Abstract
Identification of new molecular targets for the treatment of breast cancer is an important clinical goal, especially for triple-negative breast cancer, which is refractory to existing targeted treatments. The aryl hydrocarbon receptor (AhR) is a ligand-activated transcription factor known primarily as the mediator of dioxin toxicity. However, the AhR can also inhibit cellular proliferation in a ligand-dependent manner and act as a tumor suppressor in mice, and thus may be a potential anticancer target. To investigate the AhR as an anticancer target, we conducted a small molecule screen to discover novel AhR ligands with anticancer properties. We identified raloxifene, a selective estrogen receptor (ER) modulator currently used in the clinic for prevention of ER-positive breast cancer and osteoporosis in post-menopausal women, as an AhR activator. Raloxifene directly bound the AhR and induced apoptosis in ER-negative mouse and human hepatoma cells in an AhR-dependent manner, indicating that the AhR is a molecular target of raloxifene and mediates raloxifene-induced apoptosis in the absence of ER. Raloxifene selectively induced apoptosis of triple-negative MDA-MB-231 breast cancer cells compared with non-transformed mammary epithelial cells via the AhR. Combined with recent data showing that raloxifene inhibits triple-negative breast cancer xenografts in vivo (Int J Oncol. 43(3):785-92, 2013), our results support the possibility of repurposing of raloxifene as an AhR-targeted therapeutic for triple-negative breast cancer patients. To this end, we also evaluated the role of AhR expression on survival of patients diagnosed with breast cancer. We found that higher expression of the AhR is significantly associated with increased overall survival and distant metastasis-free survival in both hormone-dependent (ER-positive) and hormone-independent (ER and progesterone receptor (PR)-negative) breast cancers. Together, our data strongly support the possibility of using the AhR as a molecular target for the treatment of hormone-independent breast cancers.
Similar content being viewed by others
Main
The aryl hydrocarbon receptor (AhR) is a ligand-activated transcription factor of the Per-ARNT-Sim (PAS) protein family. Upon ligand-binding, the AhR dissociates from chaperone proteins HSP90 and XAP2,1 translocates from the cytosol to the nucleus, and heterodimerizes the AhR nuclear translocator (ARNT) to regulate AhR target genes containing functional xenobiotic response elements (XREs).1
The role of the AhR in carcinogenesis continues to evolve. The prototypical AhR ligand 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) acts as a tumor promoter in rodent models;2, 3, 4, 5 however, there is no direct evidence that AhR promotes tumorigenesis in humans. On the contrary, several ligand-dependent and -independent AhR tumor-suppressive functions have been identified. AhR knockout TRAMP (transgenic adenocarcinoma of the mouse prostate) mice develop tumors with increased severity and frequency compared with AhR-expressing counterparts,6 and treatment of TRAMP mice with the AhR ligand 6-methyl-1,3,8-trichlorodibenzofuran (6-MCDF) reduces the frequency of prostate metastases.7, 8 Likewise, formation of diethyl nitrosamine (DEN)-induced liver tumors is accelerated in the absence of the AhR in a ligand-independent manner.9 Similarly, AhR null mice10 develop cecal tumors, whereas WT AhR mice do not, which is further potentiated in the ApcMin/+ model of intestinal cancer.11 At the molecular level, the AhR inhibits cellular proliferation by inducing p27Kip1 expression,12 and deletion of the AhR ligand-binding domain results in constitutively active AhR that induces apoptosis in Jurkat T-lymphocytes.13 Taken together, these data highlight distinct pathways by which AhR can function as a tumor suppressor.
Investigation of the AhR as an anticancer target as well as identification and testing of novel anticancer AhR ligands lacking dioxin-like toxicity is a major goal of our laboratory. Identification of activators of AhR-mediated transcription from existing FDA-approved drugs may significantly expedite this goal by identifying new indications, thereby re-tooling old drugs for a new anticancer target. Importantly, the AhR is activated by a structurally diverse array of ligands,14, 15 some of which have known anticancer effects.16, 17, 18, 19 We previously used a small molecule-screening approach to identify the immunomodulatory drug leflunomide as an AhR ligand with antiproliferative effects in melanoma cells.19 Using the same approach, we identified raloxifene (Evista) as an activator of AhR signaling.
Raloxifene is a selective estrogen receptor (ER) modulator currently used in the clinic for the chemoprevention of osteoporosis. Identified through a structure activity study of a series of 3-aroyl-2-arylbenzo[b]thiophene derivatives aimed at identifying non-steroidal anti-estrogens,20 raloxifene was found to inhibit the growth of 7,12-dimethylbenz(a)anthracene (DMBA)-induced mammary tumors and reverse the inhibitory effect of 17β-estradiol (E2) on ovine prolactin-stimulated α-lactalbumin production.20, 21, 22 Subsequent studies verified the weak estrogenic potential of raloxifene, as well as the ability to inhibit bone loss in ovariectomized rats and reduce risk of vertebral fractures in post-menopausal women with osteoporosis.23, 24, 25 Prior to 2007, the only approved-indication of raloxifene was for prevention and treatment of osteoporosis; however, the results of the study of tamoxifen and raloxifene (STAR) trial demonstrated that raloxifene is as effective as tamoxifen in reducing the risk of invasive breast cancer and is associated with decreased risk of fractures and stroke compared with tamoxifen.26, 27 Thus, raloxifene is also used in the prevention of breast cancer in post-menopausal women with increased risk of developing the disease.23, 27 Raloxifene inhibits the growth of carcinogen-induced mammary tumors in mice.20, 21, 28 In addition, ER-independent effects of raloxifene have been demonstrated in prostate and breast cancer cells.29, 30, 31, 32
Based on our small molecule-screening results, we investigated whether raloxifene had anticancer effects mediated through the AhR, and found that it induces apoptosis in an AhR-dependent manner. Our results suggest that the AhR can be targeted to induce apoptosis in ER-negative breast cancer cells, which may have important clinical implications for the treatment of both hormone-independent and triple-negative breast cancers expressing the AhR.
Results
Raloxifene induces transcriptional activation of the AhR
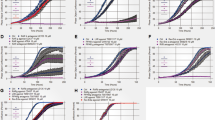
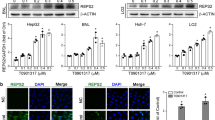
We conducted a small molecule screen to identify novel activators of AhR-mediated transcription. Breast cancer drug raloxifene was identified in this screen and elicited a dose-dependent activation of XRE-driven luciferase reporter (Figure 1a). We were immediately interested in the possibility of AhR-dependent effects of raloxifene in ER-negative breast cancer cells. To evaluate this possibility we used ER-negative MDA-MB-231 breast cancer cells, which express similar levels of AhR as mouse and human hepatoma cell lines (Figure 1b). We then characterized the profile of AhR activation by raloxifene in these cell lines. In both Hepa1 and MDA-MB-231 cells, raloxifene induced AhR nuclear localization similar to TCDD (Figures 1c and d). Interestingly, AhR exhibited partial nuclear localization in MDA-MB-231 cells in the absence of ligand, which was potentiated by TCDD and raloxifene (Figure 1d). We next evaluated the ability of raloxifene to activate the endogenous AhR target genes CYP1A1 and NQO1 by semi-quantitative RT-PCR. Raloxifene significantly increased the expression of these genes in Hepa1 cells (Figure 1e). Activation of both CYP1A1 and NQO1 required functional ARNT (Figure 1f), indicating the requirement for canonical AhR signaling. Raloxifene significantly increased the expression of CYP1A1 in both human HepG2 hepatoma cells (Figure 1g) and MDA-MB-231 breast cancer cells (Figure 1h). Taken together, these results indicated that raloxifene activates AhR signaling in liver and breast cancer cells.
Raloxifene activates the AhR. (a) XRE reporter gene activation in Hepa1 cells after 18-h treatment.14 Results are the mean±s.e.m. of three independent experiments. (b) Western blot depicting relative levels of AhR in mouse WT Hepa1 and human HepG2 hepatoma cells and the estrogen receptor-negative breast cancer cell line MDA-MB-231. GAPDH is shown as a loading control. (c and d) AhR localization in Hepa1 (c) and MDA-MB-231 (d) cells treated with DMSO, 0.1% (v/v), TCDD (1 nM), or raloxifene (20 and 30 μM, respectively) for 4 h. Bars indicate equal image size within respective panels. (e) Raloxifene activates AhR target genes CYP1A1 and NQO1 in mouse WT Hepa1 cells. (f) Activation of AhR target genes CYP1A1 and NQO1 by raloxifene is dependent on the presence of transcriptionally active ARNT. (g) Raloxifene activates AhR target genes CYP1A1 and NQO1 in human HepG2 cells. (h) Activation of the AhR target gene CYP1A1 by raloxifene in MDA-MB-231 cells. GAPDH expression is shown as a control for all semi-quantitative RT-PCR experiments, and cycle numbers of PCR reaction sampling (non-saturated) are indicated. All treatments for RT-PCR analysis were performed overnight
Raloxifene is an AhR ligand
To determine whether raloxifene is a ligand of the AhR, we first performed molecular docking studies with raloxifene using an optimized homology model of the human AhR ligand-binding pocket. A raloxifene-guided optimization step of the model previously developed in our laboratory33 produced significant changes at the level of volume and area of the binding pocket, which were attributed to the different size and shape of raloxifene compared with the planar and symmetric TCDD scaffold (Figure 2a). The optimized binding pocket extended towards the Gβ and Iβ sheets in order to allocate the 4-hydroxyphenyl-benzothiophene and piperidyl rings of raloxifene (Figure 2a). TCDD was able to dock into the pocket with a score of −21.8, establishing a hydrogen bond (HB) between oxygen and the side chain of Gln 383 (Iβ) (Figure 2b). Instead, raloxifene docked with the lower score of −10.3 exhibiting a binding pattern different from that of TCDD, as it did not involve Gln 383 (Iβ) due to unfavorable energy in that conformation. Raloxifene established HB interactions with a ketone carbonyl, the thiol group of Cys 333 (Fα), and the 6-hydroxyl and carbonyl of the backbone of Ile 349 (Gβ) and Val 363 (Hβ), respectively (Figure 2b). The docking scores obtained predicted a lower binding affinity for raloxifene than TCDD, which was consistent with the higher concentrations of raloxifene needed to activate the AhR (Figure 1).
Raloxifene is a ligand of the AhR. (a) Comparison of the binding pocket size and shape of the raloxifene-optimized AhR PAS-B homology model and our previous (TCDD) optimized model.33 The binding pockets are characterized with ICM Pocket Finder and colored in black (original model) and gray (optimized model). The protein backbone is displayed as ribbons and colored by N to C. (b) Docking orientation of TCDD and raloxifene into the human raloxifene-optimized AhR PAS-B homology model with the protein backbone displayed as ribbon and colored by secondary structure. TCDD and raloxifene are displayed as sticks and colored by atom type, with the carbon atoms in magenta (TCDD) and orange (Raloxifene). Residues are displayed as sticks and colored by atom type with carbon atom in green. Hydrogen bonds are displayed as black dashed lines. Molecular modeling, docking, and visualization were performed with ICM v3.7-2d. (c) Competitive binding assay with [3H]-3-Methylcholanthrene and dose–response of TCDD or raloxifene. IC50 TCDD: 3.6 × 10−9 M; IC50 Raloxifene: 9.8 × 10−5 M
We next used a ligand competition assay employing cytosolic extracts from ER-negative, AhR-expressing Hepa1 cells and radiolabeled 3-methylcholanthrene ([3H]-3MC) to evaluate direct binding of raloxifene to the AhR.34 Raloxifene displaced [3H]-3MC (Figure 2c), suggesting that raloxifene binds the AhR. To provide additional evidence of direct binding, we evaluated the ability of raloxifene to delay subtilisin-mediated proteolysis of the AhR. Incubation of in vitro translated AhR with raloxifene resulted in delayed AhR proteolysis and differential formation of proteolysis products (Supplementary Figure S1). Taken together, these data suggest that raloxifene is a ligand of the AhR.
Raloxifene induces cell death in human hepatoma and breast cancer cells
During characterization of AhR activation by raloxifene, we observed inhibition of growth and signs of cell death in Hepa1, HepG2, and MDA-MB-231 cells. Specifically, treatment with raloxifene for 48 h induced dramatic cell death evidenced by cell rounding, membrane blebbing, and loss of plate adhesion (Figure 3a). Overnight incubation of MDA-MB-231 cells with raloxifene induced similar effects, with clear evidence of apoptosis as indicated by nuclear condensation and fragmentation (Figure 3b). These data suggested that raloxifene induces a growth inhibitory effect in both hepatoma and ER-negative breast cancer cells. We also performed cell viability assays to quantitatively assess the effects of raloxifene. Raloxifene significantly decreased the number of Hepa1 cells in a dose- and time-dependent manner, with 20 μM raloxifene reducing Hepa1 cell viability at 48 and 72 h by 75% and 95%, respectively (Figure 3c). Likewise, raloxifene significantly reduced viability of human HepG2 cells after 72 h (Figure 3d), while MDA-MB-231 cells treated with raloxifene exhibited significantly reduced incorporation of BrdU (Supplementary Figure S2).
Raloxifene induces cell death. (a) Raloxifene induces cell death in mouse and human hepatoma cells. Phase contrast microscopy of Hepa1 and HepG2 cells treated for 48 h with either vehicle (DMSO) or raloxifene (40 μM). (b) Raloxifene induces apoptosis in MDA-MB-231 cells. Phase contrast microscopy (top panels) and fluorescence microscopy of DAPI-stained nuclei (bottom panels) of MDA-MB-231 cells treated with DMSO or 40 μM raloxifene for 24 h. (c) Viability of Hepa1 cells treated for the indicated treatments and time points was determined by MTS assay. (d) Viability of HepG2 cells treated with vehicle or raloxifene was determined at 72 h by MTS assay. (e) Raloxifene-mediated inhibition of proliferation in rat hepatoma cells is AhR-dependent. A western blot shows relative levels of AhR in AhR expressing (5 L) and low-expressing (BP8) rat hepatoma cells. Cells were treated as indicated for 24 h after which BrdU incorporation was analyzed. BrdU results are the mean±s.e.m. of four biological replicates. (f) Stable re-expression of AhR in an AhR-low mouse hepatoma line (C12+AhR and C12, respectively) rescues the antiproliferative effects of raloxifene. Cells were treated as indicated for 72 h, and cell viability was measured by MTS assay. Results representative of three similar experiments. For all experiments, results are the mean±s.e.m. of three independent determinations unless otherwise indicated; *P<0.05, #P<0.01 ‡P<0.001. For cell photographs, bars indicate equal image size within respective experiments
Requirement of AhR expression for Raloxifene-induced apoptosis in hepatoma cells
Having determined that raloxifene is both an AhR ligand and induces significant growth inhibition and apoptosis in mouse Hepa1 and human HepG2 and MDA-MB-231 cells at concentrations consistent with AhR activation, we next determined whether AhR signaling mediates both the antiproliferative and apoptotic effects of raloxifene. To investigate the role of AhR in mediating the effects of raloxifene in hepatoma cells, we assessed cell viability and apoptosis of hepatoma cells with differential AhR expression after treatment with raloxifene. We first employed the rat hepatoma cell culture model of 5L and BP8 cells, which are AhR-positive and AhR-negative, respectively, and have been used previously to demonstrate the AhR-dependent antiproliferative effects of the AhR ligand TCDD.12, 35, 36 AhR-deficient BP8 cells treated with raloxifene exhibited significantly higher BrdU incorporation than AhR-expressing 5L cells (Figure 3e). Consistent with these data, Hepa1 cells exhibited decreased viability compared with derivative TAO cells that have 90% reduced AhR expression (data not shown). We also employed an independently generated set of mouse hepatoma cell lines with low-AhR expression (C12) and C12+AhR cells, which are stably transfected with mouse AhR.37 The viability of C12+AhR cells was significantly decreased upon treatment with raloxifene compared with C12 cells (Figure 3f). Induction of AhR target genes by raloxifene required expression of the AhR heterodimerization partner ARNT (Figure 1f). To determine whether induction of apoptosis by raloxifene in hepatoma cells also required canonical AhR signaling, we evaluated the effect of ARNT expression on raloxifene-mediated cell death. Consistent with gene induction data, cells expressing functional ARNT showed significantly decreased proliferation compared with C4 cells (data not shown).
We next determined the extent to which apoptosis contributes to raloxifene-induced growth inhibition in mouse hepatoma cells. Raloxifene strongly induced apoptosis after 48 h of treatment compared with vehicle-treated Hepa1 cells (% nuclear fragmentation for vehicle: 4.7±1.8; raloxifene: 47.0±4.0, P=0.0007) (Supplementary Figure S3).
Despite the ability of TCDD to strongly activate AhR signaling, TCDD did not induce apoptosis (data not shown), suggesting a ligand-specific effect. AhR expression was required for increased apoptosis in Hepa1 cells compared with TAO cells (Figures 4a and b), strongly suggesting that the effects were AhR-dependent. To confirm these findings, we analyzed the effects of raloxifene on induction of apoptosis in C12 and C12+AhR cells. Consistent with the observed decreases in cell viability, re-expression of AhR in C12 cells rescued induction of apoptosis by raloxifene (Figure 4c). Likewise, apoptosis induced by raloxifene was significantly increased in vT{2} cells compared with C4 cells (Figure 4d). Taken together, these results indicated that induction of apoptosis by raloxifene in mouse hepatoma cells was significantly dependent on the activity of both AhR and ARNT.
Induction of apoptosis by raloxifene is AhR-dependent. Apoptosis was quantified by analysis of nuclear fragmentation (a) and caspase 3/7 activation (b) apoptosis assays in Hepa1 and TAO cells. (a, left) Western blot depicting relative AhR expression between Hepa1 cells and TAO cells is shown with GAPDH as an equal loading control. (c) Nuclear fragmentation assay in C12+AhR and C12 cells. (c, left) Western blot depicting relative AhR expression between C12 and C12+AhR cells is shown with alpha-tubulin as an equal loading control. (d) Raloxifene-induced apoptosis requires ARNT. Apoptosis was determined by nuclear fragmentation assay. For all experiments, unless indicated otherwise, results are the mean±s.e.m. of three biological replicates and representative of at least two independent experiments. Relevant statistically significant differences are indicated
Having shown that the induction of apoptosis by raloxifene required AhR expression in Hepa1 cells, we next confirmed the effects in human cell lines. The extent of apoptosis induced by raloxifene in HepG2 cells was similar to that of Hepa1 cells (Supplementary Figure S3). To further evaluate the AhR-dependent effects of raloxifene in hepatoma cells, the effects of raloxifene were tested on human HepG2 cells stably expressing scrambled shRNA (shScram) or AhR shRNA (shAhR) (Figure 5). Consistent with our results obtained thus far, knockdown of AhR in HepG2 cells conferred a significant increase in viability compared with cells expressing shScram (Figure 5).
Induction of apoptosis by raloxifene is AhR-dependent. (a) AhR knockdown in shAhR expressing HepG2 cells significantly reduces the antiproliferative effects of raloxifene compared with shScram after 72 h. Vehicle: 5.0%±0.5 versus raloxifene: 36.2%±0.7, P<0.0001. (a) Western blot showing relative AhR levels in HepG2 cells stably expressing a shRNA against a non-targeting (shScram) sequence or AhR (shAhR) is shown with GAPDH as an equal loading control. (b) Raloxifene inhibits cell viability compared with vehicle after 24 h in human HepG2 hepatoma cells in an AhR-dependent manner. Results are the mean±s.e.m. of three independent experiments
Raloxifene-induced apoptosis in ER-negative MDA-MB-231 breast cancer cells is AhR-dependent
Raloxifene activated AhR signaling (Figures 1d and h) and induced apoptosis in MDA-MB-231 cells (Figure 3b). Given that the AhR-dependent antiproliferative effects of raloxifene were observed in ER-negative hepatoma cells, we next investigated the effects of raloxifene in ER-negative/AhR-positive breast cancer cells. To this end, we employed two independent AhR knockdown strategies in MDA-MB-231 cells. Transient knockdown of AhR significantly decreased raloxifene-induced nuclear fragmentation in MDA-MB-231 cells (Figure 6a). We also generated a stable cell line (MDA-MB-231-pTRIPZ-shAhR1) in which AhR knockdown was induced by addition of doxycycline (DOX) to the cell culture media via expression of an shAhR hairpin with a RFP reporter (Figure 6b). Real time cellular analysis revealed that MDA-MB-231 cells without DOX (normal AhR expression) exhibited increased sensitivity to raloxifene compared with MDA-MB-231 cells with AhR knockdown (Figure 6c). Likewise, increased caspase 3/7 activation by raloxifene (in MDA-MB-231-pTRIPZ-shAhR1 cells without DOX) was suppressed by suppression of AhR expression (by the addition of DOX) (Figure 6d). These data were in good agreement with our observations in hepatoma cells and, taken together, strongly indicate that the induction of apoptosis in MDA-MB-231 cells by raloxifene was significantly dependent on AhR expression. To determine whether the effects of raloxifene could be selective towards cancer cells, we compared the effects of raloxifene on MDA-MB-231 cells with MCF-10A non-transformed breast cells, both of which express similar levels of AhR. Importantly, MDA-MB-231 cells exhibited increased sensitivity to raloxifene in a dose- and time-dependent manner compared with MCF-10A cells (Figure 6e).
AhR-dependent effects of raloxifene in human hepatoma and ER-negative breast cancer. (a) Transient knockdown of AhR in MDA-MB-231 cells significantly reduces nuclear fragmentation by raloxifene (RLX, 20 μM) compared with vehicle. (b) Inducible knockdown of AhR in MDA-MB-231-pTRIPZ-shAhR treated with or without doxycycline (DOX, 2 μg/ml) was verified by western blot. Flow cytometry analysis of a co-expressed RFP reporter indicated the uniformity and extent of shRNA expression. (c) Relative cell death index (RCDI) of MDA-MB-231-pTRIPZ-shAhR cells with and without DOX treated with raloxifene. Increasing RCDI is indicative of increasing cell death and calculated by subtracting the normalized cell index of vehicle (0.1% DMSO)-treated cells from that of raloxifene-treated cells in the absence or presence of DOX. Results are the mean of two independent determinations. (d) Caspase 3/7 activity is significantly reduced in MDA-MB-231 cells with decreased AhR expression. (e) Breast cancer cells exhibit increased sensitivity to raloxifene compared with normal breast epithelial cells. MDA-MB-231 cells exhibit increased sensitivity to raloxifene compared with non-transformed MCF-10A breast epithelial cells. Cells were treated with raloxifene for 48 and 72 h and viability was determined. **P<0.001; ****P<0.0001. Results are the mean±s.d. of three independent experiments
AhR is a potential molecular target for the treatment of ER-negative breast cancer
Our results indicated that raloxifene induces apoptosis in ER-negative breast cancer cells in an AhR-dependent manner. To determine the utility of the AhR an anticancer target, we next determined whether AhR is expressed in different breast cancer subtypes. We also investigated the prognostic value of AhR expression in different breast cancer subtypes in terms of overall survival, distant metastasis-free survival, and relapse-free survival using a breast cancer-specific Kaplan–Meier Plotter analysis tool.38 We found that higher AhR expression was associated with statistically significant increased overall survival and distant metastasis-free survival (Figures 7a and b). In addition, these data indicate that AhR is expressed in both ER-positive and ER-negative cancers (Figures 7 and 8).
Effect of AhR expression on breast cancer outcomes. (a–i) Kaplan–Meier plots depicting overall survival (OS), distant metastasis-free survival (DMFS), and relapse-free survival (RFS) segregated based on AhR expression and the indicated breast cancer subtypes. The cutoff for AhR expression was selected from between the upper and lower patient quartiles, and reflected the level of AhR expression associated with the best difference between the indicated survival type. Beeswarm plots (inset within each plot) show the specific cutoff value, and patient numbers for each survival group are indicated. ER status was determined based on microarray data from individual tumor data. The horizontal dashed lines in beeswarm plots indicate an arbitrary expression level of 2000 and is shown for comparison purposes. Refer to Supplementary Figure S4 for data analyzed based on ER status determined by immunotyping
AhR expression is significantly associated with relapse-free survival in ER/PR-negative breast cancer. (a–e) Examination of patient survival data showing that AhR is significantly associated with relapse-free survival. Beeswarm plots (right panels) show the optimized AhR expression cutoffs for the respective survival curves. PR status is based on clinical determination (immunotyping) and ER status were based on microarray expression data. The cutoff for AhR expression was selected from between the upper and lower patient quartiles and reflected the level of AhR expression associated with the best difference between the indicated survival type. §, the corresponding data are presented in Figure 7; * data set too small for analysis. The horizontal dashed lines in beeswarm plots indicate an arbitrary expression level of 2000 and is shown for comparison purposes
We found that high expression of the AhR strongly correlated with better patient survival in several breast cancer contexts. For example, in a population of 1027 breast cancer patients, those in the upper 75% based on AhR expression exhibited better overall survival (Figure 7a, P<0.05). Here, the hazard ratio (HR) serves as an indication of patient prognosis, with HR values <1 indicating better survival. Further analysis showed that distant metastasis-free survival in all breast cancer patients (n=1353) based on the upper 75% of AhR expression was associated with a 1.41-fold improved prognosis (HR−1, P=0.0046; Figure 7b). Interestingly, the prognostic value of the AhR expression was significantly better for ER-positive breast cancer than ER-negative breast cancer. The expression of AhR is associated with increased overall survival, distant metastasis-free survival, and relapse-free survival in ER-positive breast cancer (Figures 7d–f). Further, the log-rank P significance of distant metastasis-free survival in patients reached 2.6e−7 based on immunotyped ER-positive expression and high AhR expression (Supplementary Figure S4B). Comparison of patient survival based on microarray and clinical determination of ER status showed similar trends (Figure 7 and Supplementary Figure S4).
To look specifically at hormone-independent breast cancer subtype, we evaluated the effect of progesterone receptor (PR) expression in addition to that of ER. In ER- and PR-negative breast cancer, relapse-free survival was significantly improved for the upper two-thirds of patients based on AhR expression (Figure 8a, HR−1=2.22, P=0.0021). We also investigated relapse-free survival in specific breast cancer subtypes based on the St. Gallen criteria,39 comprising Luminal types A and B, HER2+, and basal subtypes (Supplementary Figure S5). Luminal type A breast cancer exhibited a statistically significant difference between AhR high and low expression groups. In addition, there was a modest improvement in relapse-free survival in patients with high AhR expression in the basal subset. Taken together, our data suggest that AhR expression in both hormone-dependent and -independent cancers is associated with improved patient survival in certain contexts. Targeting these patient subsets with AhR-mediated therapeutics (such as raloxifene) is thus an exciting possibility for improving patient outcome.
Discussion
In the pursuit of new treatments for breast cancer, it is vital not only to develop new therapeutic agents for known drug targets, but also to identify and characterize novel molecular targets of existing drugs for re-tooling in order to expand the arsenal of treatment options available to breast cancer patients. While the AhR has been studied in the past as the mediator of TCDD toxicity1, 10, 40, 41 as well other environmental carcinogens, recent studies strongly suggest that the AhR has the potential to be an effective target for cancer therapeutics.6, 7, 16, 17, 42, 43, 44, 45, 46, 47 A structurally diverse array of small molecules, some of which are used in the clinic, have been shown to be ligands of the AhR.14, 15, 19, 46 Likewise, several compounds have been developed as selective modulators of the AhR, some of which show promise as anticancer agents.17
In the present study, we screened a library of pharmacologically active compounds to identify putative AhR ligands. This targeted screening approach was intentional, as identification of novel AhR activators from compounds with established clinical activity may expedite the AhR-based drug discovery process.19 During characterization of lead compounds, we found that raloxifene, a selective ER modulator used in the clinic is an AhR ligand (Figure 2) that activates AhR signaling (Figure 1). We evaluated the effects of raloxifene in Hepa1 and HepG2 cells and found that raloxifene decreased cell viability by inducing apoptosis in an AhR-dependent manner (Figures 3, 4, 5). Despite the strong AhR-dependent induction of apoptosis, we also found that overall cell viability was decreased in both AhR expressing and paired low/non-expressing cells, which likely reflected a combination of residual AhR expression in knockdown cell lines and AhR-independent effects of raloxifene.
AhR-dependent apoptosis induced by raloxifene in ER-negative hepatoma cells were extendable to MDA-MB-231 breast cancer cells. Specifically, both transient and stable AhR knockdown in MDA-MB-231 cells revealed that induction of apoptosis by raloxifene was significantly dependent upon AhR expression (Figures 6a–d). Comparison of the antiproliferative effects of raloxifene in non-transformed MCF-10A cells and MDA-MB-231 breast cancer cells showed that the latter had increased sensitivity to raloxifene, despite similar levels of AhR expression (Figure 6e). This was especially important given that the concentrations of raloxifene used in our study were higher than the bioavailable concentrations achieved using the standard dosing regimen of raloxifene (60 mg per day). It is unknown whether AhR-activating concentrations of raloxifene in ER-negative breast cancer patients can be achieved; however, animal testing indicates that acute dosing of raloxifene is very well tolerated, with no adverse effects of the drug observed at single oral doses of raloxifene up 5000 mg/kg in rats and mice and 1000 mg/kg in monkeys (Evista product insert). Further, a study evaluating bone turnover, serum lipids, and endometrium in post-menopausal women showed that raloxifene is well tolerated up to 600 mg/kg/day for 8 weeks.48 These results were confirmed in a subsequent 8-week study of 63 post-menopausal women, given 600 mg/kg/day raloxifene.49 In support of the possibility of using raloxifene for the treatment of ER-negative breast cancer, a study published while this manuscript was in review showed that raloxifene given daily by oral route to mice inhibits xenografts of triple-negative MDA-MB-231 and MDA-MB-468 breast cancer cells and also induces modest tumor regression of MDA-MB-468 cell xenografts.50
Having identified an AhR-dependent apoptotic effect of raloxifene in triple-negative MDA-MB-231 breast cancer cells, we investigated at the ability of AhR expression to predict relapse-free survival of patients. Specifically, evaluation of relapse-free survival in breast cancer patients indicated that higher levels of AhR are associated with better overall survival compared with those expressing lower levels of AhR (Figure 7a). In extending these results to ER-positive and ER-negative breast cancer subsets, we found that higher AhR expression was significantly associated with better relapse-free survival, overall survival, and distant metastasis-free survival in ER-positive breast cancers, strongly supporting a tumor suppressor role of the AhR (Supplementary Figures S4A–C and Figures 7e and f). Further, while patients with ER-negative breast cancers exhibited mixed effects with respect to AhR expression on survival outcomes (Figure 7, lower panels, and Supplementary Figures S4D–F), specific analysis of ER- and PR-negative breast cancers revealed a significant association between high AhR expression and better relapse-free survival (Figure 8). These data support a tumor suppressor role of the AhR in ER-positive and ER- and PR-negative breast cancer subsets. Thus, patients with hormone-independent breast cancers and high AhR expression may be uniquely primed for treatment with AhR modulators that activate its tumor-suppressive functions.
In conclusion, our data indicate that raloxifene is a novel ligand of the AhR and induces AhR-dependent apoptosis in ER-negative hepatoma and breast cancer cells. These results open up the possibility of re-tooling raloxifene for treatment of patients diagnosed with hormone-independent or triple-negative breast cancers. In the context of ER- and PR-negative breast cancer, the identification of the AhR as a novel anti-breast cancer target has important therapeutic implications – namely, identification of ligands of the AhR with functional effects similar to those of raloxifene. Such molecules would represent a new class of therapeutics specifically for AhR-expressing cancers such as triple-negative breast cancers.
Materials and Methods
Cell lines and reagents
All cells (except MCF10) were cultured in DMEM (Mediatech, Manassas, VA, USA) supplemented with 10% FBS (Tissue Culture Biologicals, Seal Beach, CA, USA), 100 U/ml penicillin, and 100 mg/ml streptomycin (Mediatech). MCF-10A cells were cultured with DMEM/F12 supplemented with 5% horse serum (Invitrogen, Carlsbad, CA, USA), 20 ng/ml EGF, 0.5 mg/ml hydrocortisone, 100 ng/ml cholera toxin, and 10 μg/ml insulin, 100 U/ml penicillin, and 100 mg/ml streptomycin. All cells were grown at 37 °C in a humidified 5% CO2 atmosphere. Raloxifene was purchased from Enzo (Plymouth Meeting, PA, USA). All other reagents were purchased from Sigma (St Louis, MO, USA).
AhR activation assays
Reporter gene assays, immunofluorescence studies for AhR localization, semi-quantitative Real Time PCR for analysis of raloxifene-mediated induction of AhR target genes in Hepa1, MDA-MB-231, and HepG2 cells were conducted as previously described.19, 33 Primer sequences for mouse and human GAPDH, CYP1A1, and NQO1 have been described previously.19 To evaluate the requirement for ARNT, we employed a pair of mouse hepatoma cells, one of which expresses a transcriptionally inactive mutant ARNT protein (C4 cells), and C4 cells stably re-expressing WT ARNT (vT{2} cells).
Homology modeling
The homology model of human AhR-LBD bound to TCDD was initially built as described previously by our laboratory.33 The complex was then submitted to 105 steps of ligand–protein side chain optimization using Monte Carlo (MC) simulation in the internal coordinate space with Molsoft ICM.51, 52 The resulting best energy conformation was then used to dock raloxifene. The resulting complex with docked ligand submitted again to 104 steps of MC ligand–protein side chain optimization to reach the most energetically favorable conformation of the AhR-LBD bound to raloxifene. Molecular Docking of AHR agonists TCDD and raloxifene was performed as described previously.33
In the ICM-VLS (Molsoft ICM)-screening procedure, the ligand scoring is optimized to obtain maximal separation between the binders and non-binders. Each compound is assigned a score according to its fit within the receptor; this ICM score accounts for continuum and discreet electrostatics, hydrophobicity, and entropy parameters.51, 53
Competitive binding assays
Competitive binding assays were performed as described previously.34 ER-negative WT Hepa1 cells served as the source of the whole-cell extract used for the assays and was prepared as described previously.18 [3H]-3-Methylcholanthrene was purchased from Moravek (Brea, CA, USA).
Western blot analysis
Western blot analysis for relative AhR expression was performed as described previously.18, 33
AhR knockdown
HepG2 and MDA-M231 cells stably expressing shRNAs for constitutive (pLKO.1) and inducible (pTRIPZ) knockdown of AhR were generated as described previously.18 Briefly, lentiviral particles were produced in HEK293T cells via transfection of packaging and envelope vectors psPAX2 and pMD2.G, respectively. AhR knockdown was routinely confirmed by western blot.
For transient knockdown of AhR in MDA-MB-231, cells were transfected using Dharmafect transfection reagent and siRNA duplexes against luciferase (siLuc, control) or human AhR (siAhR) (Thermo Scientific, Waltham, MA, USA). Knockdowns of AhR of greater than 75% were achieved within 24 h after transfection, and treatments were begun within 24–48 h, depending on cell density. Knockdowns were verified by western blot.
Proliferation and apoptosis assays
Cell viability was assayed using either the CellTiter AQueous Non-Radioactive Cell Proliferation Assay (MTS, Huntsville, AL, USA) or CellTiter Luminescent Cell Viability Assay (Promega, Madison, WI, USA) as described previously.18 BrdU incorporation was assayed using a BrdU Cell Proliferation Assay according to the manufacturer’s suggested protocol (Calbiochem, Billerica, MA, USA). Real time analysis of cellular proliferation by xCELLligence assay was performed as described previously.18
Apoptosis was determined by nuclear fragmentation assay as described previously.54, 55, 56, 57 Quantifications were performed in triplicate, with each count consisting of at least 300 cells. Caspase 3/7 activation was evaluated by Caspase 3/7 kit (Promega) according to the manufacturer’s suggested protocol.
Analysis of relapse-free survival in patients with ER-negative and -positive breast cancer
The effect of AhR expression on relapse-free survival in patients with ER-negative and -positive breast cancer was evaluated by Kaplan–Meier analysis using the online tool KMPLOT based on the updated 2012 data set.38 Data for AhR was accessed using AffymetrixID probe 202820_at. AhR low and high expression was determined based on the optimized expression level for the best difference between the two expression (patient survival) groups, the values of which were reported.
Data analysis
Data were analyzed with Graphpad Prism version 5.0 using Student’s t-test or one-way ANOVA with Tukey’s post test. Values of P<0.05 were considered to be statistically significant.
Abbreviations
- 3MC:
-
3-methylcholanthrene
- 6-MCDF:
-
6-methyl-1,3,8-trichlorodibenzofuran
- AhR:
-
aryl hydrocarbon receptor
- ARNT:
-
aryl hydrocarbon receptor nuclear translocator
- DEN:
-
diethyl nitrosamine
- DMBA:
-
7,12-dimethylbenz(a)anthracene
- E2:
-
estradiol, 17β-estradiol, oestradiol
- ER:
-
estrogen receptor
- HR:
-
hazard ratio
- HSP90:
-
heat shock protein 90
- PAS:
-
Per-ARNT-Sim
- PR:
-
progesterone receptor
- TCDD:
-
2,3,7,8-Tetrachlorodibenzo-p-dioxin
- TRAMP:
-
transgenic adenocarcinoma of the mouse prostate
- XAP2:
-
hepatitis B virus X-associated protein
- XRE:
-
xenobiotic response element
References
Beischlag TV, Luis Morales J, Hollingshead BD, Perdew GH . The aryl hydrocarbon receptor complex and the control of gene expression. Crit Rev Eukaryot Gene Expr 2008; 18: 207–250.
Dragan YP, Schrenk D . Animal studies addressing the carcinogenicity of TCDD (or related compounds) with an emphasis on tumour promotion. Food Addit Contam 2000; 17: 289–302.
Kociba RJ, Keyes DG, Beyer JE, Carreon RM, Wade CE, Dittenber DA et al. Results of a two-year chronic toxicity and oncogenicity study of 2,3,7,8-tetrachlorodibenzo-p-dioxin in rats. Toxicol Appl Pharmacol 1978; 46: 279–303.
Ray S, Swanson HI . Activation of the aryl hydrocarbon receptor by TCDD inhibits senescence: a tumor promoting event? Biochem Pharmacol 2009; 77: 681–688.
Viluksela M, Bager Y, Tuomisto JT, Scheu G, Unkila M, Pohjanvirta R et al. Liver tumor-promoting activity of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) in TCDD-sensitive and TCDD-resistant rat strains. Cancer Res 2000; 60: 6911–6920.
Fritz WA, Lin TM, Cardiff RD, Peterson RE . The aryl hydrocarbon receptor inhibits prostate carcinogenesis in TRAMP mice. Carcinogenesis 2007; 28: 497–505.
Fritz WA, Lin TM, Safe S, Moore RW, Peterson RE . The selective aryl hydrocarbon receptor modulator 6-methyl-1,3,8-trichlorodibenzofuran inhibits prostate tumor metastasis in TRAMP mice. Biochem Pharmacol 2009; 77: 1151–1160.
Fritz WA, Lin TM, Peterson RE . The aryl hydrocarbon receptor (AhR) inhibits vanadate-induced vascular endothelial growth factor (VEGF) production in TRAMP prostates. Carcinogenesis 2008; 29: 1077–1082.
Fan Y, Boivin GP, Knudsen ES, Nebert DW, Xia Y, Puga A . The aryl hydrocarbon receptor functions as a tumor suppressor of liver carcinogenesis. Cancer Res 2010; 70: 212–220.
Mimura J, Yamashita K, Nakamura K, Morita M, Takagi TN, Nakao K et al. Loss of teratogenic response to 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) in mice lacking the Ah (dioxin) receptor. Genes Cells 1997; 2: 645–654.
Kawajiri K, Kobayashi Y, Ohtake F, Ikuta T, Matsushima Y, Mimura J et al. Aryl hydrocarbon receptor suppresses intestinal carcinogenesis in ApcMin/+ mice with natural ligands. Proc Natl Acad Sci USA 2009; 106: 13481–13486.
Kolluri SK, Weiss C, Koff A, Gottlicher M . p27(Kip1) induction and inhibition of proliferation by the intracellular Ah receptor in developing thymus and hepatoma cells. Genes Dev 1999; 13: 1742–1753.
Ito T, Tsukumo S, Suzuki N, Motohashi H, Yamamoto M, Fujii-Kuriyama Y et al. A constitutively active arylhydrocarbon receptor induces growth inhibition of jurkat T cells through changes in the expression of genes related to apoptosis and cell cycle arrest. J Biol Chem 2004; 279: 25204–25210.
Denison MS, Nagy SR . Activation of the aryl hydrocarbon receptor by structurally diverse exogenous and endogenous chemicals. Ann Rev Pharmacol Toxicol 2003; 43: 309–334.
Hu W, Sorrentino C, Denison MS, Kolaja K, Fielden MR . Induction of cyp1a1 is a nonspecific biomarker of aryl hydrocarbon receptor activation: results of large scale screening of pharmaceuticals and toxicants in vivo and in vitro. Mol Pharmacol 2007; 71: 1475–1486.
Safe S, Qin C, McDougal A . Development of selective aryl hydrocarbon receptor modulators for treatment of breast cancer. Expert Opin Investig Drugs 1999; 8: 1385–1396.
Safe S, McDougal A . Mechanism of action and development of selective aryl hydrocarbon receptor modulators for treatment of hormone-dependent cancers (Review). Int J Oncol 2002; 20: 1123–1128.
O'Donnell EF, Kopparapu PR, Koch DC, Jang HS, Phillips JL, Tanguay RL et al. The aryl hydrocarbon receptor mediates leflunomide-induced growth inhibition of melanoma cells. PloS One 2012; 7: e40926.
O'Donnell EF, Saili KS, Koch DC, Kopparapu PR, Farrer D, Bisson WH et al. The anti-inflammatory drug leflunomide is an agonist of the aryl hydrocarbon receptor. PLoS One 2010; 5: e13128.
Jones CD, Jevnikar MG, Pike AJ, Peters MK, Black LJ, Thompson AR et al. Antiestrogens. 2. Structure-activity studies in a series of 3-aroyl-2-arylbenzo[b]thiophene derivatives leading to [6-hydroxy-2-(4-hydroxyphenyl)benzo[b]thien-3-yl] [4-[2-(1-piperidinyl)ethoxy]-phenyl]methanone hydrochloride (LY156758), a remarkably effective estrogen antagonist with only minimal intrinsic estrogenicity. J Med Chem 1984; 27: 1057–1066.
Clemens JA, Bennett DR, Black LJ, Jones CD . Effects of a new antiestrogen, keoxifene (LY156758), on growth of carcinogen-induced mammary tumors and on LH and prolactin levels. Life Sci 1983; 32: 2869–2875.
Kleinberg DL, Todd J, Babitsky G . Inhibition by estradiol of the lactogenic effect of prolactin in primate mammary tissue: reversal by antiestrogens LY 156758 and tamoxifen. Proc Natl Acad Sci USA 1983; 80: 4144–4148.
Jordan VC . Selective estrogen receptor modulation: a personal perspective. Cancer Res 2001; 61: 5683–5687.
Jordan VC, Phelps E, Lindgren JU . Effects of anti-estrogens on bone in castrated and intact female rats. Breast Cancer Res Treat 1987; 10: 31–35.
Ettinger B, Black DM, Mitlak BH, Knickerbocker RK, Nickelsen T, Genant HK et al. Reduction of vertebral fracture risk in postmenopausal women with osteoporosis treated with raloxifene: results from a 3-year randomized clinical trial. Multiple Outcomes of Raloxifene Evaluation (MORE) Investigators. JAMA 1999; 282: 637–645.
Vogel VG . Managing the risk of invasive breast cancer in women at risk for breast cancer and osteoporosis: the role of raloxifene. Clinical Interv Aging 2008; 3: 601–609.
Vogel VG, Costantino JP, Wickerham DL, Cronin WM, Cecchini RS, Atkins JN et al. Effects of tamoxifen versus raloxifene on the risk of developing invasive breast cancer and other disease outcomes: the NSABP Study of Tamoxifen and Raloxifene (STAR) P-2 trial. JAMA 2006; 295: 2727–2741.
Okamoto Y, Liu X, Suzuki N, Okamoto K, Sekimoto M, Laxmi YR et al. Increased antitumor potential of the raloxifene prodrug, raloxifene diphosphate. Int J Cancer 2008; 122: 2142–2147.
Barkhem T, Andersson-Ross C, Hoglund M, Nilsson S . Characterization of the ‘estrogenicity’ of tamoxifen and raloxifene in HepG2 cells: regulation of gene expression from an ERE controlled reporter vector versus regulation of the endogenous SHBG and PS2 genes. J Steroid Biochem Mol Biol 1997; 62: 53–64.
Kim IY, Kim BC, Seong DH, Lee DK, Seo JM, Hong YJ et al. Raloxifene, a mixed estrogen agonist/antagonist, induces apoptosis in androgen-independent human prostate cancer cell lines. Cancer Res 2002; 62: 5365–5369.
Kim IY, Seong DH, Kim BC, Lee DK, Remaley AT, Leach F et al. Raloxifene, a selective estrogen receptor modulator, induces apoptosis in androgen-responsive human prostate cancer cell line LNCaP through an androgen-independent pathway. Cancer Res 2002; 62: 3649–3653.
Stuart EC, Rosengren RJ . The combination of raloxifene and epigallocatechin gallate suppresses growth and induces apoptosis in MDA-MB-231 cells. Life Sci 2008; 82: 943–948.
Bisson WH, Koch DC, O'Donnell EF, Khalil SM, Kerkvliet NI, Tanguay RL et al. Modeling of the aryl hydrocarbon receptor (AhR) ligand binding domain and its utility in virtual ligand screening to predict new AhR ligands. J Med Chem 2009; 52: 5635–5641.
Denison MS, Rogers JM, Rushing SR, Jones CL, Tetangco SC, Heath-Pagliuso S . Analysis of the aryl hydrocarbon receptor (AhR) signal transduction pathway. Curr Protoc Toxicol 2002 Chapter 4: Unit 4.8.
Kolluri SK, Balduf C, Hofmann M, Gottlicher M . Novel target genes of the Ah (dioxin) receptor: transcriptional induction of N-myristoyltransferase 2. Cancer Res 2001; 61: 8534–8539.
Weiss C, Kolluri SK, Kiefer F, Gottlicher M . Complementation of Ah receptor deficiency in hepatoma cells: negative feedback regulation and cell cycle control by the Ah receptor. Exp Cell Res 1996; 226: 154–163.
Wang F, Zhang R, Shi S, Hankinson O . The effect of aromatic hydrocarbon receptor on the phenotype of the Hepa 1c1c7 murine hepatoma cells in the absence of dioxin. Gene Regul Syst Biol 2007; 1: 49–56.
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q et al. An online survival analysis tool to rapidly assess the effect of 22 277 genes on breast cancer prognosis using microarray data of 1809 patients. Breast Cancer Res Treat 2009; 123: 725–731.
Mihály Z, Győrffy B . Improving pathological assessment of breast cancer by employing array-based transcriptome analysis. Microarrays 2013; 2: 228–242.
Fernandez-Salguero PM, Hilbert DM, Rudikoff S, Ward JM, Gonzalez FJ . Aryl-hydrocarbon receptor-deficient mice are resistant to 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced toxicity. Toxicol Appl Pharmacol 1996; 140: 173–179.
Bunger MK, Glover E, Moran SM, Walisser JA, Lahvis GP, Hsu EL et al. Abnormal liver development and resistance to 2,3,7,8-tetrachlorodibenzo-p-dioxin toxicity in mice carrying a mutation in the DNA-binding domain of the aryl hydrocarbon receptor. Toxicol Sci 2008; 106: 83–92.
Ma Q, Whitlock JP Jr . The aromatic hydrocarbon receptor modulates the Hepa 1c1c7 cell cycle and differentiated state independently of dioxin. Mol Cell Biol 1996; 16: 2144–2150.
Matthews J, Gustafsson JA . Estrogen receptor and aryl hydrocarbon receptor signaling pathways. Nucl Recept Signal 2006; 4: e016.
Gasiewicz TA, Henry EC, Collins LL . Expression and activity of aryl hydrocarbon receptors in development and cancer. Crit Rev Eukaryot Gene Expr 2008; 18: 279–321.
Hall JM, Barhoover MA, Kazmin D, McDonnell DP, Greenlee WF, Thomas RS . Activation of the aryl-hydrocarbon receptor inhibits invasive and metastatic features of human breast cancer cells and promotes breast cancer cell differentiation. Mol Endocrinol 2010; 24: 359–369.
Goodale BC, La Du JK, Bisson WH, Janszen DB, Waters KM, Tanguay RL . AHR2 mutant reveals functional diversity of aryl hydrocarbon receptors in zebrafish. PloS One 2012; 7: e29346.
Khan S, Barhoumi R, Burghardt R, Liu S, Kim K, Safe S . Molecular mechanism of inhibitory aryl hydrocarbon receptor-estrogen receptor/Sp1 cross talk in breast cancer cells. Mol Endocrinol 2006; 20: 2199–2214.
Draper MW, Flowers DE, Huster WJ, Neild JA, Harper KD, Arnaud C . A controlled trial of raloxifene (LY139481) HCl: impact on bone turnover and serum lipid profile in healthy postmenopausal women. J Bone Miner Res 1996; 11: 835–842.
Boss SM, Huster WJ, Neild JA, Glant MD, Eisenhut CC, Draper MW . Effects of raloxifene hydrochloride on the endometrium of postmenopausal women. Am J Obstet Gynecol 1997; 177: 1458–1464.
Taurin S, Allen KM, Scandlyn MJ, Rosengren RJ . Raloxifene reduces triple-negative breast cancer tumor growth and decreases EGFR expression. Int J Oncol 2013; 43: 785–792.
Abagyan R, Totrov M, Kuznetsov D . ICM—A new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J Computat Chem 1994; 15: 488–506.
Tosh DK, Phan K, Gao ZG, Gakh AA, Xu F, Deflorian F et al. Optimization of adenosine 5'-carboxamide derivatives as adenosine receptor agonists using structure-based ligand design and fragment screening. J Med Chem 2012; 55: 4297–4308.
Totrov M, Abagyan R . Flexible protein-ligand docking by global energy optimization in internal coordinates. Proteins 1997; 29 (Suppl 1): 215–220.
Kolluri SK, Zhu X, Zhou X, Lin B, Chen Y, Sun K et al. A short Nur77-derived peptide converts Bcl-2 from a protector to a killer. Cancer Cell 2008; 14: 285–298.
Li H, Kolluri SK, Gu J, Dawson MI, Cao X, Hobbs PD et al. Cytochrome c release and apoptosis induced by mitochondrial targeting of nuclear orphan receptor TR3. Science 2000; 289: 1159–1164.
Lin B, Kolluri SK, Lin F, Liu W, Han YH, Cao X et al. Conversion of Bcl-2 from protector to killer by interaction with nuclear orphan receptor Nur77/TR3. Cell 2004; 116: 527–540.
Kolluri SK, Corr M, James SY, Bernasconi M, Lu D, Liu W et al. The R-enantiomer of the nonsteroidal antiinflammatory drug etodolac binds retinoid X receptor and induces tumor-selective apoptosis. Proc Nat Acad Sci USA 2005; 102: 2525–2530.
Acknowledgements
This work was supported by the National Institute of Environmental Health Sciences (NIEHS; RES019000A) the U.S. Army Medical Research and Materiel Command and American Cancer Society (RSG-13-132-01-CDD). EFO was supported by the Department of Defense Breast Cancer Research Program pre-doctoral fellowship (W81XWH-10-1-0160) and a NIEHS training grant (T32 ES007060). DCK was supported by a National Research Service Award (1F31CA144571-01) pre-doctoral fellowship form the National Cancer Institute. We wish to thank Jessica Phillips, Vidya Schalk, Prasad Kopparapu, Soheila Kazemi, Cathy Duong, and Erin Albertson for technical assistance, Sam Bradford for excellent flow cytometry assistance, Drs Mark Leid, Nancy Kerkvliet, Robert Tanguay, Chrissa Kioussi, Andrew Buermeyer, Joseph Beckman, and Craig Marcus for very helpful discussions and the Oregon State University Cell Imaging and Analysis Facilities and Services Cores of the Environmental Health Sciences Center grant no. P30 ES000210, NIEHS, National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by RA Knight
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
O'Donnell, E., Koch, D., Bisson, W. et al. The aryl hydrocarbon receptor mediates raloxifene-induced apoptosis in estrogen receptor-negative hepatoma and breast cancer cells. Cell Death Dis 5, e1038 (2014). https://doi.org/10.1038/cddis.2013.549
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.549
Keywords
This article is cited by
-
Carbidopa suppresses estrogen receptor-positive breast cancer via AhR-mediated proteasomal degradation of ERα
Investigational New Drugs (2022)
-
Raloxifene potentiates the effect of gefitinib in triple-negative breast cancer cell lines
Medical Oncology (2022)
-
Discovery and Mechanistic Characterization of a Select Modulator of AhR-regulated Transcription (SMAhRT) with Anti-cancer Effects
Apoptosis (2021)
-
Activation of the Ah Receptor Modulates Gastrointestinal Homeostasis and the Intestinal Microbiome
Current Pharmacology Reports (2019)
-
Loss of NR2E3 represses AHR by LSD1 reprogramming, is associated with poor prognosis in liver cancer
Scientific Reports (2017)