Abstract
The molecular mechanisms by which the endosomal sorting complexes required for transport (ESCRT) proteins contribute to the integrity of the nuclear envelope (NE) barrier are not fully defined. We leveraged the single NE hole generated by mitotic extrusion of the Schizosaccharomyces pombe spindle pole body to reveal two modes of ESCRT function executed by distinct complements of ESCRT-III proteins, both dependent on CHMP7/Cmp7. A grommet-like function is required to restrict the NE hole in anaphase B, whereas replacement of Cmp7 by a sealing module ultimately closes the NE in interphase. Without Cmp7, nucleocytoplasmic compartmentalization remains intact despite NE discontinuities of up to 540 nm, suggesting mechanisms to limit diffusion through these holes. We implicate spindle pole body proteins as key components of a diffusion barrier acting with Cmp7 in anaphase B. Thus, NE remodelling mechanisms cooperate with proteinaceous diffusion barriers beyond nuclear pore complexes to maintain the nuclear compartment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
Code availability
The MATLAB script for quantification of ESCRT subunits is available with no access restrictions at https://github.com/LusKingLab/Find_n_Fit_Spots_3D.
References
Dultz, E., Wojtynek, M., Medalia, O. & Onischenko, E. The nuclear pore complex: birth, life, and death of a cellular behemoth. Cells 11, 1456 (2022).
Makarova, M. & Oliferenko, S. Mixing and matching nuclear envelope remodeling and spindle assembly strategies in the evolution of mitosis. Curr. Opin. Cell Biol. 41, 43–50 (2016).
Ungricht, R. & Kutay, U. Mechanisms and functions of nuclear envelope remodelling. Nat. Rev. Mol. Cell Biol. 18, 229–245 (2017).
Vietri, M. et al. Spastin and ESCRT-III coordinate mitotic spindle disassembly and nuclear envelope sealing. Nature 522, 231–235 (2015).
Olmos, Y., Hodgson, L., Mantell, J., Verkade, P. & Carlton, J. G. ESCRT-III controls nuclear envelope reformation. Nature 522, 236–239 (2015).
Gu, M. et al. LEM2 recruits CHMP7 for ESCRT-mediated nuclear envelope closure in fission yeast and human cells. Proc. Natl Acad. Sci. USA 114, E2166–E2175 (2017).
von Appen, A. et al. LEM2 phase separation promotes ESCRT-mediated nuclear envelope reformation. Nature 582, 115–118 (2020).
Ventimiglia, L. N. et al. CC2D1B coordinates ESCRT-III activity during the mitotic reformation of the nuclear envelope. Dev. Cell 47, 547–563 (2018).
Olmos, Y., Perdrix-Rosell, A. & Carlton, J. G. Membrane binding by CHMP7 coordinates ESCRT-III-dependent nuclear envelope reformation. Curr. Biol. 26, 2635–2641 (2016).
Thaller, D. J. et al. An ESCRT–LEM protein surveillance system is poised to directly monitor the nuclear envelope and nuclear transport system. eLife 8, e45284 (2019).
Webster, B. M., Colombi, P., Jäger, J. & Lusk, C. P. Surveillance of nuclear pore complex assembly by ESCRT-III/Vps4. Cell 159, 388–401 (2014).
Webster, B. M. et al. Chm7 and Heh1 collaborate to link nuclear pore complex quality control with nuclear envelope sealing. EMBO J. 35, 2447–2467 (2016).
Thaller, D. J. et al. Direct binding of ESCRT protein Chm7 to phosphatidic acid-rich membranes at nuclear envelope herniations. J. Cell Biol. 220, e202004222 (2021).
Denais, C. M. et al. Nuclear envelope rupture and repair during cancer cell migration. Science 352, 353–358 (2016).
Raab, M. et al. ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death. Science 352, 359–362 (2016).
Vietri, M., Radulovic, M. & Stenmark, H. The many functions of ESCRTs. Nat. Rev. Mol. Cell Biol. 21, 25–42 (2020).
Pfitzner, A. K., Moser von Filseck, J. & Roux, A. Principles of membrane remodeling by dynamic ESCRT-III polymers. Trends Cell Biol. 31, 856–868 (2021).
Remec Pavlin, M. & Hurley, J. H. The ESCRTs—converging on mechanism. J. Cell Sci. 133, jcs240333 (2020).
McCullough, J., Frost, A. & Sundquist, W. I. Structures, functions, and dynamics of ESCRT-III/Vps4 membrane remodeling and fission complexes. Annu. Rev. Cell Dev. Biol. 34, 85–109 (2018).
Mierzwa, B. E. et al. Dynamic subunit turnover in ESCRT-III assemblies is regulated by Vps4 to mediate membrane remodelling during cytokinesis. Nat. Cell Biol. 19, 787–798 (2017).
Adell, M. A. Y. et al. Recruitment dynamics of ESCRT-III and Vps4 to endosomes and implications for reverse membrane budding. eLife 6, e31652 (2017).
Wallis, S. S. et al. The ESCRT machinery counteracts Nesprin-2G-mediated mechanical forces during nuclear envelope repair. Dev. Cell 56, 3192–3202 (2021).
Ding, R., West, R. R., Morphew, D. M., Oakley, B. R. & McIntosh, J. R. The spindle pole body of Schizosaccharomyces pombe enters and leaves the nuclear envelope as the cell cycle proceeds. Mol. Biol. Cell 8, 1461–1479 (1997).
Wälde, S. & King, M. C. The KASH protein Kms2 coordinates mitotic remodeling of the spindle pole body. J. Cell Sci. 127, 3625–3640 (2014).
Krüger, L. K., Sanchez, J. L., Paoletti, A. & Tran, P. T. Kinesin-6 regulates cell-size-dependent spindle elongation velocity to keep mitosis duration constant in fission yeast. eLife 8, e42182 (2019).
West, R. R., Vaisberg, E. V., Ding, R., Nurse, P. & McIntosh, J. R. cut11+: A gene required for cell cycle-dependent spindle pole body anchoring in the nuclear envelope and bipolar spindle formation in Schizosaccharomyces pombe. Mol. Biol. Cell 9, 2839–2855 (1998).
Lawrimore, J., Bloom, K. S. & Salmon, E. D. Point centromeres contain more than a single centromere-specific Cse4 (CENP-A) nucleosome. J. Cell Biol. 195, 573–582 (2011).
Sohrmann, M., Schmidt, S., Hagan, I. & Simanis, V. Asymmetric segregation on spindle poles of the Schizosaccharomyces pombe septum-inducing protein kinase Cdc7p. Genes Dev. 12, 84–94 (1998).
Popken, P., Ghavami, A., Onck, P. R., Poolman, B. & Veenhoff, L. M. Size-dependent leak of soluble and membrane proteins through the yeast nuclear pore complex. Mol. Biol. Cell 26, 1386–1394 (2015).
Ribbeck, K. & Görlich, D. The permeability barrier of nuclear pore complexes appears to operate via hydrophobic exclusion. EMBO J. 21, 2664–2671 (2002).
Shulga, N. & Goldfarb, D. S. Binding dynamics of structural nucleoporins govern nuclear pore complex permeability and may mediate channel gating. Mol. Cell. Biol. 23, 534–542 (2003).
Barrientos, E. C. R., Otto, T. A., Mouton, S. N., Steen, A. & Veenhoff, L. M. A survey of the specificity and mechanism of 1,6 hexanediol-induced disruption of nuclear transport. Nucleus 14, 2240139 (2023).
Kanke, M. et al. Auxin-inducible protein depletion system in fission yeast. BMC Cell Biol. 12, 8 (2011).
Zhang, X. R. et al. An improved auxin-inducible degron system for fission yeast. G3 12, jkab393 (2022).
Jaspersen, S. L. Anatomy of the fungal microtubule organizing center, the spindle pole body. Curr. Opin. Struct. Biol. 66, 22–31 (2021).
Tamm, T. et al. Brr6 drives the Schizosaccharomyces pombe spindle pole body nuclear envelope insertion/extrusion cycle. J. Cell Biol. 195, 467–484 (2011).
Fong, C. S., Sato, M. & Toda, T. Fission yeast Pcp1 links polo kinase-mediated mitotic entry to γ-tubulin-dependent spindle formation. EMBO J. 29, 120–130 (2010).
Rosenberg, J. A. et al. Ppc89 links multiple proteins, including the septation initiation network, to the core of the fission yeast spindle-pole body. Mol. Biol. Cell 17, 3793–3805 (2006).
Flory, M. R., Morphew, M., Joseph, J. D., Means, A. R. & Davis, T. N. Pcp1p, an Spc110p-related calmodulin target at the centrosome of the fission yeast Schizosaccharomyces pombe. Cell Growth Differ. 13, 47–58 (2002).
O’Toole, E. T., Winey, M. & McIntosh, J. R. High-voltage electron tomography of spindle pole bodies and early mitotic spindles in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 10, 2017–2031 (1999).
Byers, B. & Goetsch, L. Behavior of spindles and spindle plaques in the cell cycle and conjugation of Saccharomyces cerevisiae. J. Bacteriol. 124, 511–523 (1975).
Halfmann, C. T. et al. Repair of nuclear ruptures requires barrier-to-autointegration factor. J. Cell Biol. 218, 2136–2149 (2019).
Samwer, M. et al. DNA cross-bridging shapes a single nucleus from a set of mitotic chromosomes. Cell 170, 956–972 (2017).
Radulovic, M. et al. ESCRT-mediated lysosome repair precedes lysophagy and promotes cell survival. EMBO J. 37, e99753 (2018).
Skowyra, M. L., Schlesinger, P. H., Naismith, T. V. & Hanson, P. I. Triggered recruitment of ESCRT machinery promotes endolysosomal repair. Science 360, eaar5078 (2018).
Frey, S., Richter, R. P. & Görlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 314, 815–817 (2006).
Petri, M., Frey, S., Menzel, A., Görlich, D. & Techert, S. Structural characterization of nanoscale meshworks within a nucleoporin FG hydrogel. Biomacromolecules 13, 1882–1889 (2012).
Celetti, G. et al. The liquid state of FG-nucleoporins mimics permeability barrier properties of nuclear pore complexes. J. Cell Biol. 219, e201907157 (2020).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters 3, e201702000010 (2017).
Jiang, X. et al. Condensation of pericentrin proteins in human cells illuminates phase separation in centrosome assembly. J. Cell Sci. 134, jcs258897 (2021).
Zwicker, D., Decker, M., Jaensch, S., Hyman, A. A. & Julicher, F. Centrosomes are autocatalytic droplets of pericentriolar material organized by centrioles. Proc. Natl Acad. Sci. USA 111, E2636–E2645 (2014).
Woodruff, J. B. et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell 169, 1066–1077 (2017).
Feng, Z. et al. Structural basis for mitotic centrosome assembly in flies. Cell 169, 1078–1089 (2017).
Bullitt, E., Rout, M. P., Kilmartin, J. V. & Akey, C. W. The yeast spindle pole body is assembled around a central crystal of Spc42p. Cell 89, 1077–1086 (1997).
Pfitzner, A. K. et al. An ESCRT-III polymerization sequence drives membrane deformation and fission. Cell 182, 1140–1155 (2020).
Babst, M., Katzmann, D. J., Estepa-Sabal, E. J., Meerloo, T. & Emr, S. D. ESCRT-III: an endosome-associated heterooligomeric protein complex required for MVB sorting. Dev. Cell 3, 271–282 (2002).
Saksena, S., Wahlman, J., Teis, D., Johnson, A. E. & Emr, S. D. Functional reconstitution of ESCRT-III assembly and disassembly. Cell 136, 97–109 (2009).
Teis, D., Saksena, S. & Emr, S. D. Ordered assembly of the ESCRT-III complex on endosomes is required to sequester cargo during MVB formation. Dev. Cell 15, 578–589 (2008).
Henne, W. M., Buchkovich, N. J., Zhao, Y. & Emr, S. D. The endosomal sorting complex ESCRT-II mediates the assembly and architecture of ESCRT-III helices. Cell 151, 356–371 (2012).
Nguyen, H. C. et al. Membrane constriction and thinning by sequential ESCRT-III polymerization. Nat. Struct. Mol. Biol. 27, 392–399 (2020).
McCullough, J. et al. Structure and membrane remodeling activity of ESCRT-III helical polymers. Science 350, 1548–1551 (2015).
Penfield, L. et al. Regulated lipid synthesis and LEM2/CHMP7 jointly control nuclear envelope closure. J. Cell Biol. 219, e201908179 (2020).
Kinugasa, Y. et al. The very-long-chain fatty acid elongase Elo2 rescues lethal defects associated with loss of the nuclear barrier function in fission yeast cells. J. Cell Sci. 132, jcs229021 (2019).
Kume, K., Cantwell, H., Burrell, A. & Nurse, P. Nuclear membrane protein Lem2 regulates nuclear size through membrane flow. Nat. Commun. 10, 1871 (2019).
Skibinski, G. et al. Mutations in the endosomal ESCRTIII-complex subunit CHMP2B in frontotemporal dementia. Nat. Genet. 37, 806–808 (2005).
Coyne, A. N. et al. Nuclear accumulation of CHMP7 initiates nuclear pore complex injury and subsequent TDP-43 dysfunction in sporadic and familial ALS. Sci. Transl. Med. 13, eabe1923 (2021).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Hentges, P., Van Driessche, B., Tafforeau, L., Vandenhaute, J. & Carr, A. M. Three novel antibiotic marker cassettes for gene disruption and marker switching in Schizosaccharomyces pombe. Yeast 22, 1013–1019 (2005).
Bahler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951 (1998).
Harris, M. A. et al. Fission stories: using PomBase to understand Schizosaccharomyces pombe biology. Genetics 220, iyab222 (2022).
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002).
Davis, M. W. & Jorgensen, E. M. ApE, a plasmid editor: a freely available DNA manipulation and visualization program. Front. Bioinform. 2, 818619 (2022).
Huxley, C., Green, E. D. & Dunham, I. Rapid assessment of S. cerevisiae mating type by PCR. Trends Genet. 6, 236 (1990).
Murray, J. M., Watson, A. T. & Carr, A. M. Transformation of Schizosaccharomyces pombe: lithium acetate/dimethyl sulfoxide procedure. Cold Spring Harb. Protoc. 2016, pdb.prot090969 (2016).
Keeney, J. B. & Boeke, J. D. Efficient targeted integration at leu1-32 and ura4-294 in Schizosaccharomyces pombe. Genetics 136, 849–856 (1994).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kukulski, W. et al. Precise, correlated fluorescence microscopy and electron tomography of lowicryl sections using fluorescent fiducial markers. Methods Cell. Biol. 111, 235–257 (2012).
McDonald, K. High-pressure freezing for preservation of high resolution fine structure and antigenicity for immunolabeling. Methods Mol. Biol. 117, 77–97 (1999).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Mastronarde, D. N. & Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. J. Struct. Biol. 197, 102–113 (2017).
Pollard, D. A., Pollard, T. D. & Pollard, K. S. Empowering statistical methods for cellular and molecular biologists. Mol. Biol. Cell 30, 1359–1368 (2019).
Acknowledgements
We thank the Center for Cellular and Molecular Imaging, Electron Microscopy Facility at Yale School of Medicine EM for assistance, particularly M. Graham and X. Liu. We thank L.-L. Du, A. Frost, I. Hagan, T. Pollard and NBRP (YGRC), Japan (and the investigators who have deposited their strains at this resource) for yeast strains. This work was supported by the National Institutes of Health (grant nos R01 GM105672 to C.P.L. and F32 GM139285 to N.R.A.) and an Allen Distinguished Investigator Award, a Paul G. Allen Frontiers Group advised grant of the Paul G. Allen Family Foundation (M.C.K.).
Author information
Authors and Affiliations
Contributions
Conceptualization: N.R.A., M.C.K. and C.P.L. Data curation: N.R.A., I.V.S., M.C.K. and C.P.L. Formal analysis: N.R.A. and L.C. Funding acquisition: N.R.A., M.C.K. and C.P.L. Investigation: N.R.A., L.C. and W.L.C. Methodology: N.R.A., I.V.S., M.C.K. and C.P.L. Project administration: N.R.A., M.C.K. and C.P.L. Resources: N.R.A., L.C., I.V.S., W.L.C. and E.C.R. Software: I.V.S. Supervision: M.C.K. and C.P.L. Validation: N.R.A., M.C.K. and C.P.L. Visualization: N.R.A. Writing of the original draft: N.R.A., M.C.K. and C.P.L. Writing review and editing: N.R.A., I.V.S., M.C.K. and C.P.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Did2 is not detected at the SPB extrusion site, whereas Ist1 is recruited after Cmp7.
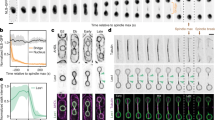
a, Fluorescence micrographs of Did2–GFP, mCherry–Atb2 and Pcp1–mCherry in a representative cell (MKSP3342) in anaphase B. Did2–GFP (left), sum projection of mCherry–Atb2/Pcp1–mCherry (middle) and magnifications (right) of ROI (top to bottom) green, magenta, merge. b, Plot of the percentage of nuclei with Did2–GFP (MKSP3342) co-localized with SPB marker at the indicated phases of the cell cycle. Points and error bars represent the mean and range, respectively, across at least three biological replicates with at least 50 nuclei/Did2–GFP/replicate, n = 372 nuclei. c, Time-lapse fluorescence microscopy of Cmp7–GFP (green) and Ist1–mCherry (magenta) in a late anaphase B cell (MKSP3521). Cell representative of population from two biological replicates. White boxes in overview micrographs indicate ROI magnified below images as single channel and composite images. Scale bars, 1 µm (overview) and 250 nm (ROI). Source numerical data are provided.
Extended Data Fig. 2 Use of the kinetochore protein Fta3 as a standard for ratiometric determination of ESCRT protein copy number at the NE.
a, Fluorescence micrographs of representative mixture of a GFP–ESCRT strain (Ist1–GFP; MKSP3365) and one expressing Fta3–GFP (MKSP3186). Overview micrograph: top left, strain expressing Ist1–GFP (green), mCherry–Atb2 (magenta), and Sad1–mCherry (magenta); bottom right, Fta3–GFP strain (green). Dotted white lines indicate the cell outline. The blue and orange boxes in the overview micrograph outline the ROIs magnified to the right of image as single channel images for Ist1–GFP and Fta3–GFP, as indicated. b, Fluorescence micrographs of representative Fta3–GFP expressing cells (MKSP3186) at indicated cell-cycle stage (top). The black boxes in overview micrographs outline the ROI magnified below. Schematic representation of Fta3–GFP appearance in the different cell-cycle stages (bottom). Number of molecules based on Lawrimore and colleagues27. c, x–y plot of Fta3–GFP focus fluorescence intensity against the expected number of molecules per focus in MKSP3186 based on Lawrimore and colleagues27. Points represent median of all values collected for that stage of the cell cycle (see b); error bars represent the 95% confidence interval. The red line represents a simple linear regression with the R2 shown; n = 366 foci across three biological replicates with at least 50 foci per replicate. Scale bars, 1 µm (overview) and 250 nm (ROI). Source numerical data are provided.
Extended Data Fig. 3 MGM4 and MGM2 reporters—the AID system allows for rapid depletion of Cmp7.
a, Micrographs of MGM4 (top left; MKSP3345) and MGM2 (bottom left; MKSP3960). Histogram of nucleoplasmic-to-cytoplasmic GFP fluorescence (N/C ratio) for individual cells are shown (right). Size between bins, 0.05. Dotted line at 0.75 bin, representing the cut-off we used to gauge robust nucleocytoplasmic compartmentalization. Quantification was performed for three biological replicates. MGM4 data replotted from Fig. 6a. MGM2, n = 178 nuclei, with at least 58 nuclei per biological replicate. Scale bars, 1 µm. b, Cmp7–AID was efficiently degraded within 15 min of treatment with 5IAA. Representative western blot (WB) of Cmp7–AID levels (detected with an antibody to AID) in total protein extracts from cells (MKSP3517) cultured in the presence of 5IAA for the indicated times. Ponceau stain is shown (bottom) to assess relative total protein loads. The relative amount of Cmp7–AID protein, normalized to T0, is shown below the WB as the mean and s.d. of three biological replicates. c, Depletion of Cmp7–AID recapitulates phenotypes observed in cmp7Δ cells, including abolishing recruitment of Ist1–GFP to the NE. Plots of the percentage of nuclei with Ist1–GFP co-localized with SPB marker at the indicated phases of the cell cycle in cells (MKSP3625) expressing Cmp7–AID and OsTIR1F74A. The cells were treated with DMSO (grey) or 5IAA (green) for 1 h. Points and error bars represent the mean and range, respectively, across at least three biological replicates with at least 50 nuclei per GFP–ESCRT per replicate. DMSO, n = 273 nuclei; and 5IAA, n = 296 nuclei. Source numerical data and unprocessed blots are provided.
Extended Data Fig. 4 Genes encoding constitutive SPB proteins and mitotically associated SPB proteins genetically interact with CMP7.
The indicated strains were serially diluted tenfold, plated on YE5S plates and incubated at the indicated temperatures. The plates were imaged after 2 d (30, 32 and 36 °C) or 4 d (22 °C). WT (MKSP399), cmp7Δ (MKSP3107), alp4-1891 (MKSP4067), cmp7Δalp4-1891 (MKSP4163), alp6-719 (MKSP3475), cmp7Δalp6-719 (MKSP4031), alp14Δ (MKSP4063), cmp7Δalp14Δ (MKSP4165), brr6.ts8 (MKSP3461), cmp7Δbrr6.ts8 (MKSP3528), cam1-116F (MKSP4061), cmp7Δcam1-116F (MKSP4167), cdc11-123 (MKSP4060), cmp7Δcdc11-123 (MKSP4169), cut11-1 (MKSP3469), cmp7Δcut11-1 (MKSP4040), gtb1-93 (MKSP4062), cmp7Δgtb1-93 (MKSP4174), kms2 DAmP (MKSP1474), cmp7Δkms2 DAmP (MKSP3683), mto1Δ (MKSP1303), cmp7Δmto1Δ (MKSP4027), mto2Δ (MKSP1305), cmp7Δmto2Δ (MKSP4151), mzt1-T27W (MKSP3476), cmp7Δmzt1-T27W (MKSP4045), plo1-24C (MKSP1461), cmp7Δplo1-24C (MKSP4048), sad1-1 (MKSP3468), cmp7Δsad1-1 (MKSP4033), sid4-SA1 (MKSP4065) and cmp7Δsid4-SA1 (MKSP4171).
Extended Data Fig. 5 Conditional depletion of Pcp1 and Ppc89 impairs nucleocytoplasmic compartmentalization without impacting mitosis progression.
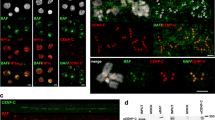
a, WB of Pcp1–AID from cells expressing OsTir1F74A treated with 5IAA as indicated. Ponceau to assess relative total protein. −OsTir1F74A (MKSP4112); +OsTir1F74A (MKSP4113); −OsTir1F74A, cmp7Δ (MKSP4114); +OsTir1F74A, cmp7Δ (MKSP4115). b, Nuclei of pcp1–AID (MKSP3773) and cmp7Δpcp1–AID (MKSP3775) strains expressing MGM4, Cut11–mCherry, mCherry–Atb2 and OsTir1F74A: top, MGM4 (green); middle, Cut11–mCherry/mCherry–Atb2 (magenta); bottom, merge. Cells were treated with DMSO/5IAA as indicated. c, Percentage of pcp1–AID (grey bars, MKSP3773) and cmp7Δpcp1–AID (teal bars, MKSP3775) cells in b with N/C fluorescence ratios >0.75 reflecting a loss of robust nucleocytoplasmic compartmentalization. Points are the mean from each biological replicate; bars and error bars represent the total mean and s.d., respectively. Points average of at least 18 nuclei per cell-cycle phase/strain/replicate, n = 649 nuclei. d, WB of Ppc89–AID levels as in a. −OsTir1F74A (MKSP4175); +OsTir1F74A (MKSP4136); −OsTir1F74A, cmp7Δ (MKSP4176); +OsTir1F74A, cmp7Δ (MKSP4135). e, Nuclei of ppc89–AID (MKSP4136) and cmp7Δppc89–AID (MKSP4135) strains expressing MGM4, Cut11–mCherry and OsTir1F74A shown as in b. f, Percentage of ppc89–AID (grey bars, MKSP4136) and cmp7Δppc89–AID (teal bars, MKSP4135) cells shown in e with N/C fluorescence ratios >0.75 reflecting a loss of robust nucleocytoplasmic compartmentalization. Points are the mean from each biological replicate; bars and error bars represent the total mean and s.d., respectively. Points represent the average of at least 11 nuclei/cell cycle phase/strain/replicate, n = 504 nuclei. g, Time-lapse of mCherry–Atb2 and Cut11–mCherry in a mitotic pcp1–AID (MKSP3773) cell and a cmp7Δpcp1–AID cell (MKSP3775) expressing OsTir1F74A. Imaging began after 1 h of treatment with DMSO/5IAA as indicated. h, Time-lapse of Cut11–mCherry in a mitotic ppc89–AID (MKSP4136) cell and a cmp7Δppc89–AID cell (MKSP4135) expressing OsTir1F74A. Imaging began after 1 h treatment with DMSO/5IAA as indicated. i, pcp1–AID cell (MKSP3773) expressing OsTir1F74A containing a monopolar spindle arranged horizontally with MGM4 (green; left), Cut11–mCherry (magenta) and mCherry–Atb2 (magenta) in middle, and merge (right) 3 h after treatment with 5IAA. Scale bars, 1 µm. Source numerical data and unprocessed blots are provided.
Extended Data Fig. 6 An ESCRT grommet and SPB-dependent diffusion barrier maintain nucleocytoplasmic compartmentalization.
A model of NE sealing and maintenance of mitotic nucleocytoplasmic compartmentalization by the ESCRT machinery and the SPB in S. pombe anaphase B. (1) At mitotic entry, the SPB (dark magenta rounded rectangle) is inserted into the NE to organize the mitotic spindle (light magenta rectangles). (2) Once the SPB is extruded to the cytosol, Cmp7 is recruited by Heh1/Lem2 (not shown). Concurrently, the SPB contributes to the formation of a proteinaceous diffusion barrier (translucent magenta) sensitive to 2.5% 1,6-HD to prevent free diffusion through the unsealed NE. (3) A second wave of ESCRT recruitment comprising Vps32 and Ist1 (multiple circles represent copy number ratio to Cmp7) occurs in late anaphase B, which generates a grommet that restricts the size of the NE hole and supports the SPB-dependent diffusion barrier. The Ist1 circle at partial opacity represents copy number increase during mitosis and into G1/S. (4) Finally, in the G1/S of the following cell cycle, the third wave of ESCRTs, Did4 and Vps24 (multiple circles represent copy number ratio to Cmp7), is recruited, displacing Cmp7 and sealing the NE. N, nucleoplasm (blue); C, cytoplasm (beige).
Supplementary information
Supplementary Video 1
CLEM and electron tomography of a WT cell in early anaphase B. Electron tomogram of an early anaphase B cell (MKSP3180) with segmentation. NE, blue; SPB, dark orange; microtubules, yellow. Scale bar, 100 nm.
Supplementary Video 2
CLEM and electron tomography of a WT cell in late anaphase B. Electron tomogram of a late anaphase B cell (MKSP3180) with segmentation. NE, blue; SPB, dark orange; microtubules, yellow. Annotations at end of the video show a NPC and SPB extrusion hole. Scale bar, 100 nm.
Supplementary Video 3
CLEM and electron tomography of a cmp7∆ cell in early anaphase B. Electron tomogram of an early anaphase B cell (MKSP3243) with segmentation. NE, blue; SPB, dark orange; microtubules, yellow. Scale bar, 100 nm.
Supplementary Video 4
CLEM and electron tomography of a cmp7∆ cell in late anaphase B. Electron tomogram of a late anaphase B cell (MKSP3243) with segmentation. NE, blue; SPB, dark orange; microtubules, yellow. Scale bar, 100 nm.
Supplementary Table 1
GFP tagging of key ESCRT machinery. ESCRT protein in S. pombe is shown (left). Orthologues in S. cerevisiae and Homo sapiens are shown (centre). Tagging approach to ensure function is shown (right).
Supplementary Table 2
List of S. pombe strains used in this study.
Supplementary Table 3
List of plasmids used in this study.
Supplementary Table 4
List of primers used in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ader, N.R., Chen, L., Surovtsev, I.V. et al. An ESCRT grommet cooperates with a diffusion barrier to maintain nuclear integrity. Nat Cell Biol 25, 1465–1477 (2023). https://doi.org/10.1038/s41556-023-01235-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01235-4
This article is cited by
-
Nuclear-import receptors as gatekeepers of pathological phase transitions in ALS/FTD
Molecular Neurodegeneration (2024)
-
A diffusion barrier limits nuclear leaks
Nature Cell Biology (2023)