Abstract
Pericentrin, a conserved centrosomal component, provides the structural scaffold to anchor numerous centrosomal proteins, and thus plays an essential role in the organization and function of the centrosome and the mitotic spindle. Although pericentrin was shown to localize in the cytoplasm and reported to be sensitive to leptomycin B (LMB), a specific inhibitor of Crm1, the regions within pericentrin that serve as signals for transporting in and out of the nucleus have not yet been identified. In this study, we identified five novel nuclear export signals (NESs) in pericentrin with diverse export activities. All of the five NESs could bind to Crm1 in a LMB-sensitive way when mediating the nuclear export of pericentrin. We also demonstrated that the region of amino acids 8-42 in pericentrin contains a tripartite nuclear localization signal (NLS) consisting of three clusters of basic amino acids. The NLS of pericentrin binds to importin β directly or via the adaptor importin α to form the import complex, which could be disrupted by RanQ69L, a dominant-negative Ran GTPase possessing high affinity for importin β. Furthermore, we found that mutation of the NESs in full-length pericentrin results in both nuclear and cytoplasmic localization, and mutation of the NLS abolishes the nuclear import of pericentrin. On the basis of these results, we suggest that the NESs and NLS of pericentrin are essential for its subcellular localization and nucleocytoplasmic trafficking during the cell cycle.
Similar content being viewed by others
Introduction
Pericentrin is a conserved component of the pericentriolar matrix that supplies the structural scaffold for anchoring numerous proteins and plays a crucial role in organization and function of centrosomes and mitotic spindles 1, 2, 3. It targets centrosomes via its PACT domain, and is located at the centrosomes throughout the whole cell cycle 4. Similar to the yeast homolog, Spc110p, pericentrin anchors γ-tubulin ring complex to centrosomes through interaction with γ-tubulin complex proteins 2 and 3, providing the microtubule nucleation sites and mediating the spindle organization in mitotic cells 5, 6. Several other proteins including cAMP-dependent protein kinase (PKA), protein kinase C (PKC) and disrupted-In-schizophrenia 1 (DISC-1) are also recruited to centrosomes by pericentrin 7, 8, 9. Moreover, pericentrin could form functional complex with pericentriolar material 1 (PCM1) at centrosomes 10.
Recent studies have revealed new functions of pericentrin in the maintenance of cell cycle integrity and regulation of DNA damage checkpoint. It was indicated that mutations in the gene PCNT result in Majewski osteodysplastic primordial dwarfism type II (MOPD II) and Seckel syndrome, two diseases in humans that are characterized by short stature and small brain size 11, 12, 13. It was suggested that the centrosome abnormality caused by PCNT mutations would be the main reason for cell growth restriction in MOPD II and Seckel syndrome. However, besides reduction or loss of centrosomal pericentrin and dysfunction of mitotic spindles, missegregation of chromosomes and defects in DNA damage signaling pathway are also present in patient cells, suggesting a link between pericentrin and the nucleus. Although there is no direct evidence to show that pericentrin is active in the nucleus, the finding of the interaction between pericentrin and chromodomain helicase DNA-binding protein 3/4 (CHD3/4), components of the multiprotein nucleosome remodeling deacetylase complex, suggests that pericentrin might have a nuclear function 14. A recent study has found that pericentrin could accumulate into the nucleus after the treatment of leptomycin B (LMB), a specific inhibitor of Crm1-mediated nuclear export 15. However, whether pericentrin contains functional nuclear export signals (NESs) and nuclear localization signals (NLSs), and how pericentrin shuttles between the nucleus and the cytoplasm have not been addressed.
Nucleocytoplasmic trafficking is an efficient mechanism to control the subcellular localization of proteins, and regulates cell proliferation and tumorigenesis 16. Proteins with masses that exceed 40 kD shuttle between the nucleus and cytoplasm in an active, signal-mediated pathway. Nuclear entry of a protein is determined by the presence of NLSs that interact with importin β directly or via the adaptor importin α for recognition 17. The NLS sites typically contain clusters of positively charged basic amino acids of lysine (K) and/or arginine (R) 17. Most of NLSs consist of either one (monopartite) or two (bipartite) stretches of basic amino acids (e.g., 126PKKKRRV132 in the SV40 large T-antigen for monopartite NLS and 155KRPAATKKAGQAKKKK170 in nucleoplasmin for bipartite NLS) 18, 19. Few proteins, like the epidermal growth factor receptor (EGFR), contain unusual tripartite NLS that has three clusters of basic amino acids 20. Conversely, nuclear exit of proteins is facilitated by the presence of NESs, which are characterized by the presence of leucines and other hydrophobic residues 21. The nuclear export process depends on the nuclear export receptor Crm1 to recognize NES sequences of proteins 21. Both nuclear import and export of proteins are regulated by the Ran GTPase 22.
In this study, we identified an unusual tripartite NLS and five classical NESs in pericentrin. We demonstrated that the NLS and NESs of pericentrin are essential for its cytoplasmic localization and nucleocytoplasmic shuttling during the cell cycle.
Results
The nucleocytoplasmic trafficking of pericentrin is mediated by Crm1 and importin β under the regulation of Ran GTPase
A previous study has shown that pericentrin was sensitive to LMB, a specific inhibitor of Crm1, and accumulated in the nucleus in LMB-treated cells 15. To confirm that pericentrin is a nucleocytoplasmic shuttling protein, we treated HeLa cells with 10 nM LMB for 6 h, and investigated the subcellular localization of pericentrin before and after LMB treatment. The immunofluorescence microscopy results showed that pericentrin predominantly localized in the cytoplasm with a strong focus on centrosomes in HeLa cells, and that the nuclear pericentrin was increased greatly after treatment with LMB, supporting the notion that pericentrin is exported from the nucleus to the cytoplasm in a Crm1-mediated pathway (Figure 1A). Since pericentrin is a large protein, which could not diffuse from the cytoplasm to nucleus passively, we proposed that pericentrin enters the nucleus by active nuclear import. To address this hypothesis, we immunoprecipitated endogenous pericentrin from HeLa cell lysates using the anti-pericentrin antibody and analyzed the pericentrin-binding proteins by western blot for the export receptor Crm1, the nuclear import receptor importin β and the regulation factor of nucleocytoplasmic transport Ran GTPase. The results showed that Crm1, importin β and Ran GTPase were all co-immunoprecipitated with pericentrin (Figure 1B), indicating that pericentrin was able to bind Crm1 and importin β in vivo under the regulation of Ran GTPase. To further confirm these results, we performed the GST pull-down assays using both GST-Crm1 and GST-importin β proteins in the lysates of the HA-pericentrin-expressing HeLa cells, and found that both GST-Crm1 and GST-importin β could specifically pull-down HA-pericentrin in the cell lysates (Figure 1C). Taken together, these data indicate that pericentrin shuttles between the cytoplasm and the nucleus, mediated by the nuclear transport receptors Crm1 and importin β under the regulation of Ran GTPase.
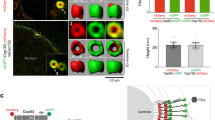
Pericentrin shuttles between the nucleus and cytoplasm. (A) Pericentrin is sensitive to LMB treatment. HeLa cells were treated with 10 nM LMB for 6 h before fixing with paraformaldehyde, and immunostained with anti-pericentrin antibody (green). The nuclei were stained with DAPI (blue). Bar, 20 μm. (B) Pericentrin binds to Crm1, importin β and Ran in vivo. HeLa cells were lysed with 1% NP-40-TB buffer and centrifuged for the supernatant. The cell lysates were incubated with protein A beads immobilized with anti-pericentrin antibody or rabbit IgG as a control at 4 °C for 2 h. The bound proteins on beads were analyzed by western blot using anti-pericentrin, Crm1, importin β and Ran antibodies. (C) Pericentrin binds to GST-Crm1 and GST-importin β in vitro. HeLa cells were transfected with HA-pericentrin for 48 h and then lysed using 0.5% NP-40-TB buffer. The cell lysates were incubated with GST (as a control), GST-Crm1 and GST-importin β beads at 4 °C for 2 h. The bound proteins on beads were analyzed by western blot using anti-HA antibody.
Identification of multiple NESs in pericentrin
Active nuclear export of proteins is usually mediated by NESs. A classical NES is a sequence consisting of about four hydrophobic amino acids (usually leucines), with a defined spacing between them 23. In searching for the potential NESs in pericentrin, we analyzed the amino acid sequence of human pericentrin and probed eight candidate NESs (cNESs) in pericentrin (Table 1). To test whether these sequences are functional NESs, we constructed a series of pericentrin fragments fused with the GFP tag, as shown in the schematic diagrams, and observed their distribution in cells before and after LMB treatment (Figure 2A). Because cNES2 and cNES3, cNES4 and cNES5 are closely located, we tested them in one single fragment, respectively (Figure 2A). To ensure that the cytoplasmic localization of the fragments was caused by their cNESs instead of other sequences, we also constructed a series of ΔcNES fragments as control (Figure 2A). Since a protein larger than 40 kD would be difficult to diffuse through nuclear envelope freely, we limited the truncated proteins with GFP tag within 40 kD to allow them diffuse freely if they do not contain special localization sequences. Our results showed that the fragments PCNT1-268 (containing cNES1), PCNT1146-1302(containing cNES4 and cNES5), PCNT1420-1558 (containing cNES6) and PCNT1596-1764 (containing cNES7) exhibited specifically cytoplasmic distribution, and relocated into the nucleus when cells were treated with LMB (Figure 2B); in contrast, the fragments PCNT997-1158 (containing cNES2 and cNES3) and PCNT1761-1880 (containing cNES8) distributed throughout the whole cells, regardless of the LMB treatment (Figure 2B and 2C). To test whether only the fragments containing functional NESs were bound to Crm1, we performed a pull-down assay using GST-Crm1 protein in the lysates of HeLa cells expressing the cNESs fragments. As expected, we found that only PCNT1-268, PCNT1146-1302, PCNT1420-1558 and PCNT1596-1764 could be pulled down by GST-Crm1 (Figure 2D), suggesting that these fragments were exported from the nucleus to the cytoplasm through interacting with Crm1. Taken together, these data indicate that cNES1, cNES4, cNES5, cNES6 and cNES7, which appeared sensitive to LMB, would be functional NESs mediated by Crm1, while cNES2, cNES3 and cNES8 do not function as NESs in the nuclear export.
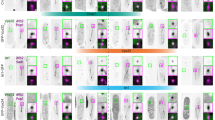
Identification of functional NESs in pericentrin. (A) Schematic diagrams of pericentrin truncated mutants. (B and C) The subcellular localization of the truncated mutants in the absence and presence of LMB. HeLa cells were transfected with the GFP-tagged truncated mutants. After 24 h, the cells transfected with truncated mutants containing the cNESs were treated with 10 nM LMB for 3 h. All the cells were fixed using paraformaldehyde for observation. The percentages of cells with predominantly nuclear (N>C), nuclear and cytoplasmic (N=C) or cytoplasmic (N<C) localization were depicted. At least 200 cells were evaluated for each sample. Data were averages of three independent experiments shown as mean ± SD. Bar, 20 μm. (D) GST-Crm1 pulls down the truncated mutants of pericentrin containing NESs specifically. HeLa cells were transfected with a series of truncated mutants of pericentrin showed in A for 24 h, and lysed with 0.5% NP-40-TB buffer. The cell lysates were incubated with GST-Crm1 beads for 2 h at 4 °C. The beads were washed four times with TB buffer and the bound proteins were analyzed by western blot using anti-GFP antibody.
Characterization of the cNESs in pericentrin
Next, we examined the nuclear export activities of the functional NESs in pericentrin through the in vivo nuclear export assay. The nuclear export assay using NES-deficient vector Rev1.4-GFP was reported previously and has been recognized to be an effective method to test the nuclear export activity of functional NESs 24. To confirm that the nuclear export assay was doable, we first used it to test seven defined NESs whose export activities are known, including NESs from MAPKK, RanBP1, c-ABL, FMRP, IκB, hdm2 and p53. The results showed that the NESs of MAPKK and RanBP1 have the strongest nuclear export activity (score 9+ and 8+/7+, respectively); the NESs of c-ABL and FMRP have a medium export activity (score 7+/6+ and 5+/4+, respectively), whereas the NESs of IκB, hdm2 and p53 have a relatively weak export activity (score 3+/2+, 2+ and 1+, respectively) (Supplementary information, Figure S1). Our data were consistent with previous findings 24 and indicated the validity of the nuclear export assay.
To test the nuclear export activity of pericentrin cNESs, we inserted the cNESs containing short segments of pericentrin into pRev1.4-GFP vector and expressed the fusion proteins in HeLa cells first (Figure 3A). We found that the insertion of cNES7 resulted in a majority of cytoplasmic relocation of Rev 1.4-GFP proteins except for some nucleolus accumulation, whereas cNES1, cNES4, cNES5 and cNES6 showed progressively weaker activity (Figure 3B). To observe the nuclear export activity of these cNESs more clearly, we treated the transfected cells with actinomycin D (Act.D) to prevent the nuclear import process of Rev1.4-cNES proteins. The results showed that the cNES7-Rev1.4-GFP in nucleolus was greatly reduced after Act.D treatment, indicating that cNES7 has strong export activity, while the fusion proteins of Rev1.4-GFP with cNES1, cNES4, cNES5 and cNES6 were more or less situated in the cytoplasm after treatment, suggesting that their export activities are relatively weak (Figure 3B). Additionally, when the cells were treated with LMB, all the fusion proteins of Rev1.4-GFP with cNESs were retained in the nucleus, indicating that all of them were sensitive to LMB treatment (Figure 3B). By comparing with the scoring system standard, we determined that cNES7 (residues 1 745 to 1 755) is a relatively strong NES (scoring 7+), whereas cNES1 (residues 245 to 254), cNES4 (residues 1 158 to 1 168), cNES5 (residues 1 169 to 1 179) and cNES6 (residues 1 540 to 1 548) are weak (scoring 3+, 2+, 2+/1+ and 1+, respectively) but functional NESs. To further confirm these identified NESs, we performed a mutation analysis. As the last two hydrophobic residues in NES sequences are generally the most conserved and most important ones for the nuclear export activity, these two residues of Rev1.4-GFP fusion proteins with cNESs were replaced by alanines, and the localization of these cNES-mutant proteins was tested in the nuclear export assay (Figure 3A). As expected, these cNES-mutant proteins localized in the nucleus even when the cells were treated with Act.D (Figure 3C), indicating that their NES functions were abolished by the mutation.
Analysis of the export activities of pericentrin NESs. (A) The sequences of the tested cNESs and their point mutations that were inserted into pRev (1.4)-GFP. The hydrophobic amino acids were shown in bold and the mutated sites were shown in red. (B) The cNESs of pericentrin exhibit different nuclear export activities. The cNESs of pericentrin were cloned into vector pRev (1.4)-GFP. HeLa cells were transfected with vectors for 24 h, and then treated with 5 μg/ml Act.D or with 10 nM LMB for 3 h. Bar, 20 μm. (C) The point mutations of cNESs lose the nuclear export activities. The mutants of cNESs were cloned into vector pRev (1.4)-GFP. HeLa cells were transfected with wild-type and mutants of cNESs for 24 h, and then treated with 5 μg/ml Act.D for 3 h. The subcellular localizations of these mutants were examined and compared with the wild-type NESs. Bar, 20 μm.
Taken together, we demonstrated that pericentrin contains five functional nuclear export sequences distributed in its N-terminal and middle regions with various nuclear export activities.
A nuclear localization signal locates at the N-terminus of pericentrin
In studying the NESs of pericentrin, we noticed that more than 80% of GFP-PCNT1-235 protein localized in nucleus, while the GFP-PCNT1-268 protein, which contains an NES sequence (cNES1), remained in the cytoplasm (Figure 2B), indicating that the N-terminus of pericentrin contains a NLS. We analyzed the sequence of pericentrin and found that the residues from 8 to 42 are rich in lysines and arginines (Table 1). To test whether it is a potential NLS, we constructed a few GFP-tagged truncated mutants within N-terminus of pericentrin and investigated their nuclear import activities in HeLa cells (Figure 4A). The data showed that, distinct from GFP-PCNT50-235, which distributed throughout the whole transfected cells, both GFP-PCNT1-235 and GFP-PCNT1-50 accumulated in the nucleus of the transfected cells (Figure 4B and 4C), indicating that the amino acid residues from 8 to 42 are responsible for the nuclear localization. In contrast, GFP-PCNT1-268, which contains an NES, showed cytoplasmic localization (Figure 4B and 4C). Since the nuclear import process is usually mediated by the nuclear import receptor importin β, we further tested the interaction of pericentrin NLS with importin β. Sepharose beads immobilized with GST-importin β protein were incubated with cell lysates from HeLa cells transfected with GFP, GFP-PCNT1-50, GFP-PCNT50-235 or GFP-PCNT1-235, respectively. The beads were isolated and analyzed by western blot with anti-GFP antibody. We found that GFP-PCNT1-50 and GFP-PCNT1-235 could efficiently bind importin β, while GFP-PCNT50-235 and the control GFP did not have any affinity for this nuclear import receptor (Figure 4D). These results suggest that pericentrin has a functional NLS located in its N-terminal region.
Identification of the NLS in the N-terminus of pericentrin. (A) The map for amino acids sequence of the NLS in pericentrin and schematic diagrams of truncated mutants within the N-terminus of human pericentrin. The positively charged amino acids lysine and arginine are shown in red. (B) The subcellular localizations of pericentrin-truncated mutants. HeLa cells were transfected with pericentrin-truncated mutants. After 24 h, cells were fixed for observation. Bar, 20 μm. (C) The nuclear accumulation percentages of pericentrin-truncated mutants were analyzed. At least 30 cells were evaluated for each sample. Data were averages of three independent experiments, shown as mean ± SD. (D) Importin β binds to GFP-PCNT1-50 and GFP-PCNT1-235, but does not have any affinity with GFP or GFP-PCNT50-235. HeLa cells were transfected with GFP, GFP-PCNT1-50, GFP-PCNT50-235 and GFP-PCNT1-235 for 24 h. The cells were lysed to form suspension and mixed with GST Sepharose beads with immobilized importin β for 2 h at 4 °C. After washing four times, the proteins that were bound to beads were eluted and analyzed by western blot using anti-GFP antibody.
As the amino acid sequence from 8 to 42 consists of three basic amino acids-rich clusters, which are amino acids 8 to 11, 16 to 28 and 36 to 42 (Figure 4A), we decided to check which cluster is crucial for the nuclear localization through expressing the truncated mutants in cells. The results showed that both GFP-PCNT8-28 and GFP-PCNT23-42 were located in nucleus efficiently, as observed for PCNT1-50, but the shorter truncated mutants PCNT6-13, PCNT14-30 and PCNT34-44 appeared to be less distributed in the nucleus, indicating that pericentrin NLS needs at least two of the three clusters to localize the protein in the nucleus.
Functional analysis of pericentrin NLS
The classical NLSs are typically composed of several positively charged amino acids, but not all of these amino acids contribute equally to the NLS function. To further investigate which amino acids in the NLS were essential for the nuclear import of pericentrin, we mutated lysine and arginine to neutral amino acid alanine within the fragment PCNT1-235 (Figure 5A). The results showed that the mutants R8A/R9A, R23A/R25A and K40A/R41A/K42A abolished PCNT1-235 nuclear accumulation (Figure 5B and 5C); in contrast, mutants R10A/K11A, R16A/K18A and K36A/K37A showed no obvious effect on the nuclear localization of PCNT1-235 (Figure 5B and 5C). Interestingly, mutation of K26A/K28A induced a subset of PCNT1-235 to localize unusually to the nucleolus (Figure 5B and 5C), suggesting that pericentrin might be involved in the function of the nucleolus. To further investigate the interaction of these mutants with importin β, we performed a GST-importin β pull-down assay using cell lysates of GFP-PCNT1-235 fusion protein-expressing HeLa cells. As shown by western blot, the mutants R8A/R9A, R23A/R25A and K40A/R41A/K42A were either unable to bind or only weakly bound GST-importin β, whereas the other mutants could still strongly bind with GST-importin β (Figure 5D). These results indicate that the basic amino acid residues R8, R9, R23, R25, K40, R41 and K42 play crucial roles in the nuclear import of pericentrin via interacting with importin β.
Mutational analysis of pericentrin NLS. (A) The map for mutational sites in the NLS of pericentrin. The positions of the lysine and arginine replaced by alanine in the NLS mutants are indicated. (B) The subcellular localizations of wild-type (WT) NLS (PCNT1-235) and its mutants in HeLa cells. Bar, 20 μm. (C) The nuclear accumulation percentages of pericentrin NLS and its mutants were analyzed. At least 30 cells were evaluated for each sample. Data were averages of three independent experiments, shown as mean ± SD. (D) The pull-down assay using GST-importin β. HeLa cells were transfected with wild-type and site-directed mutations of PCNT1-235 for 24 h, lysed and mixed with GST-importin β beads for 2 h at 4 °C. After washing four times, the proteins bound to beads were eluted and analyzed by western blot using anti-GFP antibody.
The classical NLSs usually consist of one or two basic amino acid clusters, which are termed as monopartite or bipartite NLS 25. However, our mutagenesis analysis demonstrated that the critical amino acids of pericentrin NLS locate in three basic amino acid clusters, and all these clusters of pericentrin NLS are necessary for the nuclear import process, suggesting that pericentrin contains an unusual tripartite NLS, which is quite different from the classical NLSs. Although GFP-PCNT8-28 and GFP-PCNT23-42 also showed nuclear localization, we propose that this might be due to their small sizes. Nevertheless, the transport of the macromolecule pericentrin protein from the cytoplasm into the nucleus does require the three clusters containing full NLS.
The nuclear import of pericentrin is mediated by importin α/β or importin β alone, and regulated by Ran GTPase cycle
Most basic charged NLS-containing proteins are thought to be imported into the nucleus via the importin α/β heterodimer. However, some NLS-containing proteins can directly interact with importin β 25, 26. To determine whether the interaction between pericentrin NLS and importin β is mediated by importin α, we performed in vitro protein binding assays using bacterially expressed His-GFP-PCNT1-235, GST-importin α and GST-importin β proteins. We immobilized PCNT1-235 on CNBr-activated beads and incubated the beads with importin α and β separately or with both of them. The results showed that PCNT1-235 could bind importin α or importin β individually, although the affinity was relatively low (Figure 6A and 6B). However, when both importin α and β were present, the quantity of importin α and β bound to PCNT1-235 beads increased dramatically by about three-fold (Figure 6C-6F). These results indicate that, although pericentrin NLS can bind importin β directly, importin α may facilitate the interaction between pericentrin NLS and importin β. Taken together, we propose that the nuclear import of pericentrin is mediated mainly by the importin α/β heterodimer, but without importin α, pericentrin can also be imported into the nucleus directly via importin β.
Interaction of pericentrin with importin α/β in vitro. (A) PCNT1-235 bound to importin α directly in vitro. In 200 μl TB buffer, gradient concentration of importin α (from 0.02 to 0.5 nM) was incubated with His-GFP-PCNT1-235-immobilized CNBr beads for 2 h at 4 °C. The beads were washed four times and the bound proteins were analyzed by SDS-PAGE. (B) PCNT1-235 bound to importin β directly in vitro. The experiment was performed similar to that shown in A. (C) Importin β facilitated the interaction between PCNT1-235 and importin α in vitro. In 200 μl TB buffer, gradient concentration of importin α (from 0.02 to 0.5 nM) and 0.1 nM importin β were incubated with His-GFP-PCNT1-235-immobilized CNBr beads for 2 h at 4 °C. The beads were washed four times and the bound proteins were analyzed by SDS-PAGE. (D) Importin α facilitated the interaction between PCNT1-235 and importin β in vitro. The experiment was performed similar to that shown in C. (E) The quantitative analysis of pericentrin NLS-importin α affinity in vitro. (F) The quantitative analysis of pericentrin NLS-importin β affinity in vitro.
Ran GTPase is an essential factor for the nucleocytoplasmic transport 23, 26. In the nucleus, RanGTP binds to importin β and disrupts the import complex by changing the conformation of importin β 23, 26. To determine whether the nuclear import process of pericentrin is under the regulation of Ran, we treated cells transfected with RFP-RanQ69L (a mutant defective in GTPase activity and therefore locked in the GTP-bound form) with LMB. The results showed that pericentrin in the transfected cells became insensitive to LMB and remained in cytoplasm even after LMB treatment (Figure 7A), demonstrating that RanQ69L inhibits the import process of endogenous pericentrin. We further co-transfected HeLa cells with GFP-PCNT1-235 and RFP-RanQ69L, and found that RanQ69L obviously prevented the nuclear localization of GFP-PCNT1-235 (Figure 7B). Next, we analyzed the quantity of the bound importin α/β on GFP-PCNT1-235CNBr beads in the presence of gradient concentrations of RanQ69L. The results showed that importin α/β dissociated from GFP-PCNT1-235 rapidly along with the increase of RanQ69L (Figure 7C and 7D). These data support that RanQ69L inhibited the nuclear import of pericentrin by disrupting the import complex of pericentrin.
The import process of pericentrin is regulated by Ran GTPase. (A) The endogenous pericentrin was insensitive to LMB treatment when RanQ69L was overexpressed. Cells were transfected with RFP-RanQ69L for 24 h, and then treated with LMB for 6 h. The cells before and after LMB treatment were fixed and immunostained with anti-pericentrin antibody. The arrows showed the transfected cells and the arrowheads showed the untransfected cells. Bar, 20 μm. (B) GFP-PCNT1-235 changed the distribution in cells when RFP-RanQ69L was present. GFP-PCNT1-235 was transfected into HeLa cells alone or together with RFP-RanQ69L. After 24 h, cells were fixed for observation. Bar, 20 μm. (C) The interaction between PCNT1-235 and importin α/β in vitro was inhibited by RanQ69L. In 200 μl TB buffer, gradient concentration of His-RanQ69L (from 0 to 2 nM), 0.1 nM importin α and 0.1 nM importin β were incubated with His-GFP-PCNT1-235-immobilized CNBr beads for 2 h at 4 °C. The beads were washed four times and the bound proteins were analyzed by SDS-PAGE. (D) The quantitative analysis of pericentrin NLS-importin α/β affinity in vitro when RanQ69L was present.
Mutation of the NLS or NESs caused mis-localization of full-length pericentrin
To assess the role of the NLS and NESs in the localization of the whole pericentrin molecule, we generated point mutations in the NLS and the NESs of full-length pericentrin. As expected, the wild-type pericentrin located in the cytoplasm, and shuttled between the nucleus and cytoplasm (Figure 8). We found that none of single-NES mutations could change the cytoplasmic localization of pericentrin (data not shown). However, when the entire NESs were mutated, the amount of the mutant was increased in the nucleus (Figure 8). Conversely, dysfunction of the NLS had almost no effect on the cytoplasmic distribution of pericentrin, but caused a defect in the nuclear entry of the molecule, as probed by LMB treatment (Figure 8). More importantly, our data indicated that both NESm-PCNT and NLSm-PCNT lost the activity to shuttle between the nucleus and cytoplasm. Thus, we suggest that the NESs of pericentrin are required for its cytoplasmic localization, and both NESs and NLS of pericentrin are essential for its nucleocytoplasmic trafficking.
Nucleocytoplasmic assays of wild-type and NLS- and NESs-deficient mutants of pericentrin. (A) The immunofluorescence of wild-type and NLS- and NESs-deficient mutants of pericentrin in cells treated with LMB for 3-6 h and LMB withdrawn for 3-12 h afterward. Wild-type pericentrin (WT), NESs-deficient mutant (NESm) and NLS-deficient mutant (NLSm) of pericentrin with HA tag were transfected with HeLa cells for 36 h, then the cells were treated with 10 nM LMB for 3-6 h, and after wash with PBS four times to remove LMB, the cells were grown in the fresh medium without LMB for further 3-12 h. All the cells were fixed and immunostained with anti-HA antibody. The nuclei were stained with DAPI (blue). Bar, 20 μm. (B) The cytoplasmic accumulation percentages of WT, NLSm and NESm pericentrin were analyzed. At least 30 cells were evaluated for each sample. Data were averages of three independent experiments, shown as mean ± SD.
Discussion
Previous studies have shown that pericentrin is sensitive to LMB, a specific inhibitor of Crm1-dependent nuclear export, raising the possibility that pericentrin might be a nucleocytoplasmic trafficking protein 15. In this study, we confirmed that pericentrin localizes predominantly in the cytoplasm and this subcellular distribution is sensitive to LMB. Through construction of truncated mutants and nuclear export assays using the Rev1.4-GFP system, we detected five functional NESs within pericentrin, including NES1 (245LELEALRLSL254), NES2 (1158LQDALRRLLGL1168), NES3 (1169FGETLRAAVTL1179), NES4 (1540LDEFNELAI1548) and NES5 (1745VIEKLQHELSL1755) (Table 2). There is accumulating evidence that large nucleocytoplasmic shutting proteins usually contain more than one NES in their sequences. For example, FANCA (Fanconi anemia group A protein, containing 1 455 amino acids) was reported to have five NESs 27; colon cancer-associated protein APC (adenomatous polyposis coli, containing 2 845 amino acids) contains two NESs 28, 29. In this study, we found that pericentrin contains five NESs and all of these sequences contribute to the nuclear export of this protein, although these sequences vary in the nuclear export activity. We also observed that a single-NES mutation was not sufficient to block the nuclear export of pericentrin, but, when all of the five NESs were mutated, the protein failed to be retained in cytoplasm. Therefore, we propose that increasing numbers of the NESs would be an efficient way to enhance the interaction between pericentrin and Crm1 and facilitate its effective nuclear export. However, we noticed that there were still less than 40% of the NESm-PCNT proteins localized in cytoplasm. A possible reason might be that the PACT domain, which is responsible for its centrosomal targeting 5, remains intact in NESm-PCNT. As a result, at least a part of the mutant protein can be 'held' on centrosomes stably. Moreover, many pericentrin-associated proteins, such as dynein and PKC 8, 30, predominantly localize in cytoplasm and could retain pericentrin in cytoplasm by interacting with it, although its NES signals were destroyed.
Although pericentrin appears predominantly cytoplasmic, it can still shuttle between the cytoplasm and the nucleus. In this study, we detected one NLS at the N-terminus of pericentrin. The classical NLS is characterized by one or two clusters of basic amino acids separated by a linker and recognized by importin α 31. However, the NLS of pericentrin (from amino acids 8 to 42) consists of three clusters of basic amino acids and all of the three clusters are essential for its nuclear localization. It was reported that EGFR also contains a tripartite NLS 20. Moreover, the first two clusters of the EGFR tripartite NLS were sufficient to target GFP fusion protein into the nucleus, but the nuclear targeting was weak when the fusion partner is a large protein 20. Interestingly, we found similar results with the NLS of pericentrin: GFP-PCNT8-28, which contains the first two clusters of the NLS of pericentrin, showed nuclear localization, whereas the K40A/R41A/K42A mutant of GFP-PCNT1-235 failed to target the nucleus. On the basis of these data, we suggest that moving the large molecule of pericentrin into the nucleus should require full tripartite NLS. The classical NLSs are usually recognized by importin α and interact with importin β for translocation from the cytoplasm to nucleus through the nuclear pore complex. Unlike the classical ones, the NLS of pericentrin could bind to importin β directly, although importin α facilitates their interaction, again showing that the NLS of pericentrin is not a typical NLS. The advantage of having two nuclear import pathways would be that the nuclear import process can be carried on even when importin α is absent in cells. Although pericentrin was demonstrated to interact with CHD3/4 and MTA2, components of a nuclear complex, it remains unclear whether the centrosomal pericentrin has function in nucleus 14. Our findings about the NLS of pericentrin at least indicate that pericentrin can shuttle into the nucleus, and support the hypothesis that pericentrin plays a role in nucleus. However, when and why pericentrin enters the nucleus is still unclear.
Previous study demonstrated that overexpression of pericentrin induces a series of mitotic defects, including centrosome and spindle disorganization, chromosome misalignment, genomic instability and cytokinesis failure 30. Recent studies of the human dwarfism MOPD II and Seckel syndrome implicated pericentrin in the regulation of cell cycle progression 12, 13. We also found that overexpression of wild-type pericentrin would lead to G2/M transition delay (data not shown). Dynamic nucleocytoplasmic trafficking of proteins is a common regulatory mechanism for many cellular processes such as cell cycle progression and signal transduction. In fact, many proteins involved in cell cycle checkpoints and maintenance of genome integrity such as cyclin B1, BRCA1 and p53 were shown to be regulated, at least in part, by nucleocytoplasmic transport 32, 33, 34. Although we mainly report the nucleocytoplasmic transport of pericentrin in this work, this protein may also play an important role in the cell cycle regulation, and further studies of the detailed mechanism for its roles in the cell cycle control will be needed.
In summary, we identified for the first time five functional NESs and one tripartite NLS within pericentrin that are important for its subcellular localization and its nucleocytoplasmic trafficking during the cell cycle (Table 2).
Materials and Methods
Plasmid construction
The cDNA coding (NCBI AF515282) for human pericentrin was amplified from the Human HEK 293 cDNA library (Clontech) by PCR. The truncated mutants PCNT1-235, PCNT1-268, PCNT1-50, PCNT50-235, PCNT997-1101, PCNT997-1158, PCNT1146-1302, PCNT1188-1302, PCNT1420-1533, PCNT1420-1558, PCNT1596-1739, PCNT1596-1764, PCNT1827-1880 and PCNT1761-1880 were amplified from pericentrin. All these pericentrin DNA fragments were inserted into pEGFP-C2 vector. A series of NLS mutants were generated by site-directed mutagenesis from truncated mutant PCNT1-235. The NLS-deficient mutant and NESs-deficient mutant of full-length pericentrin were generated by site-directed mutagenesis. The truncated mutants PCNT8-28, PCNT23-42, PCNT6-13, PCNT14-30 and PCNT34-44 were generated by annealing the sense and the antisense oligonucleotides and inserting them into pEGFP-C2 vector.
The pRev1.4-GFP vector was kindly provided by Dr Zhigang Lu. The cNESs sequences of pericentrin (cNES1, 245LELEALRLSL254; cNES4, 1158LQDALRRLLGL1168; cNES5, 1169FGETLRAAVTL1179; cNES6, 1540LDEFNELAI1548; and cNES7, 1745VIEKLQHELSL1755) and their site-directed mutants (cNES1-mt, 245LELEALRASA254; cNES4-mt, 1158LQDALRRLAGA1168; cNES5-mt, 1169FGETLRAAATA1179; cNES6-mt, 1540LDEFNEAAA1548; and cNES7-mt, 1745VIEKLQHEASA1755) were generated by annealing the sense and the antisense oligonucleotides and inserted into the EcoRI and SalI sites of pRev1.4-GFP vector.
Human importin β and Crm1 were cloned into pGEX4T-1. Human importin α1 and the truncated mutant PCNT1-235 were cloned into pET28a. RanQ69L was cloned into monoRFP and pET28a.
Protein purification
Escherichia coli strain BL21 (pLysS) was transformed with pET28a-GFP, pET28a-GFP-PCNT1-235, pET28a-importin α, pET28a-RanQ69L, pGEX4T-1-importin β and pGEX4T-1-Crm1. GST-importin β and GST-Crm1 were purified using glutathione sepharose-4B affinity resin (Pharmacia). His-GFP, His-GFP-PCNT1-235, His-importin α and His-RanQ69L were purified using TALON metal affinity resin (Clontech).
Cell culture, treatment and transfection
HeLa cells were cultured in DMEM (Invitrogen) with 10% fetal bovine serum (HyClone), 100 U/ml penicillin and 100 μg/ml streptomycin at 37 °C and 5% CO2. For blocking the nuclear export of NES proteins, cells were treated with 10 nM LMB (Sigma) for 3 or 6 h. For transfection, cells were grown on coverslips in 35-mm diameter culture dishes to about 80% confluence and transfected with the indicated plasmids using Lipofectamine 2000 (Invitrogen) according to the protocol provided by the manufacturer.
Immunofluorescence
The rabbit polyclonal antibody against pericentrin was purchased from Abcam and the mouse monoclonal antibody against HA was purchased from PTGLAB. The FITC-conjugated goat anti-rabbit secondary antibody was purchased from Dako and the FITC-conjugated goat anti-mouse secondary antibody was purchased from Invitrogen. The cells cultured on coverslips were fixed with 4% paraformaldehyde-PBS for 15 min at room temperature and permeabilized in 0.5% Triton X-100-PBS buffer for 3 min. Alternatively, cells were fixed in cold methanol for 5 min. The primary antibodies were diluted 1:500 (for anti-pericentrin) or 1:100 (for anti-HA) in 3% BSA-PBS, and incubated with the cells at 37 °C for 1 h or at 4 °C over night. Then, cells were washed with PBS three times. The cells were then incubated with fluorescence-dye conjugated secondary antibodies at 1:200 dilution in 3% BSA-PBS at 37 °C for 1 h. After rinsing with PBS three times, the cells were mounted onto slides with the mowiol (Sigma) mount, which contained DAPI to stain DNA.
Immunoprecipitation
HeLa cells were lysed with 1%-NP40-TB buffer (20 mM HEPES-KOH (pH 7.3), 110 mM potassium acetate, 2 mM magnesium acetate, 5 mM sodium acetate and 0.5 mM EGTA) and centrifuged for suspension. Rabbit anti-pericentrin antibody or rabbit IgG (as a control) was incubated with protein A beads (Pharmacia) and then mixed with cell lysates for 2 h at 4 °C. The beads were washed with TB buffer four times, and the proteins that were bound to beads were eluted by sample buffer and analyzed by western blot.
In vitro binding assay
His-GFP or His-GFP-PCNT1-235 (100 nmol) proteins were loaded onto CNBr-activated Sepharose 4B beads (Amersham Biosciences). The beads were then incubated with gradient concentrations of His-importin α (from 0.02 to 0.5 nM), or gradient concentrations of GST-importin β (from 0.02 to 0.5 nM), or a certain concentration of His-importin α (0.1 nM) and gradient concentrations of GST-importin β (from 0 to 0.2 nM), or a certain concentration of GST-importin β (0.1 nM) and gradient concentrations of His-importin α (from 0 to 0.2 nM), or certain concentrations of His-importin α (0.1 nM) and GST-importin β and gradient concentrations of RanQ69L (from 0 to 2 nM) in 200 μl TB buffer for 2 h at 4 °C. The beads were washed with TB buffer four times and the proteins that were bound to beads were eluted by sample buffer and analyzed by 6-15% SDS-PAGE.
Pull-down assay
HeLa cells transfected with full-length pericentrin, fragments of pericentrin or mutations of PCNT1-235, they were lysed in 0.5% NP40-TB buffer and then centrifuged at 13 000 g for 20 min. Glutathione Sepharose-4B beads immobilized with GST-Crm1 or GST-importin β were incubated with cell lysates for 2 h at 4 °C. The beads were washed with 0.5% NP40-TB buffer four times and the proteins that were bound to beads were eluted by sample buffer and analyzed by western blot.
Western blot
Proteins were separated by 6-15% SDS-PAGE and transferred onto nitrocellulose membrane. The membranes were probed with primary antibodies (1:1 000 dilution in TTBS containing 5% non-fat milk). Rabbit polyclonal antibodies against GFP, Crm1 and importin β were generated by injecting rabbits. Mouse monoclonal antibody against Ran was purchased from Sigma. After washing and blocking, the membranes were incubated with horseradish peroxidase-conjugated goat anti-rabbit secondary antibodies (Jackson) (1:5 000 dilution in TTBS containing 5% non-fat milk). The membranes were developed for visualization by enhanced chemiluminescence (Sigma) and X-ray film (Kodak).
In vivo nuclear export assay
The in vivo nuclear export assay using NES-deficient vector Rev1.4-GFP was first described in Henderson's paper and has been recognized to be an effective method to test the nuclear export activity of functional NESs 24. The basic principle is that the Rev1.4-GFP fusion protein, which contains the Rev-NLS but lacks the NES, is located in the nucleus, and the nuclear export activity would be restored by inserting a functional NES sequence. However, some NESs with relative weak export activity cannot overcome the import activity of Rev-NLS and would distribute the protein in the whole cells or even retain it in the nucleus. Act.D (Sigma) treatment can specifically prevent the nuclear import of Rev and make these weak NESs detectable. Cycloheximide (10 μg/ml; Boehringer Mannheim) was added in parallel to all samples to ensure that cytoplasmic GFP arises from nuclear export and not from newly translated protein. On the basis of the cell percentages accumulated in cytoplasm before and after Act.D treatment, the nuclear export activities are estimated to 9 degrees (from 1 to 9). The highest score of '9' applies to NESs that shift GFPs completely to the cytoplasm in 80% of cells even before Act.D treatment. Strong NESs (degree 8 and 7) can overcome the rate of import and shift GFPs completely to the cytoplasm in 51-80% and 20-50% of cells before Act.D treatment, respectively. NESs with degree 6, 5 and 4 only shift GFPs completely to the cytoplasm in less than 20% of cells before Act.D treatment, but increase to more than 80%, 51-80% and 20-50% of cells after Act.D treatment, respectively. Weak NESs with degree 3 and 2 cannot completely shift GFPs to the cytoplasm, instead, only less than 50% of the cells can shift partially to the cytoplasm before Act.D treatment, and more than 80%, and 51-80% of the cells can shift partially to the cytoplasm after Act.D treatment, respectively. Very weak NESs with degree 1 cannot normally overcome the rate of Rev-NLS-mediated import (less than 20% of cells before Act.D treatment), but are able to shift the GFP fusion protein when the import is blocked (20-50% of cells after Act.D treatment).
References
Doxsey SJ, Stein P, Evans L, Calarco PD, Kirschner M . Pericentrin, a highly conserved centrosome protein involved in microtubule organization. Cell 1994; 76:639–650.
Zimmerman W, Doxsey SJ . Construction of centrosomes and spindle poles by molecular motor-driven assembly of protein particles. Traffic 2000; 1:927–934.
Diviani D, Scott JD . AKAP signaling complexes at the cytoskeleton. J Cell Sci 2001; 114:1431–1437.
Dictenberg JB, Zimmerman W, Sparks CA, et al. Pericentrin and gamma-tubulin form a protein complex and are organized into a novel lattice at the centrosome. J Cell Biol 1998; 141:163–174.
Takahashi M, Yamagiwa A, Nishimura T, Mukai H, Ono Y . Centrosomal proteins CG-NAP and kendrin provide microtubule nucleation sites by anchoring gamma-tubulin ring complex. Mol Biol Cell 2002; 13:3235–3245.
Zimmerman WC, Sillibourne J, Rosa J, Doxsey SJ . Mitosis-specific anchoring of gamma tubulin complexes by pericentrin controls spindle organization and mitotic entry. Mol Biol Cell 2004; 15:3642–3657.
Diviani D, Langeberg LK, Doxsey SJ, Scott JD . Pericentrin anchors protein kinase A at the centrosome through a newly identified RII-binding domain. Curr Biol 2000; 10:417–420.
Chen D, Purohit A, Halilovic E, Doxsey SJ, Newton AC . Centrosomal anchoring of protein kinase C betaII by pericentrin controls microtubule organization, spindle function, and cytokinesis. J Biol Chem 2004; 279:4829–4839.
Miyoshi K, Asanuma M, Miyazaki I, et al. DISC1 localizes to the centrosome by binding to kendrin. Biochem Biophys Res Commun 2004; 317:1195–1199.
Li Q, Hansen D, Killilea A, et al. Kendrin/pericentrin-B, a centrosome protein with homology to pericentrin that complexes with PCM-1. J Cell Sci 2001; 114:797–809.
Delaval B, Doxsey S . Genetics. Dwarfism, where pericentrin gains stature. Science 2008; 319:732–733.
Griffith E, Walker S, Martin CA, et al. Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling. Nat Genet 2008; 40:232–236.
Rauch A, Thiel CT, Schindler D, et al. Mutations in the pericentrin (PCNT) gene cause primordial dwarfism. Science 2008; 319:816–819.
Sillibourne JE, Delaval B, Redick S, Sinha M, Doxsey SJ . Chromatin remodeling proteins interact with pericentrin to regulate centrosome integrity. Mol Biol Cell 2007; 18:3667–3680.
Keryer G, Di Fiore B, Celati C, et al. Part of Ran is associated with AKAP450 at the centrosome: involvement in microtubule-organizing activity. Mol Biol Cell 2003; 14:4260–4271.
Weis K . Regulating access to the genome: nucleocytoplasmic transport throughout the cell cycle. Cell 2003; 112:441–451.
Gorlich D, Kutay U . Transport between the cell nucleus and the cytoplasm. Annu Rev Cell Dev Biol 1999; 15:607–660.
Kalderon D, Richardson WD, Markham AF, Smith AE . Sequence requirements for nuclear location of simian virus 40 large-T antigen. Nature 1984; 311:33–38.
Robbins J, Dilworth SM, Laskey RA, Dingwall C . Two interdependent basic domains in nucleoplasmin nuclear targeting sequence: identification of a class of bipartite nuclear targeting sequence. Cell 1991; 64:615–623.
Hsu SC, Hung MC . Characterization of a novel tripartite nuclear localization sequence in the EGFR family. J Biol Chem 2007; 282:10432–10440.
Sorokin AV, Kim ER, Ovchinnikov LP . Nucleocytoplasmic transport of proteins. Biochemistry (Mosc) 2007; 72:1439–1457.
Yoneda Y, Hieda M, Nagoshi E, Miyamoto Y . Nucleocytoplasmic protein transport and recycling of Ran. Cell Struct Funct 1999; 24:425–433.
Pemberton LF, Paschal BM . Mechanisms of receptor-mediated nuclear import and nuclear export. Traffic 2005; 6:187–198.
Henderson BR, Eleftheriou A . A comparison of the activity, sequence specificity, and CRM1-dependence of different nuclear export signals. Exp Cell Res 2000; 256:213–224.
Lange A, Mills RE, Lange CJ, et al. Classical nuclear localization signals: definition, function, and interaction with importin alpha. J Biol Chem 2007; 282:5101–5105.
Strom AC, Weis K . Importin-beta-like nuclear transport receptors. Genome Biol 2001; 2:REVIEWS3008.
Ferrer M, Rodriguez JA, Spierings EA, et al. Identification of multiple nuclear export sequences in Fanconi anemia group A protein that contribute to CRM1-dependent nuclear export. Hum Mol Genet 2005; 14:1271–1281.
Tickenbrock L, Cramer J, Vetter IR, Muller O . The coiled coil region (amino acids 129-250) of the tumor suppressor protein adenomatous polyposis coli (APC). Its structure and its interaction with chromosome maintenance region 1 (Crm-1). J Biol Chem 2002; 277:32332–32338.
Neufeld KL, Nix DA, Bogerd H, et al. Adenomatous polyposis coli protein contains two nuclear export signals and shuttles between the nucleus and cytoplasm. Proc Natl Acad Sci USA 2000; 97:12085–12090.
Purohit A, Tynan SH, Vallee R, Doxsey SJ . Direct interaction of pericentrin with cytoplasmic dynein light intermediate chain contributes to mitotic spindle organization. J Cell Biol 1999; 147:481–492.
Fontes MR, Teh T, Kobe B . Structural basis of recognition of monopartite and bipartite nuclear localization sequences by mammalian importin-alpha. J Mol Biol 2000; 297:1183–1194.
Zhang Y, Xiong Y . A p53 amino-terminal nuclear export signal inhibited by DNA damage-induced phosphorylation. Science 2001; 292:1910–1915.
Yang J, Bardes ES, Moore JD, et al. Control of cyclin B1 localization through regulated binding of the nuclear export factor CRM1. Genes Dev 1998; 12:2131–2143.
Rodriguez JA, Henderson BR . Identification of a functional nuclear export sequence in BRCA1. J Biol Chem 2000; 275:38589–38596.
Delaval B, Doxsey SJ . Pericentrin in cellular function and disease. J Cell Biol 2010; 188:181–190.
Tibelius A, Marhold J, Zentgraf H, et al. Microcephalin and pericentrin regulate mitotic entry via centrosome-associated Chk1. J Cell Biol 2009; 185:1149–1157.
Acknowledgements
We thank Dr Zhigang Lu (Laboratory of Chemical Genomics, School of Chemical Biology and Biotechnology, Shenzhen Graduate School of Peking University, China) for the kind gift of pRev1.4-GFP vector and the pRev1.4-GFP vectors containing NESs of MAPKK, RanBP1, c-ABL, FMRP, IκB, hdm2 and p53 respectively. We thank Dr Xueliang Zhu (Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, China) and Dr Doxsey (Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605, USA) for the generous gift of pericentrin gene. We also thank other members in our laboratory for helpful comments. This work was supported by funds from the National Natural Science Foundation of China (NSFC) (30721064, 30900726 and 90913021), the National Basic Research Program of China (2010CB833705) and the National Key Scientific Program of China (2007CB914502).
Author information
Authors and Affiliations
Corresponding author
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Establish the standard of nuclear export activity using Rev1.4-GFP system. (PDF 287 kb)
Rights and permissions
About this article
Cite this article
Liu, Q., Yu, J., Zhuo, X. et al. Pericentrin contains five NESs and an NLS essential for its nucleocytoplasmic trafficking during the cell cycle. Cell Res 20, 948–962 (2010). https://doi.org/10.1038/cr.2010.89
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2010.89
Keywords
This article is cited by
-
Nuclear roles for cilia-associated proteins
Cilia (2017)
-
The GTPase RAN regulates multiple steps of the centrosome life cycle
Chromosome Research (2016)
-
Calumenin-15 facilitates filopodia formation by promoting TGF-β superfamily cytokine GDF-15 transcription
Cell Death & Disease (2013)
-
Chromatin-bound NLS proteins recruit membrane vesicles and nucleoporins for nuclear envelope assembly via importin-α/β
Cell Research (2012)
-
Potential effects of CRM1 inhibition in mantle cell lymphoma
Chinese Journal of Cancer Research (2012)