Abstract
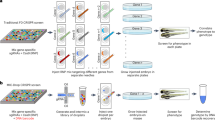
The zebrafish is a popular model organism for studying development and disease, and genetically modified zebrafish provide an essential tool for functional genomic studies. Numerous publications have demonstrated the efficacy of gene targeting in zebrafish using CRISPR/Cas9, and they have included descriptions of a variety of tools and methods for guide RNA synthesis and mutant identification. However, most of the published techniques are not readily scalable to increase throughput. We recently described a CRISPR/Cas9-based high-throughput mutagenesis and phenotyping pipeline in zebrafish. Here, we present a complete workflow for this pipeline, including target selection; cloning-free single-guide RNA (sgRNA) synthesis; microinjection; validation of the target-specific activity of the sgRNAs; founder screening to identify germline-transmitting mutations by fluorescence PCR; determination of the exact lesion by Sanger or next-generation sequencing (including software for analysis); and genotyping in the F1 or subsequent generations. Using these methods, sgRNAs can be evaluated in 3 d, zebrafish germline-transmitting mutations can be identified within 3 months and stable lines can be established within 6 months. Realistically, two researchers can target tens to hundreds of genes per year using this protocol.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Welter, D. et al. The NHGRI GWAS catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Bamshad, M.J. et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat. Rev. Genet. 12, 745–755 (2011).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Kettleborough, R.N. et al. A systematic genome-wide analysis of zebrafish protein-coding gene function. Nature 496, 494–497 (2013).
Varshney, G.K. et al. A large-scale zebrafish gene knockout resource for the genome-wide study of gene function. Genome Res. 23, 727–735 (2013).
Quach, H.N. et al. A multifunctional mutagenesis system for analysis of gene function in zebrafish. G3 (Bethesda) 5, 1283–1299 (2015).
Varshney, G.K. & Burgess, S.M. Mutagenesis and phenotyping resources in zebrafish for studying development and human disease. Brief Funct. Genomics 13, 82–94 (2014).
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat. Biotechnol. 26, 702–708 (2008).
Meng, X., Noyes, M.B., Zhu, L.J., Lawson, N.D. & Wolfe, S.A. Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat. Biotechnol. 26, 695–701 (2008).
Bedell, V.M. et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491, 114–118 (2012).
Huang, P. et al. Heritable gene targeting in zebrafish using customized TALENs. Nat. Biotechnol. 29, 699–700 (2011).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Hwang, W.Y. et al. Efficient genome editing in zebrafish using a CRISPR–Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Jao, L.E., Wente, S.R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl. Acad. Sci. USA 110, 13904–13909 (2013).
Varshney, G.K. et al. High-throughput gene targeting and phenotyping in zebrafish using CRISPR/Cas9. Genome Res. 25, 1030–1042 (2015).
Varshney, G.K., Sood, R. & Burgess, S.M. Understanding and editing the zebrafish genome. Adv. Genet. 92, 1–52 (2015).
LaFave, M.C., Varshney, G.K., Vemulapalli, M., Mullikin, J.C. & Burgess, S.M. A defined zebrafish line for high-throughput genetics and genomics: NHGRI-1. Genetics 198, 167–170 (2014).
Carrington, B., Varshney, G.K., Burgess, S.M. & Sood, R. CRISPR-STAT: an easy and reliable PCR-based method to evaluate target-specific sgRNA activity. Nucleic Acids Res. 43, e157 (2015).
Varshney, G.K. et al. CRISPRz: a database of zebrafish validated sgRNAs. Nucleic Acids Res. 44, D822–D826 (2016).
Guryev, V. et al. Genetic variation in the zebrafish. Genome Res. 16, 491–497 (2006).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Sood, R. et al. Efficient methods for targeted mutagenesis in zebrafish using zinc-finger nucleases: data from targeting of nine genes using CompoZr or CoDA ZFNs. PLoS One 8, e57239 (2013).
Xu, H. et al. Sequence determinants of improved CRISPR sgRNA design. Genome Res. 25, 1147–1157 (2015).
Moreno-Mateos, M.A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Qin, W. et al. Expansion of CRISPR/Cas9 genome targeting sites in zebrafish by Csy4-based RNA processing. Cell Res. 25, 1074–1077 (2015).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Kim, Y., Kweon, J. & Kim, J.S. TALENs and ZFNs are associated with different mutation signatures. Nat. Methods 10, 185 (2013).
Brownstein, M.J., Carpten, J.D. & Smith, J.R. Modulation of non-templated nucleotide addition by Taq DNA polymerase: primer modifications that facilitate genotyping. Biotechniques 20, 1004-6–1008-10 (1996).
Hill, J.T. et al. Poly peak parser: method and software for identification of unknown indels using Sanger sequencing of polymerase chain reaction products. Dev. Dyn. 243, 1632–1636 (2014).
Acknowledgements
We thank J. Fekecs and U. Harper for help with figures, and D. Prebilic, H. Mahon and other staff of the National Institutes of Health zebrafish facility for excellent animal care. This research is funded by the Intramural Research Program of the National Human Genome Research Institute; National Institutes of Health (S.M.B. and R.S.).
Author information
Authors and Affiliations
Contributions
G.K.V., B.C., R.S. and S.M.B. developed the protocol and wrote the manuscript, with inputs from all other authors; G.K.V., B.C., W.P., K.B., C.F., L.X., M.P.J. and J.L. performed the experiments. M.C.L. and Z.C. wrote and tested the bioinformatics pipeline, ampliconDIVider.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
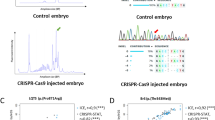
Supplementary Figure 1 Screen capture of the zebrafish CRISPR/Cas9 track hub.
This is a screen capture of the first exon of zebrafish FGF2. Most of the ZebrafishGenomics tracks are displayed. The top track shows all computationally predicted SpCas9 target sites. They are color coded by target length (gray=20, blue=19, orange=18), with darker colors having fewer predicted off-target sequences. The sequences can be seen in the left column and the predicted off-target number can be seen at the end of the sequence. The “GA CRISPR/Cas9 Targets” track shows targets that can be synthesized by Sp6 and the “GG CRISPR/Cas9 Targets” are guides that can be synthesized by T7. The black bars in the “NHGRI-1 Invariant” track indicate where the genomic sequence of the NHGRI-1 strain matches the reference sequence. Gaps represent variation. The “NHGRI-1 Variants” track shows possible SNV’s at the given position. Researchers should avoid regions with gaps or SNV’s. Green arrowhead indicates what we would consider the best target for this region.
Supplementary information
Supplementary Tables 1–3
Supplementary Table 1. sgRNA target sequences, primer sequences and amplicon sizes used to generate the data in Figures 5 and 6. Supplementary Table 2. Module manager settings for a 3130xl or 3730 capillary sequencer using a 36-cm or 50-cm array. Injection times marked in red are the variable settings between the CRISPRSTAT and fragment analysis methods. Supplementary Table 3. Barcoded M13 primer sequences. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Varshney, G., Carrington, B., Pei, W. et al. A high-throughput functional genomics workflow based on CRISPR/Cas9-mediated targeted mutagenesis in zebrafish. Nat Protoc 11, 2357–2375 (2016). https://doi.org/10.1038/nprot.2016.141
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.141
This article is cited by
-
Improved selection of zebrafish CRISPR editing by early next-generation sequencing based genotyping
Scientific Reports (2023)
-
A CRISPR-Cas9-mediated F0 screen to identify pro-regenerative genes in the zebrafish retinal pigment epithelium
Scientific Reports (2023)
-
Large-scale F0 CRISPR screens in vivo using MIC-Drop
Nature Protocols (2023)
-
A second locus contributing to the differential expression of the blue sensitive opsin SWS2A in Lake Malawi cichlids
Hydrobiologia (2023)
-
Diagnostic applications and therapeutic option of Cascade CRISPR/Cas in the modulation of miRNA in diverse cancers: promises and obstacles
Journal of Cancer Research and Clinical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.