Abstract
Autophagy is an important cellular process that controls cells in a normal homeostatic state by recycling nutrients to maintain cellular energy levels for cell survival via the turnover of proteins and damaged organelles. However, persistent activation of autophagy can lead to excessive depletion of cellular organelles and essential proteins, leading to caspase-independent autophagic cell death. As such, inducing cell death through this autophagic mechanism could be an alternative approach to the treatment of cancers. Recently, we have identified a novel autophagic inducer, saikosaponin-d (Ssd), from a medicinal plant that induces autophagy in various types of cancer cells through the formation of autophagosomes as measured by GFP-LC3 puncta formation. By computational virtual docking analysis, biochemical assays and advanced live-cell imaging techniques, Ssd was shown to increase cytosolic calcium level via direct inhibition of sarcoplasmic/endoplasmic reticulum Ca2+ ATPase pump, leading to autophagy induction through the activation of the Ca2+/calmodulin-dependent kinase kinase–AMP-activated protein kinase–mammalian target of rapamycin pathway. In addition, Ssd treatment causes the disruption of calcium homeostasis, which induces endoplasmic reticulum stress as well as the unfolded protein responses pathway. Ssd also proved to be a potent cytotoxic agent in apoptosis-defective or apoptosis-resistant mouse embryonic fibroblast cells, which either lack caspases 3, 7 or 8 or had the Bax-Bak double knockout. These results provide a detailed understanding of the mechanism of action of Ssd, as a novel autophagic inducer, which has the potential of being developed into an anti-cancer agent for targeting apoptosis-resistant cancer cells.
Similar content being viewed by others
Main
Autophagy is a highly regulated lysosomal proteolytic degradation process, which can engulf superfluous or damaged organelles and proteins through the formation of double-membraned autophagic vacuoles (autophagosomes).1 Upon formation of the autolysosomes through fusion of autophagosomes and lysosomes, all engulfed cellular components are degraded by acidic lysosomal hydrolases to synthesize new proteins.2 Although autophagy has a pivotal role in cellular housekeeping functions, it can be induced by stressors such as nutrients deprivation, oxidative stress, damaged proteins or organelles, pathogenic infection or hypoxia.2, 3 Furthermore, defects in autophagy are often linked to liver diseases, neurodegenerative diseases, inflammatory diseases, aging, cancers and metabolic syndromes.2, 4
Emerging evidence supports the role of autophagy in tumor suppression,5, 6 as increased levels of autophagy might lead to autophagic cell death (type II programmed cell death) in cancers.7 Autophagic cell death is accompanied by the formation of autophagosomes/autolysosomes without chromatin condensation.8 As cancer cells are frequently resistant to drug-mediated apoptosis after long-term chemotherapeutic treatments,9 the induction of autophagic cell death in apoptosis-defective or apoptosis-resistant tumor cells may provide an alternative therapeutic approach to tumor suppression.10 Recently, a number of clinically approved or experimental anti-tumor drugs have been shown to induce autophagy-related cell death in some malignant cells.11, 12, 13, 14 Therefore, the development and use of small-molecule autophagic inducers is an emerging approach for treatment of cancers, especially where they have become apoptosis resistant.15
We have previously identified a small-molecule, saikosaponin-d (Ssd), that exhibits anti-inflammatory effect via the suppression of NF-κB, NF-AT and AP-1 signaling.16 We recently showed that Ssd potentiated the anti-cancer potency of TNF-α through NF-κB inhibition.17 Ssd is one of the major triterpenoid saponins derived from Bupleurum falcatum L. (Umbelliferae), which is commonly prescribed for inflammatory and infectious diseases, and its active component Ssd was reported to have immuno-modulatory, anti-inflammatory, anti-bacterial, anti-viral and anti-cancer activities.18, 19
In this study, we present the first report that Ssd increased autophagic flux in HeLa and MCF-7 cancer cells, causing both apoptosis and autophagic cell death. Mechanistic studies revealed that Ssd-induced autophagy occurred by direct inhibition of sarcoplasmic/endoplasmic reticulum Ca2+ ATPase (SERCA), leading to the increase of intracellular calcium ion levels and activating the Ca2+/calmodulin-dependent kinase kinase-β (CaMKKβ)–AMP-activated protein kinase (AMPK)–mammalian target of rapamycin (mTOR) signaling cascade, endoplasmic reticulum (ER) stress and unfolded protein responses (UPR). The activation of these pathways ultimately leads to apoptotic and autophagic cell death in both cancer cells and, more importantly, apoptosis-resistant cancer cells.
Results
Ssd induces autophagy in HeLa and MCF-7 cells
Autophagy is directly involved in reducing the growth of tumor cells,20 thus chemical autophagy inducers could have the potential of inhibiting tumor cells.21 By using a panel of cancer cells, including HeLa, MCF-7, HepG2, PC3, H1299 and LLC-1, Ssd were shown to induce green fluorescent protein light-chain 3 (GFP-LC3) puncta formation (a marker of autophagy) in response to Ssd treatment (Supplementary Figures S1a and b). Among these cell types, HeLa and MCF-7 cells were chosen for more detailed mechanistic study. Results indicated that Ssd could time dependently induce autophagy as shown by the increased number of fluorescence GFP-LC3 puncta observed within the cells (Supplementary Figures S1a and b). To confirm the Ssd-induced autophagic activity, 5 mM 3-methyladenine (3-MA; an autophagic inhibitor) was added before Ssd treatment. Consistently, the addition of 3-MA decreased the Ssd-mediated autophagy as shown by the decreased percentage of cells with GFP-LC3 puncta (Figure 1a). In addition, immunocytochemical staining of endogenous LC3-II protein (a component of the autophagosome) also showed an increased level of puncta formation in cells treated with Ssd (Figure 1b).
Induction of autophagy by saikosaponin-d (Ssd). (a) GFP-LC3 detection of Ssd-mediated autophagy in HeLa and MCF-7 cells. HeLa and MCF-7 cells were transiently transfected with the GFP-LC3 plasmid for 24 h and then treated with dimethyl sulfoxide (DMSO; Ctrl), or 10 μM Ssd with or without 5 mM of autophagic inhibitor, 3-MA, for 4 h. Representative micrographs of cells that show GFP-LC3 localization. Bar chart represents the quantitation of autophagic cells. Percentages of autophagic cells demonstrated by the number of cells with GFP-LC3 dots signal (≥10 dots/cell) over the total number of GFP-positive cells in the same field. Arrows represent the cells with GFP-LC3 dots. More than 1000 GFP-positive cells were scored for each treatment. Data are the means of three independent experiments; error bars, S.D. **P<0.01; ***P<0.001 for Ssd-treated cells with and without 3-MA. (b) Direct detection of endogenous LC3-II protein by immunocytochemical analysis. HeLa cells were treated with DMSO (Ctrl) or 10 μM of Ssd for 4 h, then fixed in 4% paraformaldehyde and stained with anti-LC3-II antibodies (red). Images shown are representative of three independent experiments. Arrows indicate the cells with increased endogenous level of LC3-II in cytosolic region. All images are captured under × 60 magnification. (c) Ssd induces autophagic flux. HeLa and MCF-7 cells were treated with Ssd (10 μM) and lysosomal protease inhibitors (10 μg/mL), either alone or in combination, for the indicated time. Cell lysates were analyzed by western blot for LC3 conversion (LC3-I, 18 kDa; LC3-II, 16 kDa) and actin. (d) GFP-LC3 detection of Ssd with or without lysosomal protease inhibitors for 24 h in HeLa and MCF-7 cells. (e) Ssd stimulates the degradation of autophagic substrate p62. HeLa and MCF-7 cells were treated with Ssd (10 μM) for the indicated time. Cell lysates were analyzed using western blot for p62 and actin, respectively. Western blots images were quantified by densitometric analysis from three independent experiments and fold changes include the normalization to actin. Error bars, S.D. *P<0.05; **P<0.01 for cells treated with or without Ssd
Autophagosome accumulation may be caused by either an induction of autophagic flux via the autophagic pathway or an impairment of autophagosomes removal due to the blockage of fusion between autophagosome and lysosome.4 To differentiate between these two events, LC3-II formation in the presence of lysosomal protease inhibitors22 was assayed. Whereas Ssd or lysosomal protease inhibitors alone increased the level of LC3-II as expected, Ssd significantly increased the rate of LC3-II expression as well as its GFP-LC3 puncta formation in the presence of the inhibitors, compared with the addition of protease inhibitors alone (Figures 1c and d and Supplementary Figure S1c). These results therefore suggest that Ssd induces autophagic activity through increased autophagosome formation. p62 protein is another autophagy marker, which is selectively incorporated into autophagosomes and degraded upon autophagy induction.23 As shown in Figure 1e, decreased protein levels of p62, at least over the first 4 h of exposure, further suggest that Ssd induces autophagic flux.
Ssd-mediated autophagy is Vps34 and autophagy-related gene 7 (Atg7) dependent but Beclin1 independent
Beclin1 and Atg7 are both necessary for vesicle nucleation and elongation during autophagy.4 Gene deletion of beclin1 leads to a higher incidence of several cancers,24 whereas Atg7 knockout mice die during the neonatal starvation period.25 Immunoblots result demonstrated that the autophagy inducer, rapamycin, could increase beclin1 expression, whereas Ssd exhibited no significant effect on beclin1 expression in both HeLa and MCF-7 cells (Figure 2a). In addition, cell viability assays also showed that Ssd displayed similar cell viability curves in both control and beclin1 siRNA knockdown cells (Figure 2b). Knockdown of beclin1 also exhibited no reduction of Ssd-mediated GFP-LC3 puncta formation (Figure 2c). Given the inhibitory effect of 3-MA on Ssd-mediated autophagy induction by the PI3K inhibitor, 3-MA (Figure 1a), the role of Vps34, a beclin1-associated PI3kinase, was further studied. As shown in Supplementary Figure S2a, Ssd-induced autophagy was significantly reduced in Vps34 siRNA knockdown HeLa cells, suggesting that Vps34 is involved in Ssd-mediated autophagy but that its involvement is independent of beclin1.
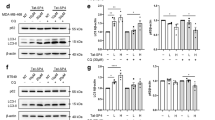
Role of Atg7 and beclin1 in Ssd-mediated autophagy. (a) Expression effect of beclin1 in response to Ssd treatment. HeLa and MCF-7 cells were treated with Ssd (10 μM) for the indicated time, whereas the cells treated with rapamycin (100 nM) for 24 h was positive control. Cell lysates were harvested and analyzed by western blot for beclin1 and actin, respectively. Error bars, S.D. *P<0.05; **P<0.01 for drug treatment compared with untreated control. (b) Ssd-mediated cell cytotoxicity is independent to beclin1 expression. HeLa and MCF-7 cells were transfected with control or beclin1 siRNA for 48 h, beclin1 knockdown efficiency was validated by western blot using anti-beclin1 antibody (upper panel). Both transfected cells were then incubated with Ssd for indicated range (0.19–100 μM) for 3 days. Cytotoxicity of Ssd on beclin1 knockdown HeLa and MCF-7 was measured by MTT (3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide) assay. (c) GFP-LC3 detection of Ssd-mediated autophagy in beclin1 knockdown cells. HeLa and MCF-7 cells were transfected with control siRNA or beclin1 siRNA together with GFP-LC3 plasmid for 48 h, treated with Ssd (10 μM) for 4 h and then fixed for fluorescence imaging and cell counting. (d) Expression effect of Atg7 in response to Ssd treatment. (e) Ssd-mediated cell cytotoxicity is dependent on Atg7 expression. (f) GFP-LC3 detection of Ssd-mediated autophagy in Atg7 knockdown cells. Bar chart represents the quantification of autophagic cells. Percentages of autophagic cells demonstrated by the number of cells with GFP-LC3 dots signal (≥10 dots/cell) over the total number of GFP-positive cells in the same field. More than 1000 GFP-positive cells were scored for each treatment. Western blot images were quantified by densitometric analysis from three independent experiments and fold changes include the normalization to actin. Error bars, S.D. P<0.001 for Ssd-treated HeLa and MCF-7 cells with or without Atg7 siRNA knockdown
Besides, Ssd and rapamycin treatment modalities do not increase the expression of Atg7 in cells (Figure 2d). However, cell viability assays showed that in Atg7 siRNA knockdown cells their sensitivity to Ssd were markedly reduced (mean LC50 for HeLa cells increased from 9.77 to 15.5 μM and for MCF-7 cells the mean LC50 increased from 9.1 to 13.6 μM) (Figure 2e). As knockdown of Atg7 significantly reduced Ssd-mediated GFP-LC3 puncta formation (Figure 2f), Atg7 is likely to have a role in Ssd-induced autophagy. Furthermore, Ssd also markedly induced GFP-LC3 puncta formation in Atg7+/+ wild-type mouse embryonic fibroblasts (MEFs) but not in Atg7−/−-deficient MEFs (Supplementary Figure S2b).
Ssd induces autophagy through activation of CaMKK-AMPK-mTOR kinase signaling cascade
Previous studies have shown that Ssd induces apoptosis through the release of reactive oxygen species,26 whereas autophagy can be induced via oxidative stress.4 However, addition of the anti-oxidant, N-acetyl-cysteine, was not able to reduce Ssd-mediated autophagy (Supplementary Figure S2c), suggesting that oxidative stress is not involved.
Nutrient deprivation can activate autophagy through the 5′-AMP-activated protein kinase (AMPK)-mTOR-dependent pathway.27 Both Ssd-treated HeLa and MCF-7 cells demonstrated the increase of AMPK phosphorylation in a time-dependent manner (Figure 3a), which was accompanied by a reduction in phosphorylated p70S6K, a downstream target of mTOR. Furthermore, a significant reduction in Ssd-induced GFP-LC3 puncta formation was also observed in cells pretreated with the AMPK inhibitor, compound C (Figure 3b and Supplementary Figure S2d).
Role of CaMKKβ-AMPK-mTOR signaling cascade in Ssd-mediated autophagy. (a) Ssd activates AMPK-mTOR signaling pathways. HeLa and MCF-7 cells were treated with Ssd (10 μM) for the indicated time and analyzed for p-AMPK, p-p70S6K, total p70S6K and actin, respectively. (b) AMPK inhibitor abrogates the Ssd-mediated autophagic effect in cancer cells. HeLa and MCF-7 cells were transiently transfected with the GFP-LC3 plasmid for 24 h and then treated with dimethyl sulfoxide (DMSO; Ctrl) or 10 μM Ssd with or without 5μM of AMPK inhibitor compound C (CC) for 4 h. The cells were then fixed for fluorescence imaging and cell counting. Bar chart represents the quantitation of autophagic cells (percentage of GFP-LC3 dots), error bars, S.D. *P<0.05. (c) CaMKKβ inhibitor abolishes the Ssd-mediated autophagic effect in cancer cells. GFP-LC3-transfected HeLa cells were treated with DMSO (Ctrl) or 10 μM Ssd with or without 25 μM of CaMKKβ inhibitor STO-609 for 4 h. The cells were then fixed for fluorescence imaging and cell counting. Bar chart represents the quantification of autophagic cells, error bars, S.D. **P<0.01. (d) Calcium chelator blocks the Ssd-induced autophagy and diminishes the Ssd-mediated cell cytotoxicity. GFP-LC3-transfected HeLa cells were treated with DMSO (Ctrl) or 10 μM Ssd in the presence of 5 mM of methyl pyruvate (MP) or 10 μM of calcium chelator BAPTA/AM (BM) for 4 h. The cells were then fixed for fluorescence imaging and cell counting. Bar chart represents the quantification of autophagic cells. The viability of HeLa cells treated with indicated concentrations of Ssd with or without 5 μM BM were measured by MTT (3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide) assay. Data are the means of three independent experiments; error bars, S.D. **P<0.01; ***P<0.001. (e) Schematic diagram to illustrate the Ssd-induced autophagy via CaMKK-AMPK-mTOR signaling cascade
Autophagy can also be induced by Ca2+-mobilizing agents by activating the CaMKKβ-AMPK-mTOR signaling cascade.28 To investigate the role of CaMKK in Ssd-induced autophagy, HeLa cells were treated with Ssd in the presence of the CaMKK inhibitor, STO-609.29 STO-609 significantly reduced the percentage of cells with GFP-LC3 puncta (Figure 3c and Supplementary Figure S2e), implicating possibly a role for Ca2+ signaling in this process.
AMPK can be activated by nutrient deprivation in cells by sensing changes in the AMP/ATP ratios.30 We therefore investigated whether the addition of Ssd can affect the mitochondrial membrane potential (ΔΨm), which controls the ATP level. Although decreased ΔΨm of HeLa cells was induced by Ssd treatment (Supplementary Figure S3), supplementation of the glycolytic intermediate (methyl pyruvate, MP) was unable to abolish Ssd-mediated GFP-LC3 puncta formation (Figure 3d), suggesting that Ssd-induced autophagy did not involve energy depletion but rather the decrease in ΔΨm is likely to be due to other effects of Ssd on the mitochondria. Addition of the intracellular Ca2+ chelator (BAPTA/AM, BM) to cells consistently abolished the formation of GFP-LC3 puncta (Figure 3d, upper panel) and markedly inhibited Ssd-induced cell death (Figure 3d, lower right panel). These results may therefore implicate a role for Ca2+ in depolarizing ΔΨm through mitochondrial Ca2+ overload.
Ssd inhibits the SERCA Ca2+ pump and increases cytosolic [Ca2+] to induce autophagy
To confirm whether Ssd can mediate autophagy by increasing cytosolic [Ca2+] levels, live-cell Ca2+ imaging was monitored and results showed that HeLa cells loaded with Fluo 3-AM displayed a marked increase in fluorescence intensity upon Ssd (10 μM) treatment (Figure 4a and Supplementary Video-1).
Ssd inhibits SERCA pump activity in turn to mobilize cytosolic calcium level. (a) Ssd mobilizes cytosolic calcium level in HeLa cells. HeLa cells seeded in 35 mm confocal disc were incubated with 5 μM of Fluo 3/AM in HBSS buffer at 37 °C for 30 min. Cells were then washed with HEPES buffer saline and incubated at 37 °C for another 10 min. Calcium signal was monitored by Applied Precision DeltaVision Elite in real-time mode for consecutive 5 min during the addition of 10 μM Ssd in HBSS buffer. Chart represents the mean intensity of fluorescence signal at 523 nm. (b) Computation docking of SERCA pump with saikosaponins. Upper panel, minimized energy pose of Ssd in SERCA pump (depicted in ribbon form and colored red). Stick model, Ssd; yellow, carbon; red, oxygen; middle panel, overlay of low-energy pose of Ssd (yellow, carbon; gray, hydrogen; red, oxygen) and thapsigargin (green) with the SERCA pump (depicted in ribbon form and colored red). Lower panel, minimized energy pose of Ssa in SERCA pump (depicted in ribbon form and colored red). Stick model, Ssa; yellow, carbon; red, oxygen. The binding pocket of the SERCA pump is represented as a translucent gray surface. (c) Inhibition of Ca2+ ATPase (SERCA) activity in skeletal muscle SR by Ssd. Experiments were measured at 25 °C (pH 7.2) using the coupled enzyme assay as described in Law et al.54 (d) Structure and activity relationship of saikosaponins on autophagy induction. GFP-LC3 transfected HeLa cells were treated with dimethyl sulfoxide (Ctrl), or 10 μM of Ssa, Ssc or Ssd, respectively for 4 h. The cells were then fixed for fluorescence imaging and cell counting. Bar chart represents the quanfication of autophagic cells. Data are the means of three independent experiments; error bars, S.D. **P<0.01; ***P<0.001
As the biological action of Ssd resembled thapsigargin, a potent inhibitor of the SERCA pump, which induces autophagy through perturbing calcium homeostasis,28 we postulated that Ssd might also function as a SERCA pump inhibitor. As such, computational virtual ligand docking studies were performed to evaluate the molecular interactions between Ssd and its structural analogues, Ssa and Ssc (Supplementary Figure S4a), with SERCA1A. Comparative analysis of the low-energy ligand conformations found the preferred site for Ssa and Ssd to be within the transmembrane domain (Figure 4b), close to the thapsigargin-binding site.31 Ssd was predicted to bind to SERCA through predominantly non-polar interactions with the hydrophobic residues lining the binding pocket, such as Phe256 and Val263, as well as Ile765 (Figure 4b, upper panel). Two hydrogen bonds were also predicted to occur between a hydroxyl group of Ssd with the backbone carbonyl group and backbone amide group of Met838 and Gly842, respectively, whereas a third hydrogen bond was made to the side-chain carboxylate moiety of Glu255. Comparison of the docking pose of Ssd with known SERCA inhibitor thapsigargin (binding score: −32) indicates that the two compounds were located in the space within the SERCA-binding pocket (Figure 4b, middle panel). Interestingly, however, thapsigargin was not predicted to make any hydrogen bonds to the SERCA protein. In the top-scoring binding pose of Ssa in the SERCA-binding site (Figure 4b, lower panel), the hydrogen bonds to Gly842 and Glu255 were absent, which might explain its weaker inhibition. The calculated binding score of −31.2 for Ssd and −28.9 for Ssa reflects notable potentially large difference in binding affinity between the two compounds and which may well explain their different potencies.
To ascertain whether the SERCA pump was suppressed by Ssd, we measured its effect using purified rabbit skeletal muscle sarcoplasmic reticulum (SR) membranes, which express the SERCA1A isoform.32 From previous studies with a range of other SERCA inhibitors, most inhibit similarly and independently of the SERCA isoform used;33, 34 the SERCA1A pump (from rabbit skeletal muscle SR) was inhibited by Ssd in a dose-dependent manner (Figure 4c), which was fitted to an allosteric dose versus effect equation, whereas Ssa and Ssc exhibited less and much lower inhibitory effects on SERCA1A activity, respectively (Supplementary Figure S4b). These findings coincided with the computation docking results of SERCA1A, which demonstrated that Ssd has a higher binding affinity and inhibitory effect on SERCA1A than Ssa, whereas Ssc displayed no inhibitory effect on SERCA1A activity. Concomitantly, GFP-LC3 puncta formation assay demonstrated that Ssd displayed an approximately∼twofolds of higher potency in autophagy induction than Ssa at 10 μM, whereas Ssc showed no autophagic effect at the same concentration (Figure 4d). The autophagic effect of Ss-a, -c and -d correlated well with SERCA inhibition and calculated binding scores on SERCA1A.
Ssd induces apoptosis and autophagic cell death simultaneously via UPR
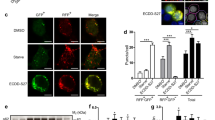
Although some anti-cancer agents can induce autophagy in various types of cancer,11 it remains controversial as to whether autophagy is in fact the driver of cell death or a pro-survival process in response to drug treatments. Consistent with our previous findings, Ssd not only induces autophagy in HepG2 cells (Figure 5a and Supplementary Figure S1a) but could also dose-dependently induce apoptosis in these cells (Figure 5b). Additional experiments were undertaken to assess whether some of the other classic hallmarks of autophagic cell death (i.e., independent of caspases and partial chromatin condensation, without nuclear fragmentation or DNA laddering11) were present with Ssd treatment. Figures 5c and d shows that HeLa cells treated with Ssd exhibited time-dependent membrane blebbing and partial chromatin condensation (but did not show any DNA laddering or nuclear fragmentation). Similar characteristics were also observed in H1299 cancer cells (Supplementary Figure S5a). Furthermore, addition of the pan-caspase inhibitor, z-VAD-FMK, did not reduce the Ssd-mediated cell death (Supplementary Figure S5b). As there are links between the autophagic and necrotic pathways,35 HeLa, MCF-7 and Atg7−/−-deficient cells were examined with Ssd in the presence of necrosis inhibitor, necrostatin. As shown in Supplementary Figure S5c, addition of 10 μM necrostatin failed to rescue the cell death from Ssd treatment, suggesting that necrosis is not involved. In addition, Atg7−/−-deficient cells that are resistant to autophagy36 were also less sensitive to Ssd-mediated cell death (mean LC50, 7.76 μM) compared with the Atg7+/+ wild-type cells (mean LC50, 4.79 μM) (Figure 5e). This was also confirmed by Annexin V staining, where Ssd-induced apoptotic cell death was significantly suppressed in Atg7 siRNA knockdowns of HeLa and MCF-7 cells (Figure 5f and Supplementary Figure S5d).
Ssd induces UPR with induction of apoptosis and autophagic cell death simultaneously. (a) Ssd induces autophagy in HepG2 cells. (b) Ssd induces apoptosis detected by Annexin V staining. HepG2 cells were incubated with medium control or 7.5–15 μM of Ssd for 24 h. The cells were then stained with Annexin V-FITC (fluorescein isothiocyanate) and propidium iodide (PI), followed by flow cytometry analysis. *P<0.05, **P<0.01 and ***P<0.001, compared with medium control treatment. (c) Ssd induces cell death with autophagic cell death morphologies. HeLa cells were incubated with dimethyl sulfoxide (Ctrl) or 10 μM Ssd for 4 h. The cells were then fixed and stained with DAPI. Left panel represents the bright-field images and right panel represents the DAPI fluorescence images. All images were captured under × 60 magnification. Arrows, autophagic cell death morphologies: membrane blebbing, chromatin condensation and no nuclear fragmentation. (d) Ssd induces autophagic cell death morphology in a time-dependent manner. Percentages of autophagic cell death demonstrated by the number of cells with cell membrane blebbing, partial chromatin condensation and no nuclear fragmentation over the total number of cells in the same field. More than 1000 cells were scored for each treatment. Results shown are representative of three independent experiments. (e) Cytotoxicity of Ssd on wild-type and ATG7−/−-deficient MEFs as measured by the MTT (3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide) assay. (f) Annexin V detection of Ssd-mediated cell cytotoxicity in Atg7 siRNA knockdown HeLa and MCF-7 cells. HeLa and MCF-7 cells were knocked down by control siRNA or Atg7 siRNA before 10 μM of Ssd treatment for 24 h. The cells were then stained with Annexin V-FITC and PI, followed by flow cytometry analysis. *P<0.05; **P<0.01 and ***P<0.001. (g) Effect of calcium chelator, BAPTA/AM, on Ssd-mediated cell cytotoxicity on HeLa cells. (h) Live-cell imaging of Ssd-mediated autophagic cell death. HeLa cells were incubated in imaging chamber at 37 °C supplied with 5% of CO2. Live-cell imaging was started to capture the GFP and bright field images after addition of 10 μM of Ssd for 4 h with 10-min time intervals. Arrows (yellow), autophagy induction with GFP-LC3 accumulation (green); arrows (red), autophagic cell death. (i) Ssd induces UPR in cancer cells. HeLa and MCF-7 cells were treated with Ssd (10 μM) for the indicated time, whereas rapamycin (100 nM) and thapsigargin (1 μM) were used as positive control. Cell lysates were harvested and analyzed by western blot for ER stress markers. (j) HeLa cells were treated with Ssd (10 μM) or thapsigargin (1 μM) for the indicated time. Reverse transcription-PCR was performed to analyze Xbp-1 mRNA splicing. (k) Effect of ER stress inhibitor, 4-phenyl-butyric acid, on Ssd-mediated cell cytotoxicity. Results shown are the means±S.E.Ms. of three independent experiments
BAPTA/AM, which can significantly abolish Ssd-mediated autophagy (see Figure 3d), was also able to reduce Ssd-mediated cell death in HeLa cells. The mean LC50 increased from 10.4 to 25.1 μM (Figure 5g). Furthermore, live-cell tracking visualized the autophagic progression as well as autophagic cell death upon Ssd treatment. GFP-LC3-transfected HeLa cells demonstrated fluorescence puncta at 10 min and reached a maximum number of GFP-puncta at 3 h and 30 min (yellow arrow) upon Ssd treatment; the cells were then killed by autophagic cell death (red arrow) (Figure 5h and Supplementary Video-2).
Disturbance of calcium homeostasis is known to cause ER stress and UPR activation, thereby inducing apoptosis and autophagy.37 Figure 5i shows that thapsigargin, (a known autophagy inducer, ER stress inducer and Ca2+ mobilizer) and Ssd both caused PERK (pancreatic ER kinase) and eIF2α phosphorylation in both HeLa and MCF-7 cells. This was accompanied by an increase in ER molecular chaperone BiP/GRP78 and ATF4 expression, as well as nuclear translocation of ATF6 (activating transcription factor 6). However, thapsigargin, but not Ssd, induced the splicing of Xbp-1 mRNA (Figure 5j), whereas only Ssd induced IRE1 (inositol-requiring transmembrane kinase/endonuclease 1)-mediated JNK and caspase 12 activation (Figure 5i), suggesting that Ssd might specifically activate the IRE1-JNK-mediated apoptotic pathway. In contrast, addition of 4-phenyl-butyric acid, a known ER stress inhibitor,38 promoted cell survival through suppressing Ssd-induced UPR activation in HeLa (mean LC50, from 8.22 to 35.8 μM) and MCF-7 cells (mean LC50, from 7.76 to 19.8 μM) (Figure 5k), implicating that Ssd-mediated cell death is caused, at least in part, through ER stress and UPR activation.
Ssd induces autophagic cell death in apoptosis-resistant cells
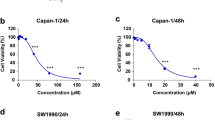
Cancer cells are frequently resistant to apoptosis.9 Therefore, the induction of autophagic cell death in apoptosis-defective or apoptosis-resistant cancer cells might also be an effective therapeutic approach.10 In this connection, we used a panel of apoptosis-defective or apoptosis-resistant cells such as caspase-3/7/8-deficient MEFs and Bax-Bak double-knockout MEFs to examine the cytotoxic effect of Ssd. As shown in Figures 6a–c, Ssd displayed similar cytotoxicity profiles for both wild-type and Caspase-3/-7/-8-deficient, or Bax-Bak DKO MEFs, implicating that Ssd can circumvent the apoptosis-resistant phenotype of cells derived from various apoptotic gene deficiencies. In contrast, the apoptosis inducer, staurosporine, was found to be much less sensitive in Bax-Bak-deficient cells (Figure 6c, right panel). In addition, blockage of Ssd-mediated autophagy by 3-MA or BAPTA/AM significantly increased the cell viability in these apoptosis-resistant Bax-Bak-/- cells (Figure 6d). However, PI-annexin V staining and MTT cytotoxicity assays demonstrated that addition of necrosis inhibitor, necrostatin, failed to abolish the Ssd-mediated cell cytotoxicity in Bax-Bak−/− cells (Figure 6e), suggesting that Ssd is solely capable of inducing autophagic cell death in these apoptosis-resistant cells. Finally, the progression of Ssd-induced autophagy and autophagic cell death in apoptosis-resistant Bax-Bak−/− cells was further visualized by live-cell imaging (Supplementary Video-3).
Ssd induces autophagic cell death in apoptosis-resistant cells. (a) Cytotoxicity of Ssd on caspase 3/7 wild-type (wt) and deficient MEFs. (b) Cytotoxicity of Ssd on caspase 8 wild-type and deficient MEFs. (c) Cytotoxicity of Ssd and staurosporine on Bax-Bak wild-type and double-knockout (DKO) MEFs. (d) Blockage of autophagy diminishes the Ssd-mediated cell death in apoptosis-resistant cells. Bax-Bak DKO MEFs were treated with dimethyl sulfoxide (Crtl) or Ssd (10 μM) with or without autophagy inhibitor, 3-MA (5 mM), or calcium chelator, BAPTA/AM (10 μM), for 24 h and then subjected to image capture. Bright-field images shown are representative of three independent experiments. All images were captured under × 10 magnification, micrographs from red rectangle represent the magnified cropped images from the particular original images. (e) PI-annexin V and MTT (3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide) analysis of the effect of necrosis inhibitor, necrostatin (Nec-1), on Ssd-mediated cell cytotoxicity in Bax-Bak DKO MEFs. Results shown are the Means±S.E.Ms. of three independent experiments. (f) Schematic diagram to illustrate the proposed molecular target and mechanisms of Ssd in autophagy induction and its subsequent effects in apoptosis and autophagic cell death
Discussion
Autophagy has been reported as a tumor suppressor mechanism and that a number of anti-cancer treatments in clinical use today may induce autophagy in tumor cells.15 Tetrahydrocannabinol, the active component of marijuana, induces ER stress and subsequent Akt/mTOR inhibition leading to autophagy in glioma cells.39 Gemcitabine, which induces autophagy-dependent cell death, is now regularly used to treat pancreatic cancer40 and the small molecule STF-62247 selectively induces autophagic cell death in renal cell carcinoma.41 Numerous clinical trials evaluating the effectiveness of mTOR inhibition alone or in combination with other therapies have been conducted by using everolimus and temsirolimus;15 therefore determining the role of autophagy during cancer therapy could be crucial in increasing the survival rate of cancer patients.
We have shown that Ssd induces calcium mobilization, leading to activation of autophagy through the CaMKKβ-AMPK-mTOR pathway (Figure 3e), and subsequently induces ER stress and UPR activation, leading to apoptosis and autophagic cell death (Figure 6f). The simultaneous induction of these programmed cell death supports the idea that both apoptosis and autophagic cell death can cross-talk and be triggered by common upstream signals.42
With respect to the slight difference between the stereochemistry of Ssa (β-OH group) and Ssd (α-OH group) at the same position of their chemical structure (Supplementary Figure S4a), results of computational docking of SERCA revealed that the α-OH group in Ssd may contribute to the overall bioactivity by inducing an optimal orientation of the side chain, leading to the hydrogen bonding with residues of Met838, Gly842 and Glu255 in SERCA, which renders the terminus of Ssd more compact to fit into the binding pocket compared with Ssa. Coincidentally, the fact that the antibodies that are specific for Ssa showed only slight cross-reactivity with Ssd (0.3%)43 further suggests that the structural stereochemistry of the 16-hydroxyl group could produce a great variation in their protein-binding affinity and therefore biological activities.
SERCA has been identified as a novel therapeutic target for cancers.44, 45 The natural product thapsigargin and its derivatives are the most commonly used SERCA inhibitors due to their high potencies.46 Thapsigargin inhibits the uptake of calcium into the sarcoplasmic and endoplasmic reticula, leading to the increase in cytosolic calcium levels. The disturbance of calcium homeostasis by thapsigargin disorganizes protein folding and induces ER stress and UPR.37 Owing to the SERCA pump’s critical role in normal cellular metabolism, agents like thapsigargin, which directly inhibits SERCA function, would likely produce significant toxicity to normal cells. Nevertheless, thapsigargin-mediated cytotoxicity can be mitigated by coupling to a targeting peptide to produce an inactive pro-drug that is only activated by prostate-specific antigen in prostate cancer.45 Therefore, Ssd can be further exploited by using pro-drug strategy similar to that of thapsigargin. Even though thapsigargin is a potent inhibitor of the SERCA, it is a rather expensive and a nonspecific necrotic agent with low yield total synthesis process.47, 48 Furthermore, thapsigargin was recently shown to induce permanent mitochondrial damage by Ca2+ overload, thereby forcing the multidrug-resistant Bax-Bak-deficient cells into necrosis.47 Although Ssd is newly identified as SERCA inhibitor, unlike thapsigargin, the necrosis inhibitor, necrostatin, failed to rescue Ssd-mediated cell death in Bax-Bak-deficient MEFs, HeLa, MCF-7, as well as autophagic-defective Atg7−/− MEFs, suggesting that necrosis is not involved. Therefore, discovery of new SERCA inhibitors like Ssd, which is able to induce apoptosis and autophagic cell death is highly desirable.
The UPR is mediated by three distinct ER-localized transmembrane proteins: PERK, IRE1 and ATF6. When cells encounter ER stress or misfolded protein accumulation, an ER-resident chaperone, GRP78, dissociates from PERK, IRE1 and ATF6, leading to autophosphorylation of PERK, IRE1 and mobilization of ATF6 for activation.37 Therefore, upregulation of GRP78, phosphorylation of PERK and eIF2α that leads to ATF4 overexpression and the nuclear translocation of ATF6 in Ssd-treated cells supported that Ssd could induce ER stress and UPR in cancer cells. Although PERK is responsible for CCAAT/enhancer-binding protein homologous protein (CHOP)-mediated apoptosis,49 activation of CHOP is only found in thapsigargin-treated cells but not in Ssd. Concomitantly, recent studies further support our findings that CHOP is not crucial for ER stress-induced apoptosis,50 suggesting that Ssd may activate ER stress-mediated apoptosis in a CHOP-independent manner.
Drug resistance is the major cause of therapeutic failure in cancer. Most, if not all, radiation- and chemotherapy-resistant cancers have apoptotic defects. Besides, the mitochondria/cytochrome c pathway of apoptosis is frequently deregulated in human cancer.11 For instance, Bax/Bak expression is severely attenuated in many malignancies,51 MEFs from double-knockout Bax–/– Bak–/– mice are resistant to a range of apoptosis inducers;42 whereas caspases-3 and -7 are critical mediators of mitochondrial events of apoptosis,52 and caspase-3, -8 and -9 are found to have a critical role in anti-cancer drug-induced apoptosis, in apoptosis-resistance and anti-cancer drug resistance.53 Our work has shown that even when caspases-3/-7/-8 and Bax/Bak genes were deleted, Ssd could still trigger caspase-independent cell death via autophagy, suggesting the potential therapeutic activity of Ssd in apoptosis-resistant cancers.
Materials and Methods
Chemicals, plasmids, small interfering RNAs and antibodies
All compounds were purchased from Sigma (St. Louis, MO, USA) unless otherwise stated. Thapsigargin, compound C, BAPTA/AM, E64D, pepstatin A, staurosporine and STO-609 were obtained from Calbiochem (San Diego, CA, USA). Saikosaponin-a/-c/-d (Ssa, Ssc and Ssd) (>98% purity, HPLC) were purchased from the China Chengdu Biotechnology Company Ltd (Chengdu, China). Antibodies against LC3B, Atg7, p70S6 kinase, phospho-p70S6 kinase (Thr389), phospho-AMPKα (Thr172), eIF2α, phospho-eIF2α (Ser51), PERK, beclin1, phosphor-JNK and JNK were purchased from Cell Signaling Technologies Inc. (Beverly, MA, USA). CHOP, p62, GRP78 and ATF4 antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). β-Actin antibodies were from Sigma. Anti-phospho-PERK (Thr980) antibodies were from BioLegend (San Diego, CA, USA). ATF6 and Caspase-12 antibodies were purchased from Novus (Littleton, CO, USA). pEGFP-LC3 reporter plasmid was a gift from Professor Tamotsu Yoshimori (Osaka University, Osaka, Japan). siRNAs targeting beclin1 or non-targeting control and necrostatin were obtained from Santa Cruz Biotechnology. siRNAs targeting Atg7 and Vps34 were obtained from Qiagen (Hilden, Germany).
Cell culture
All cells were obtained from the American Type Culture Collection (Rockville, MD, USA) unless otherwise specified. Immortalized wild-type and Atg7-deficient MEFs were kindly provided by Professor Masaaki Komatsu (Juntendo University, School of Medicine, Japan). Immortalized wild-type and Caspase 3/7-deficient MEFs were kindly provided by Professor Richard A. Flavell (Yale University School of Medicine, , USA). Immortalized wild-type and Caspase 8-deficient MEFs were kindly provided by Professor Kazuhiro Sakamaki (Kyoto University, Graduate School of Biostudies, Japan). Immortalized wild-type and Bax-Bak double-knockout MEFs were kindly provided by Professor Shigeomi Shimizu (Tokyo Medical and Dental University, Medical Research Institute, Japan). All medium were supplemented with 10% fetal bovine serum and the antibiotics penicillin (50 U/ml) and streptomycin (50 μg/ml; Invitrogen, Paisley, Scotland, UK). All cell cultures were incubated at 37 °C in a 5% humidified CO2 incubator.
Quantification of GFP-LC3 puncta
GFP-LC3 puncta was quantified as described previously.54 Cells were fixed with 4% paraformaldehyde (Sigma), permeabilized with methanol and nuclei were stained with DAPI (4,6-diamidino-2-phenylindole). Coverslips were mounted onto microscope slides with FluorSave Reagent (Calbiochem), and localization of GFP-LC3 examined under the Nikon ECLIPSE 80i microscope (Melville, NY, USA). Images were captured with CCD digital camera Spot RT3 (Diagnostic Instruments, Inc., Melville, NY, USA). Guidelines were followed to monitor autophagy;55 the percentage of cells with punctuate GFP-LC3 fluorescence was calculated by counting the number of the cells showing the punctuate pattern of GFP-LC3 (≥10 dots/cell) in GFP-positive cells over the total number of GFP-positive cells in the same field. A minimum of 1000 cells from randomly selected fields were scored per condition per experiment.
Cytotoxicity assays and apoptosis detection
Cell viability was measured using the MTT assay as described in Wong et al.16 The percentage of cell viability was calculated using the following formula: Cell viability (%)=Cells numbertreated/ Cells numberDMSO control × 100. Apoptosis was detected by Annexin V staining kit (BD Biosciences, San Diego, CA, USA). Data were obtained from three independent experiments.
Analysis of apoptosis and mitochondrial membrane potential
HepG2 cells treated with indicated concentrations of Ssd were analyzed by multiparametric flow cytometry using FITC-Annexin V and Propidium iodide staining (BD Biosciences) according to the manufacturer’s instructions. To analyze ΔΨm, HeLa cells were first stained with Rho123 (Invitrogen) at 5 μM for 20 min in the dark at room temperature. Subsequently, cells were washed with PBS three times and then treated with 10 μM Ssd from 10 to 60 min. Fluorescence images were taken by Nikon fluorescence microscope (Melville, NY, USA). Flow cytometry was carried out using a FACSCalibur flow cytometer (BD Biosciences). Data acquisition and analysis were performed with CellQuest (BD Biosciences).
RT-PCR analysis of Xbp-1 mRNA splicing
Total RNA was extracted using TRIZOL (Invitrogen). First-strand cDNA was synthesized using the High Capacity RNA-to-cDNA Master Mix (Applied Biosystems, Carlsbad, CA, USA). To detect human spliced and unspliced Xbp-1 mRNA, PCR was performed using primers 5′-CTGGAACAGCAAGTGGTAGA-3′ and 5′-CTGGGTCCTTCTGGGTAGAC-3′ as described.56 Spliced (398 bp) and unspliced (424 bp) Xbp-1 fragments were separated by electrophoresis in a 2% agarose gel with DNA bands stained with ethidium bromide and photographed.
Virtual ligand docking
Molecular docking was done using the ICM-Pro 3.6-1 program (Molsoft L.L.C., San Diego, CA, USA;57). The protein crystal structure of rabbit SERCA1A with bound thapsigargin (PDB code: 2AGV) was downloaded from the protein data bank as a basis for modeling. The complexes were evaluated with a full-atom ICM ligand-binding score58 that has been previously derived from a multi-receptor screening benchmark as a compromise between approximated Gibbs free energy of binding and numerical errors. As a reference, molecular docking of the known SERCA pump inhibitor, thapsigargin, showed a score of −32.
Measurement of SERCA activity
Purified Ca2+ ATPase (SERCA1A) was prepared from female rabbit hind leg muscle.59 ATPase activity was determined using the enzyme-coupled method using pyruvate kinase and lactate dehydrogenase as previously described in Michelangeli et al.60 All SERCA inhibition data were fitted to the allosteric dose versus effect equation using Fig P (Biosoft, Cambridge, UK):
Activity=minimum activity+(maximum activity−minimum activity)/(1+([I]/IC50)P).
Calcium imaging
In all, 2 × 106 HeLa cells were cultured in 35 mm confocal disc in a CO2 incubator set at 37 °C for 24 h and 5 mM of Fluo 3/AM/DMSO stock solution was diluted to 5 μM working solution using Hank’s Balanced Salt Solution (HBSS) and then added to cells at 37 °C for 30 min. HeLa cells were then washed three times with HEPES buffer saline and incubated at 37 °C in an imaging chamber for another 10 min. Changes in cytosolic [Ca2+] levels were monitored by following changes in fluo-3 fluorescence upon addition of 10 μM Ssd in HBSS buffer, using the real-time mode for 5 min by epifluorescence microscopy (Applied Precision DeltaVision Elite, Applied Precision Inc., GE Healthcare Company, Issaquah, WA, USA). Data Inspection Program provided by the DeltaVision software was used to measure the intensity of the fluo-3 fluorescence, and the mean fluorescence intensity was monitored at 523 nm and plotted against time (s).
Live-cell imaging
Autophagy induction was visualized in HeLa cells or Bax-Bak−/− MEFs, which were transiently transfected with GFP-LC3, and then placed on the microscope stage covered with a 37 °C chamber in which a humidified premixed gas comprising 5% CO2 and 95% air was infused. After treatment with Ssd (10 μM), the cells were observed using 60X Olympus PlanApoN 1.42 oil objective (Applied Precision Inc., GE Healthcare Company), and the fluorescence monitored at 512 nm. Both DIC and fluorescent images were acquired at 5-min intervals using high magnification wide-field epifluorescence microscopy. Images were captured as serial Z-sections of 1.0-μm interval by a Photometrics CoolSNAP HQ2 CCD camera (Applied Precision Inc., GE Healthcare Company) on the Olympus IX71-Applied Precision DeltaVision restoration microscope (Applied Precision Inc., GE Healthcare Company), and the epifluorescence images were numerically deconvolved using DeltaVision algorithms (Applied Precision Inc., GE Healthcare Company).
Statistical analysis
The results were expressed as Means±S.Ds. as indicated. The differences were considered statistically significant when the P-value was <0.05. A Student’s t-test or one-way ANOVA analysis was used for comparison among the different groups.
Abbreviations
- Ssd:
-
saikosaponin-d
- LC3:
-
light-chain 3
- GFP:
-
green fluorescent protein
- HBSS:
-
Hank’s Balanced Salt Solution
- 3-MA:
-
3-methyladenine
- SERCA:
-
sarcoplasmic/endoplasmic reticulum Ca2+ ATPase pump
- CaMKKβ:
-
Ca2+/calmodulin-dependent kinase kinase-β
- AMPK:
-
AMP-activated protein kinase
- mTOR:
-
mammalian target of rapamycin
- CHOP:
-
CCAAT/enhancer-binding protein homologous protein
- ER:
-
endoplasmic reticulum
- SR:
-
sarcoplasmic reticulum
- UPR:
-
unfolded protein responses
- MEF:
-
mouse embryonic fibroblast
- Atg:
-
autophagy-related gene
- PERK:
-
pancreatic ER kinase
- IRE1:
-
inositol-requiring transmembrane kinase/endonuclease 1
- ATF6:
-
activating transcription factor 6
References
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6: 463–477.
Rabinowitz JD, White E . Autophagy and metabolism. Science 330: 1344–1348.
Mizushima N . Autophagy: process and function. Genes Dev 2007; 21: 2861–2873.
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132: 27–42.
Levine B . Cell biology: autophagy and cancer. Nature 2007; 446: 745–747.
Mathew R, Karantza-Wadsworth V, White E . Role of autophagy in cancer. Nat Rev Cancer 2007; 7: 961–967.
Tsujimoto Y, Shimizu S . Another way to die: autophagic programmed cell death. Cell Death Differ 2005; 12 (Suppl 2): 1528–1534.
Kroemer G, Levine B . Autophagic cell death: the story of a misnomer. Nat Rev Mol Cell Biol 2008; 9: 1004–1010.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Alva AS, Gultekin SH, Baehrecke EH . Autophagy in human tumors: cell survival or death? Cell Death Differ 2004; 11: 1046–1048.
Kondo Y, Kanzawa T, Sawaya R, Kondo S . The role of autophagy in cancer development and response to therapy. Nat Rev Cancer 2005; 5: 726–734.
Hoyer-Hansen M, Bastholm L, Mathiasen IS, Elling F, Jaattela M . Vitamin D analog EB1089 triggers dramatic lysosomal changes and Beclin 1-mediated autophagic cell death. Cell Death Differ 2005; 12: 1297–1309.
Chang CP, Yang MC, Liu HS, Lin YS, Lei HY . Concanavalin A induces autophagy in hepatoma cells and has a therapeutic effect in a murine in situ hepatoma model. Hepatology 2007; 45: 286–296.
Opipari AW Jr., Tan L, Boitano AE, Sorenson DR, Aurora A, Liu JR . Resveratrol-induced autophagocytosis in ovarian cancer cells. Cancer Res 2004; 64: 696–703.
Levy JM, Thorburn A . Targeting autophagy during cancer therapy to improve clinical outcomes. Pharmacol Ther 2011; 131: 130–141.
Wong VK, Zhou H, Cheung SS, Li T, Liu L . Mechanistic study of saikosaponin-d (Ssd) on suppression of murine T lymphocyte activation. J Cell Biochem 2009; 107: 303–315.
Wong VKWZM, Zhou H, Lam KYC, Chan PL, CKM Law, PYK Yue et al. Saikosaponin-d enhances the anticancer potency of TNF via overcoming its undesirable response of activating NF-kappa B signalling in cancer cells. Evid Based Complement Altern Med 2013 2013: 14.
Tundis R, Bonesi M, Deguin B, Loizzo MR, Menichini F, Conforti F et al. Cytotoxic activity and inhibitory effect on nitric oxide production of triterpene saponins from the roots of Physospermum verticillatum (Waldst & Kit) (Apiaceae). Bioorg Med Chem 2009; 17: 4542–4547.
Hsu YL, Kuo PL, Chiang LC, Lin CC . Involvement of p53, nuclear factor kappaB and Fas/Fas ligand in induction of apoptosis and cell cycle arrest by saikosaponin d in human hepatoma cell lines. Cancer Lett 2004; 213: 213–221.
Dalby KN, Tekedereli I, Lopez-Berestein G, Ozpolat B . Targeting the prodeath and prosurvival functions of autophagy as novel therapeutic strategies in cancer. Autophagy 2010; 6: 322–329.
Turcotte S, Giaccia AJ . Targeting cancer cells through autophagy for anticancer therapy. Curr Opin Cell Biol 2010; 22: 246–251.
Mizushima N, Yoshimori T . How to interpret LC3 immunoblotting. Autophagy 2007; 3: 542–545.
Bjorkoy G, Lamark T, Pankiv S, Overvatn A, Brech A, Johansen T . Monitoring autophagic degradation of p62/SQSTM1. Methods Enzymol 2009; 452: 181–197.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol 2005; 169: 425–434.
Wang Q, Zheng XL, Yang L, Shi F, Gao LB, Zhong YJ et al. Reactive oxygen species-mediated apoptosis contributes to chemosensitization effect of saikosaponins on cisplatin-induced cytotoxicity in cancer cells. J Exp Clin Cancer Res 2010; 29: 159.
Mizushima N, Klionsky DJ . Protein turnover via autophagy: implications for metabolism. Annu Rev Nutr 2007; 27: 19–40.
Hoyer-Hansen M, Bastholm L, Szyniarowski P, Campanella M, Szabadkai G, Farkas T et al. Control of macroautophagy by calcium, calmodulin-dependent kinase kinase-beta, and Bcl-2. Mol Cell 2007; 25: 193–205.
Tokumitsu H, Inuzuka H, Ishikawa Y, Ikeda M, Saji I, Kobayashi R . STO-609, a specific inhibitor of the Ca(2+)/calmodulin-dependent protein kinase kinase. J Biol Chem 2002; 277: 15813–15818.
Liang J, Shao SH, Xu ZX, Hennessy B, Ding Z, Larrea M et al. The energy sensing LKB1-AMPK pathway regulates p27(kip1) phosphorylation mediating the decision to enter autophagy or apoptosis. Nat Cell Biol 2007; 9: 218–224.
Hoyer-Hansen M, Jaattela M . Connecting endoplasmic reticulum stress to autophagy by unfolded protein response and calcium. Cell Death Differ 2007; 14: 1576–1582.
Wu KD, Lee WS, Wey J, Bungard D, Lytton J . Localization and quantification of endoplasmic reticulum Ca(2+)-ATPase isoform transcripts. Am J Physiol 1995; 269 (3 Pt 1): C775–C784.
Michelangeli F, East JM . A diversity of SERCA Ca2+ pump inhibitors. Biochem Soc Trans 2011; 39: 789–797.
East JM, Michelangeli F . Recent advances in membrane biochemistry. Biochem Soc Trans 2011; 39: 703–706.
Jain MV, Paczulla AM, Klonisch T, Dimgba FN, Rao SB, Roberg K et al. Interconnections between apoptotic, autophagic and necrotic pathways: implications for cancer therapy development. J Cell Mol Med 2013; 17: 12–29.
Li J, Ni M, Lee B, Barron E, Hinton DR, Lee AS . The unfolded protein response regulator GRP78/BiP is required for endoplasmic reticulum integrity and stress-induced autophagy in mammalian cells. Cell Death Differ 2008; 15: 1460–1471.
Todd DJ, Lee AH, Glimcher LH . The endoplasmic reticulum stress response in immunity and autoimmunity. Nat Rev Immunol 2008; 8: 663–674.
Ozcan U, Yilmaz E, Ozcan L, Furuhashi M, Vaillancourt E, Smith RO et al. Chemical chaperones reduce ER stress and restore glucose homeostasis in a mouse model of type 2 diabetes. Science 2006; 313: 1137–1140.
Salazar M, Carracedo A, Salanueva IJ, Hernandez-Tiedra S, Lorente M, Egia A et al. Cannabinoid action induces autophagy-mediated cell death through stimulation of ER stress in human glioma cells. J Clin Invest 2009; 119: 1359–1372.
Pardo R, Lo Re A, Archange C, Ropolo A, Papademetrio DL, Gonzalez CD et al. Gemcitabine induces the VMP1-mediated autophagy pathway to promote apoptotic death in human pancreatic cancer cells. Pancreatology 2010; 10: 19–26.
Turcotte S, Chan DA, Sutphin PD, Hay MP, Denny WA, Giaccia AJ . A molecule targeting VHL-deficient renal cell carcinoma that induces autophagy. Cancer Cell 2008; 14: 90–102.
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G . Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat Rev Mol Cell Biol 2007; 8: 741–752.
Jung DW, Shibuya M, Ebizuka Y, Yoshimatsu K, Shimomura K, Sung CK . ELISA for the determination of saikosaponin a, an active component of Bupleuri Radix. Chem Pharm Bull (Tokyo) 1998; 46: 1140–1143.
He Q, Lee DI, Rong R, Yu M, Luo X, Klein M et al. Endoplasmic reticulum calcium pool depletion-induced apoptosis is coupled with activation of the death receptor 5 pathway. Oncogene 2002; 21: 2623–2633.
Denmeade SR, Isaacs JT . The SERCA pump as a therapeutic target: making a "smart bomb" for prostate cancer. Cancer Biol Ther 2005; 4: 14–22.
Elam C, Lape M, Deye J, Zultowsky J, Stanton DT, Paula S . Discovery of novel SERCA inhibitors by virtual screening of a large compound library. Eur J Med Chem 2011; 46: 1512–1523.
Janssen K, Horn S, Niemann MT, Daniel PT, Schulze-Osthoff K, Fischer U . Inhibition of the ER Ca2+ pump forces multidrug-resistant cells deficient in Bak and Bax into necrosis. J Cell Sci 2009; 122 (Pt 24): 4481–4491.
Ley SV, Antonello A, Balskus EP, Booth DT, Christensen SB, Cleator E et al. Synthesis of the thapsigargins. Proc Natl Acad Sci USA 2004; 101: 12073–12078.
Zinszner H, Kuroda M, Wang X, Batchvarova N, Lightfoot RT, Remotti H et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev 1998; 12: 982–995.
Masciarelli S, Fra AM, Pengo N, Bertolotti M, Cenci S, Fagioli C et al. CHOP-independent apoptosis and pathway-selective induction of the UPR in developing plasma cells. Mol Immunol 2010; 47: 1356–1365.
Yuan J . Divergence from a dedicated cellular suicide mechanism: exploring the evolution of cell death. Mol Cell 2006; 23: 1–12.
Lakhani SA, Masud A, Kuida K, Porter GA Jr., Booth CJ, Mehal WZ et al. Caspases 3 and 7: key mediators of mitochondrial events of apoptosis. Science 2006; 311: 847–851.
Friesen C, Uhl M, Pannicke U, Schwarz K, Miltner E, Debatin KM . DNA-ligase IV and DNA-protein kinase play a critical role in deficient caspases activation in apoptosis-resistant cancer cells by using doxorubicin. Mol Biol Cell 2008; 19: 3283–3289.
Law BY, Wang M, Ma DL, Al-Mousa F, Michelangeli F, Cheng SH et al. Alisol B, a novel inhibitor of the sarcoplasmic/endoplasmic reticulum Ca(2+) ATPase pump, induces autophagy, endoplasmic reticulum stress, and apoptosis. Mol Cancer Ther 2010; 9: 718–730.
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012; 8: 445–544.
Shang J, Lehrman MA . Discordance of UPR signaling by ATF6 and Ire1p-XBP1 with levels of target transcripts. Biochem Biophys Res Commun 2004; 317: 390–396.
Totrov M, Abagyan R . Flexible protein-ligand docking by global energy optimization in internal coordinates. Proteins 1997; Suppl 1: 215–220.
Schapira M, Totrov M, Abagyan R . Prediction of the binding energy for small molecules, peptides and proteins. J Mol Recognit 1999; 12: 177–190.
Michelangeli F, Munkonge FM . Methods of reconstitution of the purified sarcoplasmic reticulum (Ca(2+)-Mg2+)-ATPase using bile salt detergents to form membranes of defined lipid to protein ratios or sealed vesicles. Anal Biochem 1991; 194: 231–236.
Michelangeli F, Colyer J, East JM, Lee AG . Effect of pH on the activity of the Ca2++Mg2(+)-activated ATPase of sarcoplasmic reticulum. Biochem J 1990; 267: 423–429.
Acknowledgements
This work was supported by a FDCT grant from the Macao Science and Technology Development Fund (Project code: 013/2012/A1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by H-U Simon
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Wong, V., Li, T., Law, B. et al. Saikosaponin-d, a novel SERCA inhibitor, induces autophagic cell death in apoptosis-defective cells. Cell Death Dis 4, e720 (2013). https://doi.org/10.1038/cddis.2013.217
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.217
Keywords
This article is cited by
-
Saponins as cytotoxic agents: an update (2010–2021). Part II—Triterpene saponins
Phytochemistry Reviews (2023)
-
Hugan Qingzhi medication ameliorates free fatty acid-induced L02 hepatocyte endoplasmic reticulum stress by regulating the activation of PKC-δ
BMC Complementary Medicine and Therapies (2020)
-
Novel ginsenoside derivative 20(S)-Rh2E2 suppresses tumor growth and metastasis in vivo and in vitro via intervention of cancer cell energy metabolism
Cell Death & Disease (2020)
-
Saikosaponin D suppresses enterovirus A71 infection by inhibiting autophagy
Signal Transduction and Targeted Therapy (2019)
-
Autophagy and its potent modulators from phytochemicals in cancer treatment
Cancer Chemotherapy and Pharmacology (2019)