Abstract
Recessive mutations in the Mlo gene confer broad spectrum resistance in barley (Hordeum vulgare) to powdery mildew (Blumeria graminis f. sp. hordei), a widespread and damaging disease. However, all alleles discovered to date also display deleterious pleiotropic effects, including the naturally occurring mlo-11 mutant which is widely deployed in Europe. Recessive resistance was discovered in Eth295, an Ethiopian landrace, which was developmentally controlled and quantitative without spontaneous cell wall appositions or extensive necrosis and loss of photosynthetic tissue. This resistance is determined by two copies of the mlo-11 repeat units, that occur upstream to the wild-type Mlo gene, compared to 11–12 in commonly grown cultivars and was designated mlo-11 (cnv2). mlo-11 repeat unit copy number-dependent DNA methylation corresponded with cytological and macroscopic phenotypic differences between copy number variants. Sequence data indicated mlo-11 (cnv2) formed via recombination between progenitor mlo-11 repeat units and the 3′ end of an adjacent stowaway MITE containing region. mlo-11 (cnv2) is the only example of a moderated mlo variant discovered to date and may have arisen by natural selection against the deleterious effects of the progenitor mlo-11 repeat unit configuration.
Similar content being viewed by others
Introduction
Powdery mildews are a group of fungal diseases caused by obligate biotrophic species (species that derive nutrients from living tissue). Their distribution is world-wide and they infect most staple crops. The disease is characterised by white epidermal colonies composed of mycelia and spore chains. Powdery mildews stunt plant growth and hence reduce yield, while necrotic and chlorotic discoloration of surface tissue downgrades crop quality. In temperate regions barley powdery mildew, caused by Erysiphe graminis f. sp. hordei (Bgh), is a significant pathogen1.
In agriculture, the commonest form of resistance in plants to biotrophic fungi follows the race-specific gene-for-gene principle first demonstrated by Flor2, where recognition is governed by the direct or indirect interaction between the product of a host disease resistance gene (R-gene) and the complementary product of a pathogen avirulence gene3. While such R-genes are easy to select for, they are easily overcome by the pathogen as avirulence genes readily mutate, resulting in lack of recognition by a plant genotype carrying the corresponding R-gene. Successive rounds of wide scale monocultures leads to isolates harbouring large numbers of virulence genes, with virulence to almost all R-genes deployed in Europe present in just two isolates4.
Advances in molecular genetic techniques have renewed interest in identifying alternatives to major R-genes to improve crop productivity. Among these are broad-spectrum partial resistance genes. Partial resistance may be developmentally apparent or present in all growth stages and is characterised by reduced numbers of successful infection sites, slower disease progression and fewer successful fruiting bodies. Partial resistance is common in plants but often absent in modern barley cultivars, where genetic bottlenecks and breeding between elite cultivars has led to extinction of diversity in some genomic regions and increased linkage disequilibrium5,6. Combining partial resistance genes phenotypically is challenging because of their quantitative nature but improvements in genetic marker technologies has allowed key barley QTL to be identified4,7,8.
Broad-spectrum resistance to powdery mildew in plants also occurs as single, naturally occurring genes with a major effect. Only two are currently known: RPW8, identified in Arabidopsis conferring resistance to diverse powdery mildew species9,10. The second involves Mlo, a widely conserved plant gene that negatively regulates plant defence responses and might be regarded as a powdery mildew susceptibility gene11,12. Recessive mutations in Mlo confer durable resistance to all races of Bgh. Such mutations were originally discovered in barley as artificial mutants from the 1940’s onwards and subsequently as a spontaneously occurring mutant in accessions collected in Ethiopia in the late 1930’s, designated mlo-1113. mlo alleles act early during infection and are characterised by rapid formation of large cell wall appositions (CWA) that correlate with resistance in epidermal cells, although a direct connection has not been demonstrated14.
mlo mutant based resistance has proved robust since first incorporated into European spring barley cultivars over thirty years ago13 and in Europe the naturally occurring mlo-11 domain is the most widely used form of this resistance. mlo-11 possesses a normal Mlo gene, with resistance conditioned by an upstream tandem repeat array15,16. This repeat array consists of copies of 5′ Mlo regulatory sequence and the first five exons of the Mlo gene, with the structure suggestive of ‘rolling-circle’- DNA replication used by plant viruses and transposons. Piffanelli et al.15 proposed a resistance mechanism involving aberrant transcriptional read-through from the upstream repeats supressing Mlo transcription.
Despite widespread deployment, mlo mutants are not without pleiotropic effects. Both induced mutants and mlo-11 exhibit spontaneous necrosis that share a common QTL co-localising with mlo17. Excessive cell death in the absence of any pathogen leads to necrotic leaf spotting, associated loss of photosynthetic area and a reduction in yield18,19. Breeding efforts in Europe have combined mlo with genes that compensate for these effects but it has been implicated in increased susceptibility to facultative diseases such as spot blotch20 and Ramularia leaf spot21, although in the latter disease progression may be affected by environmental conditions21,22.
In the process of screening for new sources of resistance to powdery mildew, we uncovered a new variant of mlo-11. This variant, designated mlo-11 (cnv2), exhibits partial resistance to powdery mildew in seedling leaves and effective resistance in adult leaves, without the pleiotropic effects of existing mlo alleles. In this study we investigate the phenotypic, structural, gene expression and epigenetic differences of the variant compared to cultivars possessing the wild-type Mlo gene or the standard mlo-11 domain.
Results
Macroscopic Eth295 powdery mildew symptoms
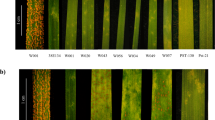
In experiments using single Australian Bgh isolates, Art-001, Wag-001 and Will-005, landrace Eth295 developed fewer colonies in detached leaves which progressed more slowly than the susceptible control cultivar (cv.) Baudin, with an infection type (IT) 2–3 at 7 dpi (see Fig. 1 and Supplementary Fig. 1 for isolate Wag-001) based on the scale of Kølster et al.23. A standard mlo-11 control cultivar, Westminster, was resistant and showed water soaked lesions, suggesting underlying or mesophyll cell death, with occasional colonies stemming from infection through stomatal cells. Detached seedling leaf inoculations gave an average ratio of Baudin to Eth295 colonies of 6.5, with a large sample standard deviation across 3 isolates, s = 4.30. In seedling tests at IPK Gatersleben six isolates produced no symptoms and two (CH4.8 and D4/6) very few colonies (data not shown) and therefore further phenotypic characterisation of adult leaves was not conducted. In detached fifth leaf inoculations, Eth295 showed a consistently more resistant phenotype than Baudin by a factor of 11.28, s = 0.31.
Macroscopic detached leaf Bgh powdery mildew colony counts.
The graph depicts colonies per cm2 for barley cv. Baudin, landrace Eth295 and cv. Westminster at 7 dpi for seedling and fifth barley leaves inoculated with Bgh isolate Wag-001. Error bars are standard errors for five biological replicates per experiment with each experiment repeated three times. Significant differences were determined using the Student’s t-test, ***P < 0.001.
Whole plant Eth295 assays inoculated at the 5th leaf stage showed fewer colonies compared to detached leaves. For example, at 14 dpi only three colonies were observed on eight Eth295 plants, while Baudin controls showed abundant sporulation, IT 4, with >50% of the leaf area infected (Fig. 2A).
Bgh disease symptoms on barley leaves inoculated with Bgh isolate Wag-001.
Figure (A) Whole plant macroscopic symptoms at 14 dpi on (from top to bottom) cv. Baudin, landrace Eth295 and, cv. Westminster. Figure (B–F) Microscopic Bgh disease symptoms on detached barley fifth leaves at 48 hpi inoculated with isolate Wag-001. B) Eth295 depicting limited secondary hyphal growth. (C) Eth295 illustrating a secondary cell wall apposition (CWA), haustorium (Hau) and conidium (C). (D) Baudin with rapidly developing hyphae. (E) Germinating conidium on Westminster with secondary and tertiary appressorial lobes. (F) Westminster, showing CWA restricted to epidermal cells immediately below a germinating conidium. Figure (G–I) Microscopic mesophyll cell death on uninfected detached barley fifth leaves. (G) Westminster stained with DAB showing the accumulation of hydrogen peroxide in numerous mesophyll cells. (H) Spontaneous CWA in cv. Westminster not observed in landrace Eth295. (I) Eth295 depicting hydrogen peroxide restricted to a few cells, in this example two cells. Samples (B,C,E) were triple stained with Evan’s blue, aniline blue and calciflour white. Samples D and F-I were DAB stained. Scale bars indicate 200 μm.
Cytological features of Eth295
Microscopic examination of 5th leaf 48 hpi showed unsuccessful spore penetration rates of the epidermal cell wall between cv. Westminster, landrace Eth295 and cv. Baudin were ≥99%, 85–90% and 25–30%, respectively. In Eth295, successful penetration was accompanied by secondary hyphae elongation (Fig. 2B,C). In general landrace Eth295 showed no auto-fluorescence beneath epidermal cells indicating a lack of mesophyll cell death compared to Westminster, while growth was limited compared to Baudin (Fig. 2D). In Westminster, fungal germlings produced an appressorial germ tube and attempted penetration from an initial appressorial lobe failed to penetrate the epidermal call, leading to second and third unsuccessful lobes (Fig. 2E) which are a feature of mlo based resistance24.
DAB staining of leaves 48 hpi was performed to detect CWA (rounded protuberances) and accumulation of hydrogen peroxide (H2O2), often a precursor of cell death. No epidermal accumulation of H2O2 was evident in Eth295 at attempted penetration sites, a feature shared with standard mlo-11 cultivars (Westminster, Grange and Henley). Where penetration was successful, Eth295 showed secondary CWA in surrounding cells (Fig. 2C). Mesophyll accumulation of H2O2 was rarely observed, typically involving 2–8 superficial mesophyll cells. In Westminster, areas of mesophyll accumulation of H2O2 were large and more numerous than Eth295, typically involving 50 or more cells when viewed from above, while trypan blue staining confirmed extensive cell death throughout the mesophyll layer (Supplementary Fig. 1E,F).
Uninfected 5th leaf segments of cv. Westminster mlo-11 also exhibited areas of accumulation of H2O2 (in the range of 10–30 cells, Fig. 2G) while spontaneous papillae were also observed in Westminster and not in Eth295 (Fig. 2H). In Eth295, accumulation was rarely observed and confined to 2–4 superficial mesophyll cells (Fig. 2H). Cell death sites occurred on average at 31 sites per cm2 in Westminster compared to 4 sites for Eth295 (Supplementary Fig. 1G).
Involvement of mlo in Eth295 resistance
Landrace Eth295 was crossed with cv. Baudin to investigate the genetics of Bgh resistance. All F1 progeny were fully susceptible to Bgh. One hundred and seventeen F2 lines were inoculated to determine the resistance segregation. Twenty seven individuals were resistant and 91 susceptible, indicating a 1:3 ratio. Chi squared was 2.67, p > 0.01, for the null hypothesis of no difference between the observed and expected segregation controlled by a single recessive locus. No F2 individuals showed disease symptoms intermediate to the parental phenotypes.
The best known recessive resistance in barley is mlo conditioned and therefore this locus was studied to establish any contribution to the Eth295 phenotype. Transient complementation with overexpressed Mlo was carried out by bombarding pUbi_Mlo_nos into epidermal leaf cells, followed by inoculation with Bgh. Both Eth295 and the control line cv. Ingrid BC mlo5 showed an increased susceptibility index by a factor >5 (Fig. 3), which is indicative of mlo mediated resistance4. A slight increase in susceptibility was observed in the negative control cv. Golden Promise. This corresponds to the average effect found in a population of non-mlo accessions4 and may be regarded as a marginal gain in susceptibility from over-expressing Mlo.
Biolistic complementation of mlo alleles with a construct overexpressing the Mlo gene.
The graph depicts epidermal cell susceptibility indices for barley lines bombarded with pUbiGUS and pUbi_Mlo_nos. Control samples were bombarded with pUbiGUS only. Haustoria were counted 48 hr after inoculation with Swiss field isolate CH4.8 avrMla9 by light microscopy. The graph compares results from barley landrace Eth295 with control genotypes cv. Ingrid mlo5, cv. Roland Mla9 and cv. Golden Promise 2012 Mlo. The susceptibility index was calculated as (ΣGUS-stained epidermal cells containing at least one haustorium/ΣGUS-stained epidermal cells). The complementation factor (CF, the ratio of the means of total haustoria in pUbi_Mlo_nos bombarded samples to the control samples) is given for each treatment pair. Error bars are standard errors based on using seven leaf segments per complementation experiment and each experiment was replicated a minimum of four times. Significant differences were determined using the Student’s t-test; ***P < 0.0005 and *P < 0.05.
As mlo-11 is a known naturally occurring form of mlo, we conducted initial PCR experiments using two diagnostic genetic markers designed against the mlo-11 domain15. These were inconclusive: The first primer pair amplifies a stowaway MITE located immediately 5′ to the mlo-11 repeats. Eth295 exhibited a short wild-type amplicon of 381 bp, rather than the 441 bp in cv. Westminster (Supplementary Fig. 2). The second, targeting the mlo-11 repeat units, showed weak and inconsistent amplification. The forward primer used to amplify this repeat, ADUP7, possesses a homopolymeric 3′ end. Non-specific amplification was excluded by amplicon sequencing which indicated at least one copy of the mlo-11 repeat unit was present.
To resolve the number of mlo-11 repeat units in landrace Eth295, new primers were developed that targeted Mlo exon 5 and digital PCR conducted to determine copy number variation (CNV). Digital PCR is a highly sensitive method for absolute quantification of nucleic acids without the need for standard curves. The PCR sample is partitioned into thousands of droplets, with each droplet containing a target sequence detected by fluorescence. Digital PCR is more accurate than conventional real-time qPCR which relies on relative quantification, while variations in amplification efficiency and signal saturation may affect results. The results indicated Eth295 contained two repeat units and the negative control Baudin no repeat units. Westminster displayed a high copy number. Despite 32 separate measurements, the number was only reliably estimated at 11–12 repeats (average = 11.92, s = 0.44).
To test whether the mlo-11 repeat region and the Mlo locus was intact in Eth295, genetic linkage between a MITE SNP (Supplementary Fig. 2) and the resistance locus was examined. Only one recombinant was detected in 117 F2 progeny. To confirm mlo-11 repeat subunit mediated resistance is cis-regulatory in Eth295, rather than a result of excision and reinsertion at a different site, PCR targeting read-through transcript from the mlo-11 repeat units to Mlo exon 6 was positive (image not shown). No microsatellites 3′ to Mlo (HVMLOH1A, GBM1448, WMS6, HVM67, GBM1388 and GBM1324) were polymorphic between Eth295 and Westminster, except Bmag0138b, which lies >25 cM from the Mlo locus19,25.
Intermediate levels of Mlo and mlo-11 aberrant transcript expression in landrace Eth295
Expression data revealed landrace Eth295 expressed more than double the amount of aberrant transcripts at first and fifth leaf stages compared to cv. Westminster (Supplementary Fig. 4). Expression levels in cv. Baudin were zero; consistent with this cultivar lacking mlo-11 repeats. Total Mlo transcripts were over three times higher in Eth295 than Westminster in seedling leaves and five times higher in fifth leaves. However, Westminster total Mlo transcripts were composed of entirely of aberrant transcript, with little or no normal Mlo transcription.
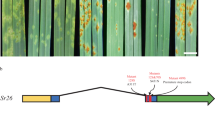
At the first leaf stage, Eth295 expression of normal Mlo transcript was less than half the level of Baudin, once aberrant mlo-11 repeat transcripts are taken into account. At fifth leaf stage, Eth 295 expressed one fifth of the amount of Mlo compared to Baudin. These relative amounts are consistent with lower colony counts at the fifth leaf stage in Eth295. Figure 4 depicts the organisation and relative expression levels of Eth295, Westminster and Baudin Mlo and mlo-11 repeat unit loci.
Schematic representations of domain organisation and transcript expression levels in Mlo-11 variants.
(A) Landrace Eth295. (B) Cultivar Westminster. (C) Wild type Mlo gene in Bgh susceptible cv. Baudin for comparison. Grey bar widths indicate relative expression levels of aberrant transcripts between the truncated mlo-11 repeats and Mlo and of Mlo itself (see Supplementary Fig. 4). Yellow rectangles represent the promoter region of Mlo and green rectangles represent merged exons. The mlo-11 arrangement in B is based on Peterhansel et al.16. DNA regions not drawn to scale. The line break indicates the exact distance between the 3′-proximal repeat unit and the Mlo wild-type gene copy is not resolved.
Expression of Mlo is DNA methylation dependent
DNA methylation levels of both the Mlo promoter and 5′ UTR regions were studied to examine methylation dependency. The restriction enzyme McrBC was used as a sensitive methylation detection method, rather than conventional methylation-dependent endonucleases. McrBC detects a high proportion of methyl cytosines by recognition of two short half sites with the consensus sequence (G/A)mC and, in addition, restricts asymmetric and hemi-methylated (single stranded) sites26. Digestion with McrBC followed by qPCR showed that cv. Westminster (together with cvs. Grange and Henley, data not shown) exhibited low qPCR amplification levels compared to Eth295 and non-mlo cv. Baudin at both read-through promoter sequences and 5′ UTR regions, indicating these regions are highly methylated. No evidence for methylation at the control actin promoter was found. Westminster methylation was strongest at the Mlo promoter (Fig. 5), resulting in negligible amplification (∆∆Cq = 0.008, s = 0.003). Landrace Eth295 displayed methylation levels intermediate between Westminster and Baudin, consistent with transcript levels at the 5′ UTR and the intron-spanning section of Mlo exons 6 and 7. Bisulphite sequencing is a process that involves treatment of genomic DNA to convert cytosine residues to uracil, but leaves 5-methylcytosine residues unaffected. Bisulphite sequencing of the promoter region also showed a higher level of DNA methylation and total number of methylated cytosine bases in Westminster, a low level in Baudin and an intermediate level in Eth295 for all the CG, CHG and CHH contexts (where H = A, T, or C, Fig. 5).
DNA methylation status of Mlo in landrace Eth295, cv. Baudin and cv. Westminster.
(A) DNA methylation status was determined by digestion with McrBC followed by quantitative PCR at the promoter and UTR region of Mlo gene. Undigested genomic DNA was used as a control. Four biological replicates were used for each accession and real time PCR was performed with two McrBC digest replicates and four qPCR technical replicates. Error bars are standard errors and significant differences are determined using the Student’s t-test; ***P < 0.001, **P < 0.01 and *P < 0.05. (B) Methylation status of cytosine residues determined by bisulphite sequencing of a section of the promoter region of Mlo (−440 to −390 bp from the start codon). The percentage of mC in three contexts (CG, CHG and CHH) was calculated by sequencing of eighteen clones from each accession.
Discussion
In this study we demonstrated Ethiopian landrace Eth295 possesses effective broad spectrum resistance to Bgh in adult leaves. Subsequent genetic analysis and complementation with overexpressed Mlo implicated a recessive mlo allele as underlying the resistance. Unlike known mlo alleles, the Eth295 variant does not show the pleiotropic effects of spontaneous necrosis and loss of photosynthetic tissue.
The resistance in seedling Eth295 leaves is quantitative in both the number of colonies and in the extent of colony growth. In detached leaf experiments, leaves exhibited proportionally higher numbers of powdery mildew colonies than adult fifth leaves. This is consistent with developmental control of spontaneous mlo CWA described by Wolter et al.27. However, Eth295 does show higher levels of infection than standard mlo cultivars, where colony development is only occasionally seen as a result of infection of stomatal subsidiary cells28 or as a result of drought-relief stress related to genetic background29. Notably, Eth295 is more resistant in intact (whole plant) assays than in severed leaves inoculated in-vitro. As mlo resistance is associated with physiological processes that prevent successful appressorial penetration of host epidermal cells, the condition of tissues in detached leaves may affect the timing and success of the resistance response.
Cytological studies showed Eth295 exhibits penetration resistance in common with standard mlo-11, represented in this study by the cultivar Westminster, with CWA observed below attempted penetration sites. Principal differences were presence in Eth295 of CWA in adjacent epidermal cells where penetration was successful at the initial site. Extensive necrosis of surrounding cells and the collapse of underlying mesophyll cells were not observed in Eth295. Similarly, in uninfected leaves, Eth295 exhibited no spontaneous CWA compared to Westminster and limited spontaneous necrosis, typically restricted to two to four cells. Partial seedling resistance similar to Eth295 has been previously observed in chemically induced mlo-12 & mlo-28 mutants of Mlo wild-type plants30. Infected leaves indicated this resistance was accompanied by lower levels of mesophyll cell death than other mlo alleles. However, moderate cell death was still evident and contrasts with Eth295. The phenotype of Eth295 supports the hypothesis of Buschges et al.12 that the Mlo protein negatively regulates multiple defence-related processes with the severity of the phenotype dependent on the strength of the mutation in inhibiting Mlo protein formation.
We examined the structure and gene expression levels of the mlo-11 subunit domains and the Mlo gene itself in Eth295 and cv. Westminster to determine whether any variances might explain their phenotypic differences. Digital PCR was conducted to determine copy number variation (CNV), rather than qPCR as imperfect amplification efficiencies and signal saturation limits CNV measurement in samples with high copy numbers. This revealed a six-fold difference between Eth295 and Westminster, with 2 copies in Eth295 and 11–12 in Westminster. The copy number in Westminster could not be more precisely fixed as such CNV counts represent the limit of whole number precision for this technique31.
Eth295 contains a distinctive short MITE amplicon compared to Westminster and other mlo-11 lines but no mlo-11 repeat unit SNPs were detected between Eth295 and cv. Westminster. Together with a lack of polymorphic microsatellites immediately downstream from Mlo, a recent shared ancestry rather than an independent origin appears likely. Eth295 contains a SNP shared with Westminster at the 3′ end of the MITE containing amplicon. The region immediately 5′ to this SNP does not align between short and long versions of the amplicon due to an insertion consisting of contiguous sections of Mlo exon 5 and intron 5 (Supplementary Fig. 2). The shared SNP and short MITE containing amplicon in Eth295 may be explained by unequal crossing over and gene conversion between progenitor repeat units or recombination between these repeat units and the wild-type Mlo gene during meiosis, followed by a second recombination event with a wild type Mlo line MITE region. However, the most parsimonious explanation for the Eth295 arrangement is direct recombination just before the shared SNP with a homologous 5′ promoter region present between the mlo-11 repeat units.
In this study, relative Mlo expression levels between Eth295 and cv. Westminster corresponded to phenotypic differences. Piffanelli et al.15 were unable to detect an involvement of DNA methylation in gene expression and suggested read-through transcripts from mlo-11 repeats, which lack a transcription termination signal, interfere with transcription from the downstream native Mlo promoter. However, in Eth295 mlo-11 read-through transcripts and Mlo itself are expressed at twice the levels found in Westminster, which showed no Mlo gene expression. Disruption of transcription machinery assembly by such read-through is therefore unlikely to be a major mechanism controlling Mlo expression in mlo-11.
Methylation of repetitive DNA is a common plant defence mechanism against viruses and transposons32. The involvement of methylation in the expression of the Mlo gene conditioned by different mlo-11 subunit domains was shown by qPCR following restriction with McrBC and direct detection of methylation at the Mlo promoter by bisulphite sequencing. These corresponded to intermediate Mlo qPCR transcript levels in landrace Eth295 and no Mlo expression in cv. Westminster, indicating a direct relationship with mlo-11 repeat unit copy number. As repetitive DNA is associated with the formation of double stranded RNA leading to RNA-directed methylation via small interfering RNAs, future analyses may elucidate the role of that process and of different histone modification pathways in mlo-11 epigenetic regulation33,34,35.
Landrace Eth295 is a selection of HOR 2543 (Hordeum vulgare L. convar. deficiens var. nudideficiens Körn.), collected by the ‘Zentralinstitut für Genetik und Kulturpflanzenforschung Gatersleben’ in Germany. No collection information is available in public databases for Eth295, however it is plausible to speculate this landrace may have arisen by natural selection against the pleiotropic effects of standard mlo-11. The existence of mlo-11 (cnv2) suggests rare mlo-11 repeat copy number recombinants offer breeders an opportunity to fine tune the balance between copy number, pleiotropic effects and resistance levels, allowing the incorporation of such alleles into cultivars without selecting genetic backgrounds that compensate for the pleiotropic effects.
Methods
Bgh maintenance and plant growth
Mono-conidial Australian Bgh cultures from Australian field isolates were produced and pathotyped as described by Tucker et al.36. Powdery mildew isolates were maintained and propagated on detached leaf sections of cv. Baudin inserted into 50 mgL−1 benzimidazole agar plates and grown in a Contherm Biosym 6200CP4 incubator (16 °C and 10 °C under a 12 h light and dark cycle, respectively). Isolates were sub-cultured at 10 days or before leaves senesced.
Barley plants for seedling leaf detached leaf infection assays were grown in vermiculite fertilised with Nitrophoska Perfect (EuroChem Antwerpen NV, Belgium) and under light shelves at 300 μmol m−2 s–1. Plants for detached 5th leaf or whole plant infection assays were grown in soil with Nitrophoska Perfect in a controlled temperature room (18–22 °C) and a 12 h photoperiod at 450 μmol m−2 s–1.
Detached leaf and whole plant disease assays
For all experiments, a minimum of five biological replicates were used and the experiments repeated three times. Landrace Eth295 was obtained from the Australian Grains Genebank (Horsham, VIC) and was identified as having prominent adult plant-like resistance in primary disease screens against Australian isolates. Cultivar Westminster was used as a standard mlo-11 control with cvs. Grange and Henley as additional comparative replicates (Limagrain-Nickerson, UK). Cultivar Baudin was used as a positive control. All experiments in Western Australia were conducted with isolate Wag-001. Two different pathotypes, Art-001 and Will-005 were used in comparative replicate experiments. Isolates CH4.8, RiIII, Ro93a, MH21, D35/3, D35/2, D4/6, D2/4 were used for inoculations at IPK Gatersleben. Detached leaves were inoculated as described by Tucker et al.36. Whole plant inoculations at the fifth leaf stage were performed by placing plants into trays enclosed in 80 cm high polythene tents and spores allowed to settle from above the tents. Inoculated plants were sealed and kept within the polythene tents for 24 hr then placed in a controlled temperature glasshouse with a maximum temperature 22–24 °C.
Inoculated detached leaves were scored macroscopically at 7 days post inoculation (dpi), unless otherwise indicated and rescored at 10 dpi to confirm more resistant disease reactions. A five point scale was used based on that devised based on Mains et al.37 and Kølster et al.23, where 0 = no visible mycelium, 1 = sparse mycelial development with no sporulation, 2 = mycelial present with very few spore chains, 3 = moderate mycelial development, discrete lesions with sporulation and 4 = amorphous mycelial development and abundant sporulation. Colony counts were obtained from 2 cm wide transects across detached leaf plates.
Cytology
Bgh infected leaves were sampled at 48 hpi by taking 0.5 cm2 leaf segments from 10 independent leaves per genotype. A modified 3,3-diaminobenzidine (DAB) uptake was carried out as described by Thordal-Christensen et al.38. Triple staining with Evans blue, aniline blue and calcofluor white was performed following the protocol developed by Felix Mauch’s Group at the University of Fribourg39. Trypan blue retention on 0.3 mm wide mesophyll cross sections was performed as described by van Wees40 using Farmer’s fluid (acetic acid: ethanol: chloroform at 1:6:3) rather than chloral hydrate solution to de-stain the tissue. DAB staining of mesophyll cells was performed on epidermal strips as described by Thordal-Christensen et al.38.
Genetic analysis
Dominance of the Eth295 resistance phenotype was determined among five F1 Baudin x Eth295 progeny by inoculating detached 1st and 5th leaves. Whole plant F2 progeny were inoculated at the 5th leaf. To confirm the location of the resistance locus (also see mlo complementation below) F2 genotyping was performed with the nearest Mlo genetic marker. This was a unique Eth295/Baudin T/C SNP within the MITE at position 7659 relative to GenBank accession Y14573 (Supplementary Fig. 2) amplified by primers Mlo6 and Mlo10 described by Piffanelli et al.15. This polymorphism served as a CAPS marker for PCR products following restriction with HinfI and resolution on a 2.5% agarose gel.
mlo complementation
Leaf segments of accession Eth295 were co-bombarded with a mixture of GUS- reporter plasmid pUbiGUS41 and pUbi_Mlo_nos carrying wild type Mlo12 as described by Spies et al.4. Ingrid BC mlo5 was used as a positive mlo control; cvs. Golden Promise and Roland Mla9 were used as negative Mlo controls. Each accession was bombarded with or without pUbi_Mlo_nos, using 7 leaf segments per treatment and each experiment replicated a minimum of four times. Four hours after bombardment leaf segments were inoculated with Bgh isolate CH4.8 avrMla9 at a density of approx. 150 conidia/mm2 and haustoria in GUS stained (transformed) epidermal cells counted 48 h after inoculation. The susceptibility index (SI) was calculated as (ΣGUS-stained epidermal cells containing at least one haustorium/ΣGUS-stained epidermal cells).
Eth295 mlo-11 locus structure
The mlo-11 locus was initially examined by PCR with previously published MITE and mlo-11 repeat primer pairs, Mlo6-Mlo10 and ADUP7-Mlo6, respectively15. PCR products were run out on a 1.5% agarose gel and Sanger sequenced to identify nucleotide differences. The following microsatellites were tested for polymorphism between Eth295 and Westminster 3′ to the Mlo gene: HVMLOH1A, GBM1448, WMS6, HVM67, GBM1388, GBM1324 and Bmag0138b19,25.
Copy number variation (CNV) of landrace Eth295 was determined with QuantStudio™ 3D Digital PCR System (Life Technologies, Carlsbad, CA, USA). The mlo-11 cultivar Westminster was used as the standard mlo-11 domain control and cv. Baudin as a wild-type Mlo control. Cultivars Grange and Henley provided additional standard mlo-11 domain controls. Sample DNA concentrations were measured with a Quibit 2.0 Fluorometer (Life Technologies). Primers MloEx5_F and MloEx5_R were designed to Mlo exon 5, together with a VIC Taqman probe MloEx5T. These primers detect mlo-11 subunits together with the WT Mlo gene. Reference single copy actin primers (designed from GenBank accessions AY145451 and CAJX010010156) were Actin F and Actin R, combined with a FAM labelled Taqman probe. DNA restriction with MseI was used to cleave linked copies of the target exon for dilution into separate PCR partitions. Digital PCR DNA partition, PCR amplification, chip reading and data analyses were performed according to manufacturer’s recommendations. For each accession, four biological replicates were used, with two MseI digests per biological replicate, followed by four technical replicates per MseI digest. To confirm read-through transcription between Mlo-11 and Mlo, forward primer MloPro_F2 and reverse primer MloEx6_R were used in conventional PCR. These primers amplify a 1650 bp DNA region -227 bp relative to the Mlo transcriptional start site through to Mlo Exon 6. All primer sequences used in this study are provided in Supplementary Fig. 3 and illustrated with Geneious v. R842.
Gene expression
Four biological replicates of leaf tissue were sampled from leaves of each accession at the 1st and 5th leaf stages. Total RNA was extracted using TRIzol® Reagent (Life Technologies) following the manufacturer’s protocol. cDNA was synthesised from 3 ug total RNA using a RNA QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany). Quantitative real-time PCR (qPCR) was performed on a C1000TM Thermal Cycler (Bio-Rad, Hercules, CA, USA) with SYBR green detection (QuantiTect SYBR Green PCR Kit) with three technical replicates. Primers were developed to target mlo-11 aberrant transcripts only by amplifying the mlo-11 repeat region read-through promoter sequence with the primers MloPro_F3 and MloPro_R1. These amplify a 138 bp section upstream of the Mlo 5′ UTR. A second set of primers amplified total Mlo (Mlo and mlo-11 read-through) transcripts from an intron-spanning section of Mlo exons 6 and 7, as the mlo-11 repeat units are composed of Mlo exons one to five15. These transcripts were detected with the primers MloEx6_F and MloEx7_R, which amplify a 100 bp region. Internal reference primers used were Actin F and R, as described for CNV above. Relative transcript levels were calculated by the ∆∆Ct method factoring in primer efficiencies.
Cytosine methylation by McrBC-quantitative PCR
Methylation of the promoter region of Mlo and the mlo-11 repeat units were detected by McrBC (NEB, Ipswich, MA) digestion followed by qRT-PCR. McrBC detects a high proportion of methyl cytosines by recognition of two short half sites with the consensus sequence (G/A)mC and, in addition, restricts asymmetric and hemi-methylated (single stranded) sites26. For each accession, two 500 ng aliquots of DNA from four biological replicates were treated with McrBC. Undigested controls were treated with water. qPCR was performed with four technical replicates per sample and DNA methylation levels quantitatively calculated by the ∆∆Ct method. Detection primers for Mlo and mlo-11 5′ UTR were forward primer MloUTR_F and reverse primer MloUTR_R. These primers amplify a 234 bp DNA region. Primers for the promoter region before the 5′ UTR were forward primer MloPro_F1 and reverse primer MloPro_R1. These primers amplify a 311 bp DNA region. Primers for the actin promoter region were forward primer ActPro_F and reverse primer ActPro_R, which amplify a 297 bp DNA region. As the Mlo promoter and 5′ UTR sequences are also identical in all mlo-11 repeat units, the qPCR results represent the overall (Mlo and mlo-11) methylation status.
Cytosine methylation bisulphite sequencing analysis
Bisulphite sequencing was performed based on published protocols34,35 with genomic DNA extracted from landrace Eth295, cv. Baudin and cv. Westminster. 200 ng of each accession was subjected to bisulphite CT conversion using an EZ DNA Methylation-Direct kit (Zymo Research, Irvine, CA). Bisulphite converted DNA was amplified by PCR using primers BiSu_F and Bisu_R. Untreated DNA was used for control reactions. Amplified fragments were cloned into the pGEM-T vector (Promega, Madison, WI) and eighteen ligated clones from each accession sequenced.
Additional Information
How to cite this article: Ge, X. et al. Tempered mlo broad-spectrum resistance to barley powdery mildew in an Ethiopian landrace. Sci. Rep. 6, 29558; doi: 10.1038/srep29558 (2016).
References
Braun, U. & Cook, R. T. A. Taxonomic manual of Erysiphales (powdery mildews). 1–707 (CBS Biodiversity Series, Utrecht, 2012).
Flor, H. H. Current status of the gene-for-gene concept. Annu. Rev. Phytopathol. 9, 275–296 (1971).
Staskawicz, B. J., Ausubel, F. M., Baker, B. J., Ellis, J. G. & Jones, J. D. Molecular genetics of plant disease resistance. Science 268, 661–667 (1995).
Spies, A. et al. Allele mining in barley genetic resources reveals genes of race-non-specific powdery mildew resistance. Front. Plant Sci. 2, 1–22 (2012).
Tondelli, A. et al. Structural and temporal variation in genetic diversity of European spring two-row barley cultivars and association mapping of quantitative traits. Plant Genome 6 (2013).
Hamblin, M. T. et al. Population structure and linkage disequilibrium in U.S. barley germplasm: Implications for association mapping. Crop Sci. 50, 556–566 (2010).
Aghnoum, R. et al. Basal host resistance of barley to powdery mildew: connecting quantitative trait loci and candidate genes. Mol. Plant Microbe Interact. 23, 91–102 (2010).
Hickey, L. T. et al. Mapping quantitative trait loci for partial resistance to powdery mildew in an Australian barley population. Crop Sci. 52, 1021–1032 (2012).
Xiao, S. et al. Broad-spectrum mildew resistance in Arabidopsis thaliana mediated by RPW8. Science 291, 118–120 (2001).
Wang, W., Wen, Y., Berkey, R. & Xiao, S. Specific targeting of the Arabidopsis resistance protein RPW8.2 to the interfacial membrane encasing the fungal haustorium renders broad-spectrum resistance to powdery mildew. Plant Cell 21, 2898–2913 (2009).
Acevedo-Garcia, J., Kusch, S. & Panstruga, R. Magical mystery tour: MLO proteins in plant immunity and beyond. New Phytol. 204, 273–281 (2014).
Büschges, R. et al. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell 88, 695–705 (1997).
Jørgensen, I. H. Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63, 141–152 (1992).
Skou, J. P., Jørgensen, J. H. & Lilholt, U. Comparative studies on callose formation in powdery mildew compatible and incompatible barley. J. Phytopathol. 109, 147–168 (1984).
Piffanelli, P. et al. A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430, 887–891 (2004).
Peterhänsel, C. & Lahaye, T. Be fruitful and multiply: gene amplification inducing pathogen resistance. Trends Plant Sci. 10, 257–260 (2006).
Behn, A., Hartl, L., Schweizer, G. & Baumer, M. Molecular mapping of QTLs for non-parasitic leaf spot resistance and comparison of half-sib DH populations in spring barley. Euphytica 141, 291–299 (2005).
Kjær, B., Jensen, H. P., Jensen, J. & Jørgensen, J. H. Associations between three ml-o powdery mildew resistance genes and agronomic traits in barley. Euphytica 46, 185–193 (1990).
Thomas, W. T. B. et al. Identification of a QTL decreasing yield in barley linked to Mlo powdery mildew resistance. Mol. Breed. 4, 381–393 (1998).
Kumar, J., Hückelhoven, R., Beckhove, U., Nagarajan, S. & Kogel, K.-H. A compromised Mlo pathway affects the response of barley to the necrotrophic fungus Bipolaris sorokiniana (teleomorph: Cochliobolus sativus) and its toxins. Phytopathology 91, 127–133 (2001).
McGrann, G. R. D. et al. A trade off between mlo resistance to powdery mildew and increased susceptibility of barley to a newly important disease, Ramularia leaf spot. J. Exp. Bot. 65, 1025–1037 (2014).
Makepeace, J. C. et al. Associations between fungal and abiotic leaf spotting and the presence of mlo alleles in barley. Plant Pathol. 56, 934–942 (2007).
Kølster, P., Munk, L., Stølen, O. & Løhde, J. Near-Isogenic barley lines with genes for resistance to powdery mildew. Crop Sci. 903–907 (1986).
Zeyen, R. J. et al. Inhibiting phenylalanine ammonia lyase and cinnamyl-alcohol dehydrogenase suppresses Mla1 (HR) but not mlo5 (non-HR) barley powdery mildew resistances. Physiol. Mol. Plant Pathol. 47, 119–140 (1995).
Varshney, R. K. et al. A high density barley microsatellite consensus map with 775 SSR loci. Theor. Appl. Genet. 114, 1091–1103 (2007).
Pieper, U. et al. The GTP-dependent restriction enzyme McrBC from Escherichia coli forms high-molecular mass complexes with DNA and produces a cleavage pattern with a characteristic 10-Base pair repeat. Biochemistry 41, 5245–5254 (2002).
Wolter, M., Hollricher, K., Salamini, F. & Schulze-Lefert, P. The mlo resistance alleles to powdery mildew infection in barley trigger a developmentally controlled defence mimic phenotype. Mol. Gen. Genet. 239, 122–128 (1993).
Jørgensen, J. H. & Mortensen, K. Primary infection by Erysiphe graminis f. sp. hordei of barley mutants with resistance genes in the ml-o locus. Phytopathology 67, 678–685 (1977).
Baker, S. J. et al. Temporary partial breakdown of mlo-resistance in spring barley by sudden relief of soil water-stress under field conditions: the effects of genetic background and mlo allele. Plant Pathol. 47, 401–410 (1998).
Piffanelli, P. et al. The barley MLO modulator of defense and cell death is responsive to biotic and abiotic stress stimuli. Plant Physiol. 129, 1076–1085 (2002).
Hindson, B. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610 (2011).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Mao, H. et al. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat. Commun. 6 (2015).
Greaves, I. K., Groszmann, M., Wang, A., Peacock, W. J. & Dennis, E. S. Inheritance of trans chromosomal methylation patterns from Arabidopsis F1 hybrids. Proceedings of the National Academy of Sciences 111, 2017–2022 (2014).
Tucker, M. A., Jayasena, K., Ellwood, S. R. & Oliver, R. P. Pathotype variation of barley powdery mildew in Western Australia. Australas. Plant Pathol. 1–7 (2013).
Mains, E. B. & Dietz, S. M. Physiologic forms of barley mildew, Erysiphe graminis hordei Marchal. Phytopathology 20, 229–239 (1930).
Thordal-Christensen, H., Zhang, Z., Wei, Y. & Collinge, D. B. Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J. 11, 1187–1194 (1997).
Mauch, F. Staining of Arabidopsis thaliana leaves with trypan blue or aniline blue. Available at: http://www.unifr.ch/plantbiology/eng/Home/research/mauch-group/protocols/trypan-blue Date of access: 14/06/2016 (2015).
van Wees, S. Phenotypic analysis of Arabidopsis mutants: trypan blue stain for fungi, oomycetes and dead plant cells. Cold Spring Harb. Protoc. pdb.prot4982 (2008).
Schweizer, P., Pokorny, J., Abderhalden, O. & Dudler, R. A transient assay system for the functional assessment of defense-related genes in wheat. Mol. Plant Microbe Interact. 12, 647–654 (1999).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Acknowledgements
This research was funded by the Grains Research & Development Corporation through CUR00017 and CUR00023 P8. The authors acknowledge the excellent technical assistance of Gabi Brantin for transient Mlo complementation experiments and support by Dr Daniela Nowara for Bgh isolate screening of Eth295.
Author information
Authors and Affiliations
Contributions
X.G. and S.R.E. discovered the Eth295 Bgh resistance and performed subsequent Bgh isolate and segregating population phenotyping. X.G. and W.D. conducted the molecular experiments and X.G. microscopy. Z.Z.L. contributed to F2 mapping and SSR identification and F.J.L.-R. to bisulphite sequencing. P.S. managed experiments at IPK Gatersleben. X.G., W.D. and S.R.E. designed experiments. S.R.E. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ge, X., Deng, W., Lee, Z. et al. Tempered mlo broad-spectrum resistance to barley powdery mildew in an Ethiopian landrace. Sci Rep 6, 29558 (2016). https://doi.org/10.1038/srep29558
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29558
This article is cited by
-
Development of mlo-based resistance in tetraploid wheat against wheat powdery mildew
Theoretical and Applied Genetics (2019)
-
Transgenic crops and beyond: how can biotechnology contribute to the sustainable control of plant diseases?
European Journal of Plant Pathology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.