Arthritis Res. Ther. 12, R133 (2010).
Grabiec, A. M., Tak, P. P. & Reedquist, K. A. Function of histone deacetylase inhibitors in inflammation. Crit Rev. Immunol. 31, 233–263 (2011).
Beier, U. H. et al. Histone deacetylases 6 and 9 and sirtuin-1 control FOXP3+ regulatory T cell function through shared and isoform-specific mechanisms. Sci. Signal. 5, ra45 (2012).
de Zoeten, E. F., Wang, L., Sai, H., Dillmann, W. H. & Hancock, W. W. Inhibition of HDAC9 increases T regulatory cell function and prevents colitis in mice. Gastroenterology 138, 583–594 (2010).
de Zoeten, E. F. et al. Histone deacetylase 6 and heat shock protein 90 control the functions of FOXP3+ T-regulatory cells. Mol. Cell Biol. 31, 2066–2078 (2011).
Li, B. et al. FOXP3 interactions with histone acetyltransferase and class II histone deacetylases are required for repression. Proc. Natl Acad. Sci. USA 104, 4571–4576 (2007).
van Loosdregt J. et al. Regulation of TREG functionality by acetylation-mediated FOXP3 protein stabilization. Blood 115, 965–974 (2010).
van Loosdregt, J. et al. Rapid temporal control of FOXP3 protein degradation by sirtuin-1. PLoS ONE 6, e19047 (2011).
Navarro, M. N., Goebel, J., Feijoo-Carnero, C., Morrice, N. & Cantrell, D. A. Phosphoproteomic analysis reveals an intrinsic pathway for the regulation of histone deacetylase 7 that controls the function of cytotoxic T lymphocytes. Nat. Immunol. 12, 352–361 (2011).
Barnes, P. J. Targeting the epigenome in the treatment of asthma and chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 6, 693–696 (2009).
Grabiec, A. M. et al. Histone deacetylase inhibitors suppress inflammatory activation of rheumatoid arthritis patient synovial macrophages and tissue. J. Immunol. 184, 2718–2728 (2010).
Niederer, F. et al. SIRT1 overexpression in the rheumatoid arthritis synovium contributes to proinflammatory cytokine production and apoptosis resistance. Ann. Rheum. Dis. 70, 1866–1873 (2011).
Gillespie, J. et al. Histone deacetylases are dysregulated in rheumatoid arthritis and a novel histone deacetylase 3-selective inhibitor reduces interleukin-6 production by peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Rheum. 64, 418–422 (2012).
Chen, X. et al. Requirement for the histone deacetylase HDAC3 for the inflammatory gene expression program in macrophages. Proc. Natl Acad. Sci. USA 109, E2865–E2874 (2012).
Mullican, S. E. et al. Histone deacetylase 3 is an epigenomic brake in macrophage alternative activation. Genes Dev. 25, 2480–2488 (2011).
Watson, P. J., Fairall, L., Santos, G. M. & Schwabe, J. W. Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature 481, 335–340 (2012).
Prinjha, R. K., Witherington, J. & Lee, K. Place your BETs: the therapeutic potential of bromodomains. Trends Pharmacol. Sci. 33, 146–153 (2012).
Nicodeme, E. et al. Suppression of inflammation by a synthetic histone mimic. Nature 468, 1119–1123 (2010).
Grabiec, A. M., Korchynskyi, O., Tak, P. P. & Reedquist, K. A. Histone deacetylase inhibitors suppress rheumatoid arthritis fibroblast-like synoviocyte and macrophage IL-6 production by accelerating mRNA decay. Ann. Rheum. Dis. 71, 424–431 (2012).
Lundh, M. et al. Histone deacetylases 1 and 3 but not 2 mediate cytokine-induced beta cell apoptosis in INS-1 cells and dispersed primary islets from rats and are differentially regulated in the islets of type 1 diabetic children. Diabetologia 55, 2421–2431 (2012).
Kruidenier, L. et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488, 404–408 (2012).
Vojinovic, J. et al. Safety and efficacy of an oral histone deacetylase inhibitor in systemic-onset juvenile idiopathic arthritis. Arthritis Rheum. 63, 1452–1458 (2011).
US National Library of Medicine. ClinicalTrials.gov [online] (2012).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Levy, D. et al. Lysine methylation of the NF-κB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-κB signaling. Nat. Immunol. 12, 29–36 (2011).
Gu, B. & Zhu, W. G. Surf the post-translational modification network of p53 regulation. Int. J. Biol. Sci. 8, 672–684 (2012).
Tanikawa, C. et al. Regulation of histone modification and chromatin structure by the p53–PADI4 pathway. Nat. Commun. 3, 676 (2012).
Liu, X. et al. Methyltransferase Set7/9 regulates p53 activity by interacting with Sirtuin 1 (SIRT1). Proc. Natl Acad. Sci. USA 108, 1925–1930 (2011).
Ptashne, M. On the use of the word 'epigenetic'. Curr. Biol. 17, R233–R236 (2007).
Bird, A. Perceptions of epigenetics. Nature 447, 396–398 (2007).
Das, J. et al. Digital signaling and hysteresis characterize Ras activation in lymphoid cells. Cell 136, 337–351 (2009).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Author information
Authors and Affiliations
Contributions
A. M. Grabiec and K. A. Reedquist contributed equally to all stages of the preparation of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Postulated mechanism of HDAC inhibitor action in animal models of arthritis (DOC 54 kb)
Supplementary Table 2
Biological roles of HDAC isoforms in cells relevant to RA pathology (DOC 58 kb)
Rights and permissions
About this article
Cite this article
Grabiec, A., Reedquist, K. The ascent of acetylation in the epigenetics of rheumatoid arthritis. Nat Rev Rheumatol 9, 311–318 (2013). https://doi.org/10.1038/nrrheum.2013.17
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2013.17
This article is cited by
-
Epi-revolution in rheumatology: the potential of histone deacetylase inhibitors for targeted rheumatoid arthritis intervention
Inflammopharmacology (2024)
-
Signaling pathways in rheumatoid arthritis: implications for targeted therapy
Signal Transduction and Targeted Therapy (2023)
-
Inhibition of histone deacetylase 6 suppresses inflammatory responses and invasiveness of fibroblast-like-synoviocytes in inflammatory arthritis
Arthritis Research & Therapy (2021)
-
Epigenetic regulation of inflammation in periodontitis: cellular mechanisms and therapeutic potential
Clinical Epigenetics (2020)
-
SAMD9 is a (epi-) genetically regulated anti-inflammatory factor activated in RA patients
Molecular and Cellular Biochemistry (2019)