Abstract
Agrobacterium tumefaciens has the unique ability to mediate inter-kingdom DNA transfer, and for this reason, it has been utilized for plant genetic engineering. To increase the transformation frequency in plant genetic engineering, we focused on gamma-aminobutyric acid (GABA), which is a negative factor in the Agrobacterium-plant interaction. Recent studies have shown contradictory results regarding the effects of GABA on vir gene expression, leading to the speculation that GABA inhibits T-DNA transfer. In this study, we examined the effect of GABA on T-DNA transfer using a tomato line with a low GABA content. Compared with the control, the T-DNA transfer frequency was increased in the low-GABA tomato line, indicating that GABA inhibits T-DNA transfer. Therefore, we bred a new A. tumefaciens strain with GABA transaminase activity and the ability to degrade GABA. The A. tumefaciens strain exhibited increased T-DNA transfer in two tomato cultivars and Erianthus arundinacues and an increased frequency of stable transformation in tomato.
Similar content being viewed by others
Introduction
Agrobacterium is a genus of gram-negative bacteria that includes strains that are able to transfer genes that cause tumors (Agrobacterium tumefaciens or A. vitis) or hairy root (A. rhizogenes). A. tumefaciens, which induces crown gall disease in plants at the junction of the root and shoot, has been thoroughly studied, and the molecular mechanisms of gene transfer by this species have been elucidated1. A. tumefaciens harbors the Ti plasmid, which includes vir genes and transfer DNA (T-DNA) regions. The T-DNA regions contain oncogenic genes, such as indole-3-acetic acid (IAA), cytokinin and opine synthesis genes. Phenolic compounds and sugars exuded from the plant root induce vir gene expression2,3,4,5,6, after which the T-DNA region is excised by VirC and VirD. The excised single-stranded T-DNA forms a T-DNA complex with VirD and VirE. This T-DNA complex is introduced into plant cells via the type IV secretory system and then enters the plant nuclei through the intercellular transport system. The VirD and VirE proteins are subsequently stripped off, and the T-DNA is integrated into the plant nucleus. Expression of the T-DNA region integrated into the plant genome causes crown gall disease.
Although A. tumefaciens causes plant disease, its unique ability to transfer DNA presents the possibility that useful traits can be introduced into crops7,8, indicating potential for use in plant genetic engineering. To adapt the bacterium for plant genetic engineering, many efforts have been made to remove the oncogenic abilities of A. tumefaciens and invent a binary vector system9,10,11,12, representing the first step in adaption for plant genetic engineering. The next step is to expand the host range and increase the transformation frequency. Upregulation of vir gene expression is an effective strategy for broadening the host range of the bacterium and increasing transformation. Application of vir gene inducers2,3,4,5,6, utilization of super-binary vectors13,14,15 and employing a ternary transformation system16 improve the transformation efficiency. Depression of negative effectors of Agrobacterium-plant interactions is also effective. For instance, the phytohormone ethylene is a negative factor in Agrobacterium-plant interactions17,18,19,20. The ability of A. tumefaciens strains to reduce ethylene evolution from plants increases the transformation frequency21,22,23,24.
Gamma-aminobutyric acid (GABA) is a negative regulator of Agrobacterium-plant interaction through the quorum-sensing (QS) signal, which induces horizontal transfer of the Ti plasmid25,26. GABA imported into A. tumefaciens moderates crown gall disease symptoms via degradation of the QS signal. In tumors, accumulation of opines induces a QS signal, which enhances conjugation of the Ti plasmid. High accumulation of GABA depresses the QS signal, resulting in inhibition of Ti plasmid conjugation27, and then GABA plays a role in tumor at a later stage of the Agrobacterium-plant interaction. In a GABA-rich tobacco line, crown gall disease symptoms are less severe than in the wild type25. An atu2422-defeated A. tumefaciens strain, which lacks the ability to take up GABA, was shown to cause severe symptoms in tobacco26. However, no significant differences in vir gene expression or T-DNA transfer are observed between the atu2422-defeated strain and the wild-type strain26. From these results, it was concluded that GABA controls crown gall disease through a pathway independent of vir gene expression and T-DNA transfer. In contrast, a recent study showed that the expression of vir genes decreases in her1 (an Arabidopsis thaliana mutant line), which shows higher accumulation of GABA28. The accumulated GABA suppresses vir gene expression, which is essential for T-DNA transfer. Therefore, the accumulation of high GABA levels may inhibit T-DNA transfer via vir gene suppression.
In this study, we examined whether GABA affects T-DNA transfer using a low-GABA tomato line that we produced and previously characterized29. Additionally, we bred a new A. tumefaciens strain showing GABA transaminase activity (GabT) and the ability to degrade GABA. Although the atu3300 gene, which exhibits high similarity to the GABA transaminase gene, is located in the linear chromosome of A. tumefaciens strain C5830, activity of this gene has not yet been reported31. We then cloned the GABA transaminase gene (gabT) from Escherichia coli K12 32 and introduced it into A. tumefaciens. The ability of T-DNA transfer in the new A. tumefaciens strain was evaluated in two tomato cultivars (‘Micro-Tom’ and ‘Moneymaker’) and the grass E. arundinaceus, which is a potential biomass plant, and stable transformation was examined in tomato (‘Micro-Tom’).
Results
Inoculation of A. tumefaciens stimulates GABA accumulation during co-cultivation
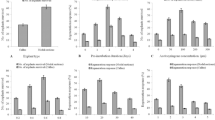
To evaluate the effect of A. tumefaciens inoculation on GABA accumulation, the cotyledon segments from 7-day-old tomato seedlings were used, and both un-inoculated and inoculated segments were prepared. After 3 days of co-cultivation, GABA accumulation in tomato (Micro-Tom) cotyledon segments was measured. In tomato cotyledon segments inoculated with A. tumefaciens (black bar, WT, Cotyledon), the GABA level was 5 times higher than in un-inoculated segments (white bar, WT, Cotyledon) (Fig. 1A). To determine whether wounding caused GABA accumulation, intact seedlings (white bar, WT, Seedling) and tomato cotyledon segments (white bar, WT, Cotyledon) were compared. In the un-inoculated treatment, the GABA content in the cotyledon segments was the same as in the intact seedlings. These results indicate that GABA accumulation was stimulated by A. tumefaciens infection, rather than that by wound stress.
(A) GABA content in intact seedlings and cotyledon segments of Micro-Tom. White and black indicate un-inoculated and inoculated, respectively. Bars indicate the standard deviation (n = 3). Different letters indicate significant differences by Tukey’s test (P < 0.01). (B) Classification of GUS-stained cotyledon explants. GUS-stained tomato cotyledons were categorized based on the stained area, as follows: 20% or more of the area, 10% or more, 5% or more, or less than 5%. The frequency of each category of GUS-stained tomato explants is shown. Bacterial strains exhibiting significant differences (Student’s t-test and Kruskal-Wallis test; P < 0.01, n = 80) are indicated with different letters. WT and RNAi refer to the non-transgenic line and the RNAi-SlGADall line, respectively. In GAD RNAi lines, the expression of GAD1, GAD2, and GAD3 was much lower than in WT.
GABA affects T-DNA transformation in tomato
If accumulation of GABA during co-cultivation inhibits T-DNA transfer, the frequency of T-DNA transfer should be increased in low-GABA plants. We investigated whether GABA affects T-DNA transfer using the GAD RNAi transgenic S. lycopersicum cv. Micro-Tom (RNAi-SlGADall)29. In the RNAi-SlGADall line, the expression levels of GAD genes (GAD1, GAD2, and GAD3) involved in GABA synthesis were reduced, and the accumulation of GABA was low compared with the non-transgenic line (Fig. 1A). To evaluate the T-DNA transfer efficiency, 80 tomato cotyledon segments were prepared from 7-day-old seedlings and inoculated with A. tumefaciens GV2260 (pIG121-Hm). The uidA gene was used as an indicator of T-DNA transfer. After 3 days of co-cultivation, tomato segments were stained with the GUS substrate, and the stained area was analyzed with ImageJ24 (https://imagej.nih.gov/ij/). The degree of staining was categorized into four classes, and the frequency of each class was calculated. In the low-GABA tomato line, the frequency of the staining class “10% or more” was increased compared with non-transgenic tomato (Fig. 1B). The same tendency was observed in three repetitions. This result showed that inhibition of GABA accumulation during co-cultivation induced by A. tumefaciens inoculation increased the T-DNA transfer level.
Introduction of the gabT gene into A. tumefaciens results in GabT activity
In this study, we used A. tumefaciens strain GV2260, which has the same chromosome as strain C58 33. GV2260 has the atu3300 gene in its linear chromosome30. Atu3300 is predicted by Pfam (http://pfam.sanger.ac.uk) to include an Aminotrans_3 (aminotransferase class-III) domain, which is characteristic of aminotransferases34. The amino acid sequence of Atu3300 was compared with GabT from E. coli K12, P. syringae pv. tomato DC3000 and P. aeruginosa PAO (Fig. 2A); these GabTs display activity and function in cell growth and plant-microbial interactions32,35,36. Four of the GabTs (with the exception of Atu3300) were well conserved and exhibited two conserved motifs (Thr-Phe-Ala-Lys-Ser-Ile-Ala) and (Leu-Arg-Ile-Leu-Val) that correlated well with the consensus sequence of aminotransferases from Salmonella typhimurium37, E. coli K12, and Saccharomyces cerevisiae32. In contrast, Atu3300 did not contain these conserved motifs. Therefore, we assumed that Atu3300 shows very low or no GABA transaminase activity. The GABA accumulation induced by A. tumefaciens inoculation during co-cultivation inhibited T-DNA transfer (Fig. 1), which suggests that A. tumefaciens exhibits GabT activity and that degradation of GABA would be effective for increasing the T-DNA transfer frequency. To introduce GABA transaminase activity to A. tumefaciens, the GABA transaminase gene (gabT) was cloned from E. coli K12 and into the broad-host-range plasmid pBBR1MCS-538, and the resulting plasmid was designated pBBRgabT. The gabT gene was expressed under the control of the lac promoter21 (Fig. 2B). Then, GabT activity was measured by monitoring glutamic acid accumulation in the reaction buffer. Only the reaction buffers containing the A. tumefaciens GV2260 lysate (pIG121-Hm, pBBRgabT) linearly increased the glutamic acid content (Fig. 2C). This result indicated that A. tumefaciens GV2260 exhibited no GABA transaminase activity, and we succeeded in conferring GabT activity on A. tumefaciens by introducing the gabT gene from E. coli K12.
(A) Amino acid sequence comparison of GABA transaminase from E. coli K12 (gabTSCA772438), P. syringae pv. tomato DC3000 (gabT1: gabT1-PSPTO0259, gabT2: gabT2-PSPTO0301), P. aeruginosa PAO1 (gabT-PA0266). “ClustalW2” (http://www.ebi.ac.uk/Tools/msa/clustalw2/) and “Jalview” (http://www.jalview.org) were used for calculation of amino acid multiple sequence alignment and display the alignment result, respectively. The residues were coloured according to their physicochemical properties as follows; Aliphatic/Hydrophobic, Aromatic, Positive, Negative, Hydrophilic, Conformationally Special and Cystein. Arrowheads indicate 3 of the 12 invariant amino acid residues among 16 aminotransferases with highly homologous peptides. The red box indicates a conserved motif (Ser [or Thr]-X-X-Lys) in the pyridoxalphosphate-binding peptide of aspartate aminotransferase (AAT) and histidinol-phosphate transaminases. The blue box indicates near identity between the homologous peptides of the histidinol-phosphate transaminases from E. coli K12 and Saccharomyces cerevisiae. (B) Construction of a plasmid for the expression of GABA transaminase (gabT) in A. tumefaciens. HindIII and XbaI fragments (ca. 1.6 kb) containing the GABA transaminase gene from E. coli K12 were ligated into the HindIII and XbaI sites of the broad-host-range plasmid pBBR1MCS-5, resulting in pBBRgabT. The expression of the GABA transaminase gene gabT was under the control of the lac promoter. MCS: multiple cloning site. (C) Detection of GABA activity in A. tumefaciens. Glutamic acid accumulation in the reaction buffer was measured according to the method of Akihiro et al.48. The open and closed circles indicate A. tumefaciens GV2260 (pBBRgabT, pIG121-Hm) and A. tumefaciens GV2260 (pBBR1MCS-5, pIG121-Hm), respectively. Bars represent the standard deviation (n = 3).
GabT activity enhances the T-DNA transfer ability of A. tumefaciens
To evaluate the effect of GabT on A. tumefaciens, two tomato cultivars (‘Micro-Tom’ and ‘Moneymaker’) and the grass E. arundinaceus were used. The uidA gene was employed as an indicator of T-DNA transfer (Fig. 3A). Compared with A. tumefaciens without GabT, the transformation frequency associated with a high degree of staining (10% or more) was increased approximately 2.1 or 4.0 times by the inoculation of A. tumefaciens carrying GabT into ‘Micro-Tom’ and ‘Moneymaker’, respectively (Fig. 3A–C). In E. arundinaceus, to estimate the T-DNA transfer frequency, we measured the number of blue spots on the surfaces of calli per 1 g of calli. The number of GUS spots per 1 g of calli increased four-fold when A. tumefaciens GV2260 (pIG121-Hm, pBBRgabT) was used (Fig. 3D). These results clearly showed that A. tumefaciens with GabT activity increased the frequency of T-DNA transfer. Therefore, degradation of GABA during co-cultivation is an effective method for increasing T-DNA transfer.
(A) GUS-stained explants of tomato (Micro-Tom). Explants were prepared from 7-day-old seedlings. After 3 days of co-cultivation, the explants were stained. (B) Estimation of T-DNA transfer for A. tumefaciens with GabT in Micro-Tom (tomato). GUS-stained tomato cotyledons were categorized based on the stained area, as follows: 20% or more of the area, 10% or more, 5% or more, or less than 5%. Different letters indicate significant differences according to Student’s t-test or the Kruskal-Wallis test; P < 0.01 (n = 80). (C) Assessment of T-DNA transfer for A. tumefaciens with GabT in ‘Moneymaker’. GUS-stained cotyledons were categorized as follows: 20% or more of the area, 10% or more, 5% or more, or less than 5%. Different letters indicate significant differences according to Student’s t-test and the Kruskal-Wallis test; P < 0.01 (n = 80). (D) Occurrence of T-DNA transformation in E. arundinaceus. The number of GUS-stained spots per 1 g of E. arundinaceus calli was counted for each treatment. The bars indicate the standard deviation (n = 3). Different letters indicate values that were significantly different according to Student’s t-test (P < 0.05). Control: A. tumefaciens GV2260 (pBBRMCS1–5, pIG121-Hm); gabT: A. tumefaciens GV2260 (pBBRgabT, pIG121-Hm).
GabT activity in A. tumefaciens enhances stable transformation
Because A. tumefaciens with GabT activity exhibited an increased T-DNA transfer frequency, we assumed that inoculation of the strain with GabT would also increase the frequency of stable transformation. To examine this hypothesis, almost 100 explants were inoculated with A. tumefaciens GV2260 (pIG121-Hm, pBBRgabT) or A. tumefaciens GV2260 (pIG121-Hm, pBBR1MCS-5). Transformation techniques were used to characterize or add gene functions to the plants. On the other hand, transformation procedure causes chromosome doubling and multi-copy insertion, which would change the phenotype, such as size, growth speed, color, and so on. Therefore, chromosome doubling and multi-copy insertion must be excluded when the phenotype or trait of transgenic plant are evaluated. The stable transformation frequency was calculated with diploid and single-copy-number plants. The stable transformation frequencies were 10.1 ± 0.7% and 4.3 ± 1.0% (mean ± SD of three repetitions using 100 explants in each experiment) (Table 1) in A. tumefaciens GV2260 (pIG121-Hm, pBBRgabT) and A. tumefaciens GV2260 (pIG121-Hm, pBBR1MCS-5), respectively (Fig. 4A). A. tumefaciens with GabT exhibited approximately 2.5 times the stable transformation frequency of A. tumefaciens without GabT. No significant differences in the appearance of diploidy were noted in the two types of A. tumefaciens (Fig. 4B), and all of the lines we obtained were independent and did not contain a cloned plant (Fig. 4C). The appearance of double and multiple copy lines was the same between the two A. tumefaciens strains (Fig. 4D). These results showed that A. tumefaciens with GabT activity exhibited an increase in stable transformation, without effects on ploidy and copy number.
(A) Effect of GabT activity in A. tumefaciens on stable transformation. Error bar shows the standard deviation (SD) (n = 3). Different characters indicate significant differences by Student’s t-test (P < 0.01). (B) Frequency of diploid appearance. Rooting shoots were checked for polyploidy. Closed and open boxes represent diploid and polyploid, respectively. Bars indicate the standard deviation (n = 3). (C) Southern blot analysis. (D) The frequency of each copy number in diploid rooting shoots. The T-DNA copy number inserted into the genome was detected through Southern blotting analysis. Bacterial strains are indicated as follows: Control: A. tumefaciens GV2260 (pBBR1MCS-5, pIG121-Hm); gabT: A. tumefaciens GV2260 (pBBRgabT, pIG121-Hm). Different letters indicate values that were significantly different by Tukey’s test (P < 0.05).
Discussion
Our results showed that A. tumefaciens infection increased the GABA contents of tomato cotyledon segments during co-cultivation. Based on this result, we assumed that the GABA accumulated during co-cultivation affected T-DNA transfer. To test this hypothesis, we used the GAD RNAi tomato, which showed a low GABA level during co-cultivation, and A. tumefaciens with the ability to degrade GABA. Compared with the control, the T-DNA transfer frequency was increased in the GAD RNAi tomato line (Fig. 1B). A. tumefaciens with GabT activity (Fig. 2B) also increased the T-DNA transfer in ‘Micro-Tom’, ‘Moneymaker’, and E. arundinaceus (Fig. 3). The frequency of stable transformation was also increased in ‘Micro-Tom’ (Table 1). These results clearly show that the GABA accumulated during co-cultivation inhibited both T-DNA transfer and stable transformation.
The present study clearly showed that GABA that accumulates during co-cultivation, induced by inoculation with A. tumefaciens, inhibits T-DNA transfer (Figs 1 and 3). These results suggest that GABA affects an early step of the Agrobacterium-plant interaction. On the other hands, previous studies conclude that GABA plays a role in tumors at a later stage of the Agrobacterium-plant interaction, and not in earlier stages26. These inconsistent results might be attributed to differences in the applied evaluation of T-DNA transfer methods, as in the previous study, T-DNA transfer was evaluated based on the proportion of stained leaf discs per infection26, whereas we evaluated the frequency of T-DNA transfer in each tomato segment or 1 g of E. arundinaceus calli. These methods allowed us to evaluate the T-DNA transfer in detail. Based on these results, we concluded that the high accumulation of GABA induced by A. tumefaciens infection inhibits T-DNA transfer and that GABA is involved in both the early and later stages of the Agrobacterium-plant interaction.
The genome of A. tumefaciens strain C58 encodes atu3300 in its linear chromosome30. This gene has 68.1% similarity and 23.7% identity with GABA transaminase from E. coli (GabT). Atu3300 included an aminotransferase class-III and was predicted to have GabT activity. However, although Atu3300 has an aminotransferase class-III domain, it did not show GabT activity (Fig. 2C). In comparisons of GabT with several aspartate, tyrosine, and histidinol-phosphate transaminases, previous studies predicted that two motifs, containing Lys268 and Arg398, are involved in active-site formation in the GabT protein32,39. The GabT proteins from E. coli K12, P. syringae pv. tomato DC3000 and P. aeruginosa PAO have these motifs. However, Atu3300 does not contain these motifs (Fig. 2A). According to the present study and previous work, the lack of GabT activity in A. tumefaciens appears to result from the lack of these motifs in Atu3300 (Fig. 2A), indicating that the two motifs in GabT containing Lys268 and Arg398 are essential for this activity.
Removal of GABA is more effective than vir gene stimulation. Our results indicated that the reduction of GABA during co-cultivation increased T-DNA transformation under high vir gene expression induced by 200 μM acetosyringone (Figs 1B and 3B,C,D), which is sufficient to fully induce the vir gene20. Although the possibility that GABA is involved in vir gene expression cannot be discarded because of two contradictory results26,28, our results show that GABA has a stronger effect on T-DNA transfer than stimulation of vir gene expression. As the QS signal accumulates in tumor, which is a late stage of the Agrobacterium-plant interaction40, GABA should inhibit T-DNA transfer independently of a QS signal in an early stage. This conclusion suggests the existence of an “unknown pathway” independent of the QS signal. The unknown pathway involved in GABA function might have a stronger influence on T-DNA transformation.
Depending on the species or cultivar, the effect of inhibiting negative factors seems to differ. In ‘Moneymaker’ and E. arundeinaceus, the new A. tumefaciens strain with GabT activity is more effective than the previously used strain that inhibits ethylene during co-cultivation24. In addition to GABA25,26 and ethylene17,18,19,20, salicylic acid, cytokinin, auxin and abscisic acid are also known negative factors in the Agrobacterium-plant interaction41,42,43,44. Selecting negative factors based on the species involved might be important for increasing transformation and broadening host ranges. Moreover, removing multiple such negative factors would have an effect on the transformation frequency and adaptation to a wide range of host plants.
We succeeded in producing an A. tumefaciens strain with improved potential for transformation by imbuing it with the ability to remove GABA, which is a negative factor in the Agrobacterium-plant interaction. Removal of GABA increased the transformation frequency approximately 2.5 times. Therefore, this newly bred bacterium enables us to decrease the number of cotyledons used for transformation by 60%. Thus, this strain allows us to reduce the time and the labor required for transformation. Based on this result, we conclude that this new strain might be a useful tool for plant genetic engineering.
For plant molecular breeding, the genetic modification technique is very important, and Agrobacterium-mediated transformation is the most frequently used method. Therefore, substantial effort has been expended to adapt Agrobacterium-mediated transformation to a wide variety of plants. However, species and genotypes recalcitrant to genetic transformation still exist, and the improvement has been required. Transformation process include the three steps: first step is T-DNA transfer, second step is selection of transgenic cells, and the third step is regeneration from the transgenic cells. Previous study indicated japonica rice showed higher transformation frequency than indica rice. Comparing with japonica rice, T-DNA transfer frequency was very low in indica rice45. This result suggested that improvement of this process was effective. Indeed, improvement of T-DNA transfer increased stable transformation46,47. Inevitable, we focused on improvement of T-DNA transfer through new bred A. tumefaciens strain. Our new bred A. tumefaciens strain with GABA degradation ability improved T-DNA transfer in two genotypes of tomato and E. arundinaceus, and increased stable transformation in tomato ‘Micro-Tom’. We succeeded in producing an A. tumefaciens strain with improved potential for transformation by imbuing it with the ability to degrade GABA. The strain created in this study might represent a new system for improving the transformation frequency in recalcitrant species and genotypes.
Materials and Methods
Bacterial strains and culture conditions
E. coli K12 and DH5α were grown at 37 °C in Luria Broth (LB) medium (1% bacto-tryptone, 0.5% yeast extract, and 0.5% NaCl). A. tumefaciens strain GV2260, a derivative of the C58 strain, was grown at 28 °C in LB medium. Antibiotics were added at the following final concentrations: ampicillin at 100 μg ml−1 for E. coli K12 and A. tumefaciens and gentamicin at 50 μg ml−1, kanamycin at 50 μg ml−1, and spectinomycin at 50 μg ml−1 for A. tumefaciens.
A. tumefaciens growth conditions
A. tumefaciens GV2260 was cultured on solid LB medium at 28 °C for 2 days. A single colony was then picked and cultured in 2 ml of LB medium at 28 °C at 200 rpm for 2 days until the pre-culture reached the stationary phase. After pre-culture, 15 μl of culture medium was added to 15 ml of LB medium containing antibiotics, and culturing was continued for 20–22 hours at 28 °C, with shaking at 200 rpm. After the culture reached an OD600 of 0.8–1.0, to collect bacterial cells, the culture was centrifuged at 3,000 × g for 10 min.
Construction of the gabT expression plasmid
The gabT gene was cloned from the genome of E. coli K12 via polymerase chain reaction (PCR) using the primers gabTF (5′-aagcttaatgaacagcaataaagagtt-3′) and gabTR (5′-tctagactactgcttcgcctcatcaaaac-3′). The 1294-bp amplified fragment was inserted into the pCRTOPO vector (Invitrogen, Carlsbad, CA, USA) to generate pCRgabT. We checked the sequence using an ABI Sequence Analyzer (Applied Biosystems, MA, USA), and the sequence was found to be identical to accession number 6061113 in the National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm.nih.gov) database. The gabT fragment was subcloned into the multiple cloning site of the broad-host-range plasmid pBBR1MCS-538 using HindIII and XbaI (New England Biolabs, Hirchin, UK) to generate a lacZ::gabT translational fusion (pBBRgabT) (Fig. 2B).
GabT activity in A. tumefaciens
A pellet of A. tumefaciens cells was re-suspended in 100 μl of BugBuster Master mix (Novagen, MA, USA) for lysate preparation. The protein concentration of the lysate was measured by a BCA Protein Assay Kit (Novagen, MA, USA). The protein content was adjusted to 100 μg per reaction mixture. The reaction mixture contained 0.1 M bicine-NaOH, 0.1 M pyridoxal phosphate, 10 mM 2-ketoglutarate, 10 mM GABA, and a protease inhibitor cocktail. The reaction mixture was incubated at 37 °C for 0, 10, 20, 30, 60, 120 or 180 min. GabT metabolizes GABA to glutamate; therefore, to estimate GabT activity, we detected the glutamate concentration in the reaction mixture using a Yamaki glutamate assay kit (Yamaki, Tokyo, Japan)48.
Plant material
Non-transgenic tomato seeds (Solanum lycopersicum ‘Micro-Tom’ or ‘Moneymaker’) and a GAD-suppressed ‘Micro-Tom’ transgenic line (RNAi-SlGADall) that we used in a previous study29 were employed in this study. ‘Moneymaker’ exhibits medium-sized fruits and is a commercialized cultivar. The seeds were washed with 70% ethanol for 10 seconds, sterilized with 5% hypochlorous acid containing 10% Triton X-100 for 45 min, and washed three times with sterilized water. After the third wash, the seeds were kept in water for 2 days. The sterilized tomato seeds were sown on Murashige and Skoog (MS) medium49 containing 15 g l−1 sucrose (Wako, Tokyo, Japan) and 0.3% Gelrite (Wako, Tokyo, Japan) and then grown for 7 days.
Calli of E. arundinaceus, known as a high biomass producer, were kindly provided by Prof. Masahiro Mii of Chiba University, Japan. The calli induced from the seeds on MS medium containing 1 g l−1 casamino acids, 2 mg l−1 2,4-dichlorophenoxyacetic acid (2, 4-D), 0.2 mg l−1 6-benzylaminopurine (BAP), 30 g l−1 4-O-α-D-glycopyranosyl-D-glycopyranose (maltose H) (Wako, Tokyo, Japan) and 0.3% Gelrite were subcultured for 2 weeks before A. tumefaciens inoculation.
Measurement of GABA content
To measure the GABA content, tomato seedlings and cotyledon segments were prepared. Approximately 50 mg of powdered sample was added to 500 μl of 8% (w/v) trichloroacetic acid and mixed via vortexing for 30 sec. The mixture was subsequently centrifuged at 10,000 × g for 20 min at 4 °C, and 300 μl of the supernatant was transferred to a new tube and mixed vigorously with 400 μl of pure diethyl ether for 10 min. The solution was then centrifuged at 10,000× g for 10 min at 4 °C, after which the upper phase of the diethyl ether was removed and the previous step was repeated. After centrifugation again at 10,000× g for 10 min at 4 °C, the upper phase was removed. To completely remove the remaining diethyl ether, the samples were incubated under a draft of air for 30 min. A 30 μl aliquot of the lower phase was transferred to a new 1.5 ml tube and dried using an evaporator (CVE3100, TOKYO RIKAKIKAI, Tokyo, Japan). The dried samples were washed with 150 μl of sterile distilled water. After washing, the samples were dissolved in 0.1 N HCl for amino acid analysis (JLC-500/V2, Japan Electron Optics Laboratory, Tokyo, Japan).
Agrobacterium-mediated T-DNA transfer
Bacterial cells were re-suspended in liquid MS medium containing 30 g l−1 glucose and 200 μM acetosyringone (Wako, Tokyo, Japan) at pH 5.2, and the OD600 was adjusted to 0.4–0.5. Cotyledons of 7-day-old tomato seedlings were cut into four pieces and subjected to inoculation with A. tumefaciens. Eighty explants were subjected to each treatment. The inoculated explants were cultured on co-cultivation medium (pH 5.2) containing MS salts, 30 g l−1 glucose, 200 μM acetosyringone and 0.3% Gelrite (Wako, Tokyo, Japan) at 25 °C for 3 days in the dark. After 3 days of co-cultivation, the tomato explants were assayed histochemically for β-glucuronidase (GUS) activity using GUS staining solution containing 100 mM NaPO4, 10 mM EDTA, 2.5 mM potassium ferrocyanide, 2.5 mM potassium ferrocyanide, 0.1% Triton X-100, and 0.5 mg ml−1 X-Gluc.
Calli of E. arundinaceus that had been subcultured for 2 weeks were also inoculated with A. tumefaciens. After co-cultivation, the GUS activity of E. arundinaceus calli was histochemically assayed with GUS staining solution, as described above.
Stable tomato transformation
After 3 days of co-cultivation, ‘Micro-Tom’ cotyledon segments were placed on callus-induction medium (MS medium containing 0.3% Gelrite, 1.5 mg l−1 zeatin, 100 mg l−1 kanamycin, and 375 mg l−1 augmentin [GlaxoSmithKline, MDX, UK]) for 4 weeks. Calli that formed from the segments were cultured on shoot-induction medium (MS medium containing 0.3% Gelrite, 1.0 mg l−1 zeatin, 100 mg l−1 kanamycin, and 375 mg l−1 augmentin) for 4 weeks. The shoots were then placed on rooting medium, which consisted of half-strength MS medium, 0.3% Gelrite (Wako, Tokyo, Japan), 100 mg l−1 kanamycin, and 375 mg l−1 augmentin, for 2 weeks. Tissues were subcultured every 10–14 days. The ploidy of the rooting shoots was checked via flow cytometry.
Ploidy analysis
One square centimeter of leaf was cut from the rooting shoots and chopped in 250 μl of nucleus-extraction solution (CyStain UV Precise P, Sysmex, Hyogo, Japan). To purify the nucleus extraction solution, 1 mm2 mesh was used. After purification, 1 ml of staining solution (CyStain UV Precise P, Sysmex, Hyogo, Japan) was added, followed by incubation for 1 min. This solution was applied to an Attune focusing analyzer (ABI, MA, USA), and diploid plants were selected. The diploid plants were planted on solid medium and acclimatized.
Southern blot analysis
Genomic DNA was extracted from young tomato leaves using the Maxwell 16 System DNA Purification kit (Promega, WI, USA). The purified DNA was digested with HindIII, then electrophoretically separated in a 0.8% agarose gel and transferred to Gene Screen Plus nylon membranes (Roche Diagnostics, Basel, Switzerland) with 20 × saline-sodium citrate buffer. After ultraviolet cross-linking, the membranes were hybridized in a solution containing 7% sodium dodecyl sulfate, 50% deionized formamide, 50 mM sodium phosphate (pH 7.0), 2% blocking solution, 0.1% N-lauroylsarcosine, 0.75 M NaCl, and 75 mM sodium citrate at 42 °C overnight. For hybridization, a digoxigenin (DIG)-labeled DNA probe specific for nptII (0.8 Kb) was used. A DIG-labeled probe was generated using DIG-High Prime, and the DIG signal was detected according to the manufacturer’s protocol (Roche Diagnostics, Basel, Switzerland).
Additional Information
How to cite this article: Nonaka, S. et al. An Agrobacterium tumefaciens Strain with Gamma-Aminobutyric Acid Transaminase Activity Shows an Enhanced Genetic Transformation Ability in Plants. Sci. Rep. 7, 42649; doi: 10.1038/srep42649 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Nester, E. W. Agrobacterium: nature’s genetic engineer. Front Plant Sci. 5, 730, 10.3389/fpls.2014.00730. eCollection 2014. (2015).
Stachel, S. E., Messens, E., van Montagu, M. & Zambryski, P. Identification of the signal molecules produced by wounded plant cells that activate T-DNA transfer in Agrobacterium tumefaciens . Nature 318, 624–629 (1985).
Stachel, S. E., Nester, E. W. & Zambryski, P. C. A plant cell factor induces Agrobacterium tumefaciens vir gene expression. Proc. Natl. Acad. Sci. USA 83, 379–383 (1986).
Cangelosi, G. A., Ankenbauer, R. G. & Nester, E. W. Sugars induce the Agrobacterium virulence genes through a periplasmic binding protein and a transmembrane signal protein. Proc. Natl. Acad. Sci. USA 87, 6708–6712 (1990).
He, F. et al. Molecular basis of ChvE function in sugar binding, sugar utilization, and virulence in Agrobacterium tumefaciens . J. Bacteriol. 191, 5802–5813 (2009).
Hu, X., Zhao, J., DeGrado, W. F. & Binns, A. N. Agrobacterium tumefaciens recognizes its host environment using ChvE to bind diverse plant sugars as virulence signals. Proc. Natl. Acad. Sci. USA 110, 678–683 (2013).
Pereira, A. A transgenic perspective on plant functional genomics. Transgenic Res. 9, 245–260 (2000).
Tyagi, A. K. & Mohanty, A. Rice transformation for crop improvement and functional genomics. Plant Sci. 158, 1–18 (2000).
Zambryski, P. et al. Ti plasmid vector for the introduction of DNA into plant cells without alteration of their normal regeneration capacity. EMBO J. 2, 2143–2150 (1983).
Hoekema, A., Hirsch, P. R., Hooykaas, P. J. J. & Schilperoort, R. A binary plant vector strategy based on separation of vir- and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303, 179–180 (1983).
Bevan, M. Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res. 12, 8711–8721 (1984).
Komari, T. et al. Binary vectors and super-binary vectors. Methods Mol Biol. 343, 15–41 (2006).
Komari, T. Transformation of cultured cells of Chenopodium quinoa by binary vectors that carry a fragment of DNA from the virulence region of pTiBo542. Plant Cell Rep. 9, 303–306 (1990).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 6, 271–282 (1994).
Ishida, Y. et al. High efficiency transformation of maize (Zea mays L.) mediated by Agrobacterium tumefaciens . Nat. Biotechnol. 14, 745–750 (1996).
van der Fits, L., Deakin, E. A., Hoge, J. H. C. & Memelink, J. The ternary transformation system: constitutive virG on a compatible plasmid dramatically increases Agrobacterium-mediated plant transformation. Plant Mol. Biol. 43, 495–502 (2000).
Davis, M. E., Miller, A. R. & Lineberger, R. D. Studies on the effects of ethylene on transformation of tomato cotyledons (Lycopersicon esculentum Mill.) by Agrobacterium tumefaciens . J. Plant Physiol. 139, 309–312 (1992).
Ezura, H., Yuhashi, K. I., Yasuta, T. & Minamisawa, K. Effect of ethylene on Agrobacterium tumefaciens-mediated gene transfer to melon. Plant Breed. 119, 75–79 (2000).
Han, J. S. et al. Agrobacterium-mediated transformation of bottle gourd (Lagenaria siceraria Standl.). Plant Cell Rep. 23, 692–698 (2005).
Nonaka, S. et al. Ethylene production in plants during transformation suppresses vir gene expression in Agrobacterium tumefaciens . New Phytol. 178, 647–656 (2008a).
Nonaka, S. et al. 1-Aminocyclopropane-1-carboxylate deaminase enhances Agrobacterium tumefaciens-mediated gene transfer into plant cells. Appl. Environ. Microbiol. 74, 2526–2528 (2008b).
Ntui, V. O. et al. An efficient Agrobacterium tumefaciens-mediated genetic transformation of “Egusi” melon (Colocynthis citrullus L.). Plant Cell Organ. Cult. 103, 15–22 (2010).
Hao, Y. T., Charles, C. & Glick, B. R. ACC deaminase increases the Agrobacterium tumefaciens-mediated transformation frequency of commercial canola cultivars. FEMS Microbiol. Lett. 307, 185–190 (2010).
Someya, T., Nonaka, S., Nakamura, K. & Ezura, H. Increased 1-aminocyclopropane-1-carboxylate deaminase activity enhances Agrobacterium tumefaciens-mediated gene delivery into plant cells. Microbiologyopen 2, 873–880 (2013).
Chevrot, R. et al. GABA controls the level of quorum-sensing signal in Agrobacterium tumefaciens . Proc. Natl. Acad. Sci. USA 103, 7460–7464 (2006).
Haudecoeur, E. et al. Proline antagonizes GABA-induced quenching of quorum-sensing in Agrobacterium tumefaciens . Proc. Natl. Acad. Sci. USA 106, 14587–14592 (2009).
Deeken, R. et al. An integrated view of gene expression and solute profiles of Arabidopsis tumors: a genome-wide approach. Plant Cell 18, 3617–3634 (2006).
Lang, J. et al. The plant GABA signaling downregulates horizontal transfer of the Agrobacterium tumefaciens virulence plasmid. New Phytol. 210, 974–983 (2016).
Takayama, M. et al. Tomato glutamate decarboxylase genes SlGAD2 and SlGAD3 play key roles in regulating γ-aminobutyric acid levels in tomato (Solanum lycopersicum). Plant Cell Physiol. 56, 1533–1545 (2015).
Wood, D. W. et al. The genome of the natural genetic engineer Agrobacterium tumefaciens C58. Science 294, 2317–2323 (2001).
Khan, S. R. & Farrand, S. K. The BlcC (AttM) lactonase of Agrobacterium tumefaciens does not quench the quorum-sensing system that regulates Ti plasmid conjugative transfer. J. Bacteriol. 191, 1320–1329 (2009).
Bartsch, K., von Johnn-Marteville, A. & Schulz A. Molecular analysis of two genes of the Escherichia coli gab cluster: nucleotide sequence of the glutamate: succinic semialdehyde transaminase gene (gabT) and characterization of the succinic semialdehyde dehydrogenase gene (gabD). J. Bacteriol. 172, 7035–7042 (1990).
Deblaere, R. et al. Efficient octopine Ti plasmid-derived vectors for Agrobacterium-mediated gene transfer to plants. Nucleic. Acids. Res. 13, 4777–4788 (1985).
Shen, B. W. et al. Crystal structure of human recombinant ornithine aminotransferase. J. Mol. Biol. 277, 81–102 (1998).
Chou, H. T., Kwon, D. H., Hegazy, M. & Lu, C. D. Transcriptome analysis of agmatine and putrescine catabolism in Pseudomonas aeruginosa PAO1. J. Bacteriol. 190, 1966–1975 (2008).
Park, D. H. et al. Mutations in γ-aminobutyric acid (GABA) transaminase genes in plants or Pseudomonas syringae reduce bacterial virulence. Plant J. 64, 318–330 (2010).
Hsu, L. C., Okamoto, M. & Snell, E. E. L-Histidinol phosphate aminotransferase from Salmonella typhimurium. Kinetic behavior and sequence at the pyridoxal-P binding site. Biochimie. 71, 477–489 (1989).
Kovach, M. E. et al. Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176 (1995).
Mehta, P. K., Hale, T. I. & Christen, P. Evolutionary relationships among aminotransferases. Tyrosine aminotransferase, histidinol-phosphate aminotransferase, and aspartate aminotransferase are homologous proteins. Eur. J. Biochem. 186, 249–253 (1989).
Lang, J. & Faure, D. Functions and regulation of quorum-sensing in Agrobacterium tumefaciens . Front Plant Sci. 5, 14, doi: 10.3389/fpls.2014.00014 (2014).
Yuan, Z. C. et al. The plant signal salicylic acid shuts down expression of the vir regulon and activates quormone-quenching genes in Agrobacterium . Proc. Natl. Acad. Sci. USA 104, 11790–11795 (2007).
Hwang, H. H. et al. Agrobacterium-produced and exogenous cytokinin-modulated Agrobacterium-mediated plant transformation. Mol. Plant Pathol. 11, 677–690 (2010).
Lee, C. W. et al. Agrobacterium tumefaciens promotes tumor induction by modulating pathogen defense in Arabidopsis thaliana . Plant Cell. 21, 2948–2962 (2009).
Rico, A. et al. Agroinfiltration reduces ABA levels and suppresses Pseudomonas syringae-elicited salicylic acid production in Nicotiana tabacum . PLoS One, 10.1371/journal.pone.0008977 (2010).
Tie, W. et al. Reasons for lower transformation efficiency in indica rice using Agrobacterium tumefaciens-mediated transformation: lessons from transformation assays and genome-wide expression profiling. Plant Mol. Biol. 78, 1–18, doi: 10.1007/s11103-011-9842-5 (2012).
Faizal, A. & Geelen, D. Agroinfiltration of intact leaves as a method for the transient and stable transformation of saponin producing Maesa lanceolata . Plant Cell Rep. 31, 1517–1526, doi: 10.1007/s00299-012-1266-4 (2012).
Nanasato, Y. et al. Improvement of Agrobacterium-mediated transformation of cucumber (Cucumis sativus L.) by combination of vacuum infiltration and co-cultivation on filter paper wicks. Plant Biotechnol. Rep. 7, 267–276 (2013).
Akihiro, T. et al. Biochemical mechanism on GABA accumulation during fruit development in tomato. Plant Cell Physiol. 49, 1378–1389 (2008).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Acknowledgements
We appreciate the help of Prof. Mii (Chiba University, Japan) for kindly providing the E. arundinaceus calli. We also thank Prof. Nakamura (Chiba University, Japan) and Prof. Mitsui (Tohoku University, Japan) for the gift of the pEKH2 plasmid and the pBBR1MCS-5 plasmid, respectively. This research was supported in part by grants from the New Energy and Industrial Technology Development Organization (NEDO, Grant Number ADD21105) to HE, a Grant-in-Aid for Young Scientists (B) (Grant Number 24780001) from JSPS KAKENHI to SN, and a Cooperative Research Grant from the Gene Research Centre of the University of Tsukuba to HE and SN.
Author information
Authors and Affiliations
Contributions
S.N., S.Z. and H.E. conceived the experiments. S.N. and S.Z. conducted the experiments. All authors analyzed the results, wrote and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Nonaka, S., Someya, T., Zhou, S. et al. An Agrobacterium tumefaciens Strain with Gamma-Aminobutyric Acid Transaminase Activity Shows an Enhanced Genetic Transformation Ability in Plants. Sci Rep 7, 42649 (2017). https://doi.org/10.1038/srep42649
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep42649
This article is cited by
-
Enigma of recalcitrance to tissue culture in the oilseed crop Sesamum indicum L.—a review
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
Evaluation of potential impacts on biodiversity of the salt-tolerant transgenic Eucalyptus camaldulensis harboring an RNA chaperonic RNA-Binding-Protein gene derived from common ice plant
Transgenic Research (2021)
-
Efficient base editing in tomato using a highly expressed transient system
Plant Cell Reports (2021)
-
Development of acetosyringone-inducible Gateway® and Golden Gate expression vectors for heterologous gene expression in Agrobacterium tumefaciens
In Vitro Cellular & Developmental Biology - Plant (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.