Abstract
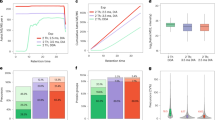
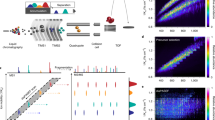
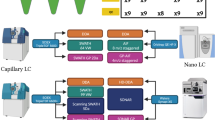
We present a data-independent acquisition mass spectrometry method, ultradefinition (UD) MSE. This approach utilizes ion mobility drift time-specific collision-energy profiles to enhance precursor fragmentation efficiency over current MSE and high-definition (HD) MSE data-independent acquisition techniques. UDMSE provided high reproducibility and substantially improved proteome coverage of the HeLa cell proteome compared to previous implementations of MSE, and it also outperformed a state-of-the-art data-dependent acquisition workflow. Additionally, we report a software tool, ISOQuant, for processing label-free quantitative UDMSE data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aebersold, R. & Mann, M. Nature 422, 198–207 (2003).
Geromanos, S.J. et al. Proteomics 9, 1683–1695 (2009).
Michalski, A., Cox, J. & Mann, M. J. Proteome Res. 10, 1785–1793 (2011).
Kelstrup, C.D., Young, C., Lavallee, R., Nielsen, M.L. & Olsen, J.V. J. Proteome Res. 11, 3487–3497 (2012).
Venable, J.D., Dong, M., Wohlschlegel, J., Dillin, A. & Yates, J.R. Nat. Methods 1, 39–45 (2004).
Geiger, T., Cox, J. & Mann, M. Mol. Cell. Proteomics 9, 2252–2261 (2010).
Panchaud, A., Jung, S., Shaffer, S.A . Aitchison, J.D. & Goodlett, D.R. Anal. Chem. 83, 2250–2257 (2011).
Gillet, L.C. et al. Mol. Cell. Proteomics 11, O111.016717 (2012).
Silva, J.C. et al. Anal. Chem. 77, 2187–2200 (2005).
Lee, S. et al. Int. J. Mass Spectrom. 309, 154–160 (2012).
Valentine, S.J. et al. J. Proteome Res. 10, 2318–2329 (2011).
Baker, E.S. et al. J. Proteome Res. 9, 997–1006 (2010).
Geromanos, S.J., Hughes, C., Ciavarini, S., Vissers, J.P.C. & Langridge, J.I. Anal. Bioanal. Chem. 404, 1127–1139 (2012).
Maclean, B. et al. Anal. Chem. 82, 10116–10124 (2010).
Cox, J. & Mann, M. Nat. Biotechnol. 26, 1367–1372 (2008).
Serang, O. & Noble, W. Stat. Interface 5, 3–20 (2012).
Silva, J.C., Gorenstein, M.V., Li, G.-Z., Vissers, J.P.C. & Geromanos, S.J. Mol. Cell. Proteomics 5, 144–156 (2006).
Vizcaíno, J.A. et al. Nucleic Acids Res. 41, D1063–D1069 (2013).
Wiśniewski, J.R., Zougman, A., Nagaraj, N. & Mann, M. Nat. Methods 6, 359–362 (2009).
Tenzer, S. et al. ACS Nano 5, 7155–7167 (2011).
Giles, K. et al. Rapid Commun. Mass Spectrom. 18, 2401–2414 (2004).
MacLean, B. et al. Bioinformatics 26, 966–968 (2010).
Podwojski, K. et al. Bioinformatics 25, 758–764 (2009).
Sakoe, H. & Chiba, S. Inst. Electr. Comm. Eng. Japan 136 (1970).
Ester, M., Kriegel, H.P., Sander, J. & Xu, X. in 2nd Int. Conf. Knowl. Discov. Data Min. (eds. Simoudis, E., Han, J. & Fayyad, U.) 226–231 (AAAI Press, 1996).
Li, G.-Z. et al. Proteomics 9, 1696–1719 (2009).
Stanley, J.R. et al. Anal. Chem. 83, 6135–6140 (2011).
Cleveland, W.S. J. Am. Stat. Assoc. 74, 829 (1979).
Nesvizhskii, A.I., Keller, A., Kolker, E. & Aebersold, R. Anal. Chem. 75, 4646–4658 (2003).
Acknowledgements
We thank R. Spohrer for excellent sample preparation; W. Thompson and K. Giles for discussions on ion-mobility separations and data-independent acquisition methods; A. Savidor and A. Gabashvili for assistance in the DDA analysis; all ISOQuant beta testers for their continuing critical evaluation of the software; and H. Vissers and K. Richardson for discussions on data evaluation. This work was supported by grants from the Deutsche Forschungsgemeinschaft (INST 371/23-1 FUGG) to S.T., H.S. and the BMBF (e:Bio Express2Present, 0316179C) to S.T., as well by as the Forschungszentrum Immunologie (FZI), the Naturwissenschaftlich-Medizinische Forschungszentrum (NMFZ) and the Focus Program Translational Neurosciences (FTN) of the Johannes Gutenberg University Mainz.
Author information
Authors and Affiliations
Contributions
U.D. and S.T. designed and performed the DIA experiments. Y.L. performed the DDA experiments and DDA data evaluation. J.K. designed and wrote the software. J.K. and S.T. developed algorithms. U.D., J.K., P.N. and S.T. analyzed the DIA data. U.D., J.K., H.S. and S.T. contributed to overall design of the project. All authors prepared and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1–4 and 9, and Supplementary Notes 1 and 2 (PDF 5277 kb)
Supplementary Table 5
List of identified proteins in tryptic HeLa cell lysate after analysis with MSE, HDMSE and UDMSE. (XLSX 4155 kb)
Supplementary Table 6
MaxQuant report of the DDA-analysis of tryptic HeLa cell lysate. (XLSX 4175 kb)
Supplementary Table 7
ISOQuant quantification Excel report for hybrid proteome sample using high confidence settings. (XLSX 1820 kb)
Supplementary Table 8
High confidence and MaxID settings for ISOQuant processing. (XLSX 11 kb)
Supplementary Software
Supplementary Software (ZIP 30644 kb)
Rights and permissions
About this article
Cite this article
Distler, U., Kuharev, J., Navarro, P. et al. Drift time-specific collision energies enable deep-coverage data-independent acquisition proteomics. Nat Methods 11, 167–170 (2014). https://doi.org/10.1038/nmeth.2767
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2767
This article is cited by
-
SigB modulates expression of novel SigB regulon members via Bc1009 in non-stressed and heat-stressed cells revealing its alternative roles in Bacillus cereus
BMC Microbiology (2023)
-
Prediction of peptide mass spectral libraries with machine learning
Nature Biotechnology (2023)
-
Physiological, epigenetic, and proteomic responses in Pfaffia glomerata growth in vitro under salt stress and 5-azacytidine
Protoplasma (2023)
-
The inhibition of putrescine synthesis affects the in vitro shoot development of Cedrela fissilis Vell. (Meliaceae) by altering endogenous polyamine metabolism and the proteomic profile
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
Microporous membrane and culture medium affect in vitro seedling development of Dalbergia nigra (Vell.) Ex Benth. (Fabaceae) by modulation of the protein profile and accumulation of ethylene and CO2
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)