Abstract
Wild wheat species Aegilops peregrina (UpUpSpSp), harbours resistance to various diseases including leaf rust and stripe rust. Inheritance studies in a recombinant inbred line population of wheat-Ae. peregrina introgression line IL pau16061 revealed the transfer of a single major dominant gene conditioning all stage resistance, herein temporarily designated as LrAp. Genomic in situ hybridisation of IL pau16061, resistant and susceptible RILs with U- and S-genome DNA probes confirmed that the introgression with leaf rust resistance is from the Up genome of Ae. peregrina. Fluorescence in situ hybridisation using chromosome specific probes identified Up genome introgression to be on the long arm of wheat chromosome 6B. To genetically map LrAp, bulked segregant analysis was combined with resistance gene enrichment sequencing (MapRenSeq). Five nucleotide binding leucine-rich repeat contigs distinguished resistant and susceptible bulks and single nucleotide polymorphism (SNP) markers from these contigs co-segregated with LrAp. All five RenSeq NB_ARC contigs showed identity with the long arm of wheat chromosome 6B confirming the introgression on 6BL which we propose is a compensating translocation from Ae. peregrina chromosome 6UpL due to homoeology between the alien and wheat chromosomes. The SNP markers developed in this study will aid in cloning and marker assisted gene pyramiding of LrAp.
Similar content being viewed by others
Introduction
Wheat is the prime food commodity of major part of the world’s population with an annual harvest of 733 million tons in 2015. It is a leading source of calories as well as protein both for humans and livestock1. The limited genetic diversity at the farmers’ field increases genetic vulnerability to various biotic and abiotic stresses2. Almost wherever wheat is grown, production is significantly constrained by one or more of the three rust diseases3. These include leaf rust (Puccinia triticina Eriks); stem rust (P. graminis Pers. f. sp. tritici); and stripe rust (P. striiformis Westend f. sp. tritici). Among the three wheat rust pathogens, leaf rust cause less damage as compared to stripe and stem rust, however, due to its recurrent global occurrence, it would be anticipated that total annual losses incurred would be greater than any other rusts4,5. In India, stem rust and stripe rust are geographically constrained, while leaf rust is endemic across all production areas. The disease not only reduce the kernel weight but also lessen number of kernels per head6,7. The frequent emergence of novel pathotypes of P. triticina (Pt) and switching of virulence patterns constitute the main hurdles for its management8,9.
Wild species of wheat, especially members of the tertiary gene pool, carry an immense diversity of disease resistance (R) genes that could enable more sustainable disease control10,11. Till date, 79 genes for leaf rust resistance have been designated and ~44% are from wild progenitor and non-progenitor species12. These genes include Lr21, Lr22a, Lr32, Lr39/Lr41, Lr40, Lr42, Lr43 (Ae. tauschii); Lr28, Lr35, Lr36, Lr47, Lr51, Lr66 (Ae. speltoides); Lr63 (T. monococcum); Lr53, Lr64 (T. dicoccoides); Lr18, Lr50 (Triticum timopheevi); Lr37 (Ae. ventricosa); Lr9, Lr76 (Aegilops umbellulata); Lr19, Lr24, Lr29 (Thinopyrum ponticum); Lr25, Lr26 (Secale cereale); Lr38 (Th. intermedium); Lr54 (Ae. kotschyi); Lr55 (Elymus trachycaulis); Lr56 (Ae. sharonensis); Lr57 (Ae. geniculata); Lr58 (Ae. triuncialis); Lr59 (Ae. peregrina) and Lr62 from Ae. neglecta11,12 besides a number of undesignated genes.
R genes from wild relatives are often linked to genes conferring undesirable traits. It typically takes many years to break this linkage drag and for R genes to be introduced into breeding programs and get deployed. Difficulties associated with genetic characterisation of alien genes include low throughput and physical resolution of cytogenetic techniques10,13,14 and limited transferability of microsatellite markers to the tertiary gene pool15. These factors limited the utilisation of these attractive resources for wheat improvement. However, the recent developments in genomic approaches and the accessibility of various genome sequences has allowed the fast access to genes in wild species.
Majority of the cloned R genes code for proteins with nucleotide-binding and leucine-rich repeat (NLR) domains16,17,18 that either in direct or indirect fashion recognise avirulence proteins. Resistance gene enrichment sequencing (RenSeq) of this particular gene class entails hooking of fragments from a genomic or cDNA library utilizing tailored biotinylated RNA oligonucleotides that complements the genes encoding for NLRs of a reference genome19,20. RenSeq on resistant and susceptible bulks developed from two populations of potato, each depicting segregation for a single R gene to Phytophthora infestans, led to the recognition of SNPs in NLR genes associated with resistance19. Converting RenSeq SNPs into Kompetitive Allele-Specific PCR (KASP) genotyping markers makes the RenSeq approach even more robust for mapping and tracking candidate R genes.
A leaf rust resistant introgression line (IL) pau16061, was developed previously from the hybridisation of the wild species Ae. peregrina (UpUpSpSp) with the wheat line WL711 (vulnerable to rust)21. In the present study, the wheat-Ae. peregrina IL pau16061 was characterised cytogenetically to identify the donor genome of the introgression with leaf rust resistance, and the resistance was mapped with SNP markers designed directly from NLRs identified through MapRenSeq.
Results
Inheritance studies
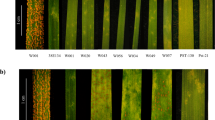
IL pau16061 was resistant to the majority of the leaf rust pathotypes prevalent in the Indian subcontinent Pt11, Pt12-5, Pt16-1, Pt77-5, Pt77-8, Pt77-10, Pt104-2, Pt104B, Pt106 and Pt162-2 at the seedling stage while WL711 was highly susceptible. The IL pau16061/WL711 F5:6 RIL population along with parental lines was tested at the seedling stage against Pt pathotype 77-5. The F1 plants obtained from this cross were resistant with near-immune infection type;, indicating the dominant behavior of the leaf rust resistance gene LrAp. Twelve seedlings from each of 160 F5:6 families were screened. All the seedlings in 69 families showed resistant infection type (;) and these RILs were classified as homozygous resistant, while all the seedlings in 81 families showed infection type 33+ and these RILs were classified as homozygous susceptible (Fig. 1)22. Ten families were found to to be segregating for the leaf rust resistance gene. Seedlings were transplanted in the field and tested for terminal disease severity. At adult plant stage, WL711 showed terminal disease severity of 80 S, whereas Ae. peregrina and IL pau16061 were highly resistant (0-TR). Out of the 160 RILs, 69 were totally free from leaf rust (0) whereas 81 showed leaf rust severity of 60S-80S, and 10 were found segregating. The segregation pattern confirmed a monogenic inheritance of leaf rust resistance (χ21.875:0.25:1.875 = 0.96, p = 0.618) and the resistance locus was tentatively designated as LrAp.
Leaf rust reaction at the seedling stage against leaf rust pathotype 77-5; 1 (Ae. peregrina); 2 (Agra Local); 3 (WL711); 4 (PBW343); 5 (IL pau16061); 6-9 (resistant progenies); 10-11(susceptible progenies). Seedling stage screening was conducted against Pt 77-5 in a glasshouse in PAU, Ludhiana, India. ITs ranging from ‘0’ to ‘2’ considered as resistant and ITs ranging from 3 to 3+ considered as susceptible38.
In situ hybridisation studies
To identify the genomic origin of the Ae. peregrina introgression in IL pau16061, i.e. whether from the Up or Sp genome, GISH was performed with genomic DNA from the U- and S-genome diploid species Ae. umbellulata and Ae. speltoides, respectively, as probes. Utilising S-genome DNA as probe, a strong signal was detected e distally in one pair of sub-metacentric chromosomes (Fig. 2a). GISH using U-genome DNA also detected a weak signal on the distal region of the long arm of a pair of chromosomes with satellites (Fig. 2b) indicating that IL pau16061 carries introgression from both the Up and Sp genomes on two different pairs of chromosomes. To identify the chromosomes carrying the U and S genome introgressions, sequential FISH was conducted on the same metaphase spreads on which U and S genome introgressions were detected with GISH. S genome introgression was confirmed to be on the long arm of wheat chromosome 2B (Fig. 2c) while the U genome introgression was identified to be on the long arm of wheat chromosome 6B (Fig. 2d). GISH using S-genome probe was also conducted on 11 homozygous resistant (HR) and 11 homozygous susceptible (HS) RILs to identify whether the U or S genome introgression carries leaf rust resistance. Strong hybridisation signals were detected in both the HR (Fig. 2e) and HS lines (Fig. 2f), indicating that the S-genome introgression does not carry leaf rust resistance. Sequential FISH also confirmed the recombinant chromosome with GISH signals to be 2B in both HR and HS RILs (Fig. 2g,h). That the U genome introgression confers leaf rust resistance was further confirmed with FISH because a very strong hybridisation site with probe Oligo-pTa535-2 at the very distal end of 6BL was missing in IL pau16061 (Fig. 2c,d) and resistant RILs (Fig. 2g) but present in the susceptible RILs (Fig. 2h). Loss of a wheat specific signal at the telomere of 6BL indicated that the distal end of 6BL in IL pau16061 and resistant lines was replaced by a U-genome introgression. In conclusion, it is U-genome introgression in 6BL that confers the resistance to leaf rust in IL pau16061.
GISH and FISH on the metaphase cells of IL pau16061 (a–d), resistant (e,g) and susceptible RILs (f, h). For GISH, the DNA was labeled with Biotin-16-dUTP and signals were detected with fluorescein-avidin DN, which fluoresces yellow-green. Chromosomes were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) and pseudocolored red. Arrowheads point to the hybridisation signals. For FISH, two probes labelled with FITC and Rhodamine were used to generate chromosome specific hybridisation signals. Chromosomes were counterstained with DAPI. (a) GISH on IL pau16061 using total genomic DNA from Aegilops speltoides; (b) GISH on IL pau16061 using total genomic DNA from Ae. umbellulata; (c,d) Sequential FISH of the same metaphase spreads using probes Oligo-pSc119.2-2 and Oligo-pTa535-2 labelled with 6-carboxyfluorescein (6-FAM) or 6-carboxytetramethylrhodamine (Tamra). Chromosomes have been identified from the banding pattern generated by the hybridization signals as reported by Tang et al.47; (e) GISH using S genome DNA as probe on a resistant RIL; (f) GISH using S genome DNA as probe on a susceptible RIL; (g) Sequential FISH using Oligo-pSc119.2-2 and Oligo-pTa535-2 probes on the same cell in the R RIL as in (e); (h) FISH using Oligo-pSc119.2-2 and Oligo-pTa535-2 probes on the same cell in the S RIL as in (f).
NLR enrichment, sequencing and assembly
For mapping of LrAp, MapRenSeq was employed, the sequential strategy of which is outlined in Supplementary Fig. 1. We performed short read NLR enrichment on IL pau16061, the recurrent susceptible parent WL711, and F5:6 RB and SB. Quantitative PCR on the enriched libraries indicated a 500 to 1,000-fold increase in NLRs relative to other genes. The enriched libraries were paired-end sequenced on the Illumina HiSeq2500 platform. De novo assembly of short reads using CLC resulted in 3,586 NLR contigs (287 of which could be annotated as complete) after running through NLR–parser (Supplementary Table S1). Paired-end read (125 bp) data of susceptible parent and bulks ranged from 1.0 to 1.2 Gb for these libraries, which corresponded to an average NLR coverage of ~100×.
Mapping LrAp with NLR markers
RenSeq BAM files of the resistant parent, susceptible parent and bulks were imported into the Savant genome browser and 3,586 NLR containing contigs were manually inspected to identify polymorphisms. We identified five contigs with SNPs with an allele frequency indicative of linkage to LrAp, i.e. a majority of SNPs derived from the susceptible parent were detected in the SB and a majority of SNPs derived from the resistant parent were detected in the RB (Supplementary Fig. 2). BLASTn searches of these five polymorphic RenSeq contigs against the IWGSC RefSeq v.1.0 assembly gave best hits with the two scaffolds of chromosome 6B. Out of five contigs, four contigs, namely ctg_1473_3, ctg_2659_1, ctg_1087_1 and ctg_765_1, showed a best BLAST hit on 6BL scaffold 119265 at positions 713.06 Mb, 713.14 Mb, 713.78 Mb and 713.96 Mb, respectively, while contig_5109_1 gave a best hit on scaffold 151695 at position 715.97 Mb (Supplementary Table S2).
To validate the putative SNPs and to map LrAp with these markers, we designed KASP assays for five SNPs in NLRs from these contigs. When these KASP markers were applied to the RIL population and the parents, they all formed distinct clusters, clearly separating IL pau16061 specific alleles present in one group and WL711 specific alleles in the other group. The primer sequences for KASP assays are listed in Supplementary Table S3.
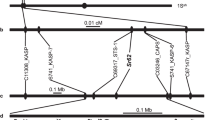
The linkage analysis of the RenSeq markers and phenotypic data indicated that LrAp was cosegregating with these loci indicating the location of all five markers on the alien segment which did not recombine with its wheat counterpart (Fig. 3). Most proximal polymorphic marker Ren_1473 was physically mapped at 713.09 Mb and distal most marker at 715.97 Mb with reference to the wheat RefSeqv1.0 (IWGSC 2018) delineating alien segment to this 2.91 Mb region at the telomeric end of long arm of wheat chromosome 6B. Distal end of chromosome 6BL would be considered as a hotspot for disease resistance genes as 3.00 Mb region encompass 52 genes out of which 34 genes have a signature NB_ARC domain of resistance genes while comparing with high confidence gene annotations in wheat RefSeqv1.1.
Mapping of Ae. peregrina leaf rust resistance gene LrAp. (a) Recombinant chromosome 6B of wheat-Ae. peregrina introgression line pau16061. Ae. peregrina introgression is depicted in blue at the distal end of 6BL. Co-segregating markers derived from NLR contigs along with leaf rust resistance gene LrAp are shown in the box. (b) Physical order of the RenSeq markers on telomeric region of 6BL according to wheat RefSeq v1.0 assembly; (c) candidate scaffolds of 6BL to which polymorphic NLR contigs were anchored from which RenSeq markers were developed.
Discussion
In wheat breeding, the identification, characterisation, mapping and deployment of new major rust resistance genes remains a crucial activity in the constant battle against evolving pathogen populations. Previously, transfer of one stripe rust and two leaf rust resistance genes from Ae. peregrina accession pau3519 to cultivated wheat have been reported21. The present study describes the cytogenetic characterisation and genetic mapping of the leaf rust resistance gene in the wheat-Ae. peregrina introgression line IL pau16061 using exome capture of NLRs (MapRenSeq). Inheritance studies showed that the leaf rust resistance to pathotype 77-5 in IL pau16061 is conferred by a single major dominant gene (LrAp). LrAp mediated resistance is expressed at both the seedling and adult plant stages making it an all stage resistance (ASR) gene. Using GISH and FISH on IL pau16061, we identified that the leaf rust resistance is conferred by an introgression on the long arm of wheat chromosome 6B from the Up genome of Ae. peregrina. Because the introgression lines were developed through induced homoeologous pairing21, most likely it is a compensating translocation from chromosome 6UPL.
Various studies have used high-throughput genotyping combined with bulked segregant analysis (BSA) to identify molecular markers associated with traits of interest23. For example, combined RNA-Seq with BSA to identify SNPs associated with the stripe rust resistance gene Yr15 introgressed into wheat from T. dicoccoides24. However, RNA-Seq is biased by the tissue sampled, the time of sampling and limited by the sequencing depth. This problem is further confounded in the case of NLRs which are often clustered in complex arrays, display extreme sequence and copy number variation between different accessions due to diversifying selection25 and are expressed at relatively low levels24,26.
To mitigate these obstacles, we combined BSA mapping with NLR exome capture and sequencing (dubbed MapRenSeq) to define NLRs associated with LrAp. This strategy was previously used in the wild diploid potatoes Solanum berthaultii and S. ruiz-ceballosii to map Rpi-ber2 and Rpi-rzc1 conferring resistance to late blight19. By capturing and sequencing long NLR fragments using Pacific Biosciences SMRT technology27, improved the resolution of NLR assembly in MapRenSeq. This allowed the fine mapping and cloning of the late blight resistance gene Rpi-amr3i in the wild diploid potato S. americanum. In another variation of the technology, RenSeq was applied to multiple independently derived ethyl methanesulfonate (EMS) mutants of the stem rust resistance genes Sr22 and Sr45 introgressed into hexaploid wheat, which allowed the direct identification of single candidate genes without fine mapping28.
In the present study, we used MapRenSeq to characterise the alien segment carrying the LrAp gene conferring leaf rust resistance in wheat IL pau16061. The approach allowed the development of five co-segregating NLR_SNP markers to LrAp. The linked contigs could be positioned by sequence homology search onto two scaffolds, Ta6BL-151695 and Ta6BL-119265, on the long arm of chromosome 6B of the IWGSC RefSeq v.1.1 assembly. LrAp thus has been introgressed from the Up genome of Ae. peregrina into the long arm of 6B. Both molecular cytogenetic studies and MapRenSeq based mapping unequivocally identified the introgression to be on the wheat chromosome arm 6BL. Previously, due to insufficient SSR marker density at terminal regions we were unable to identify any linked marker with LrAp21. However, MapRenSeq allowed direct access to NLR markers on the terminal alien segment. All the five NLR contigs polymorphic in RB and SB did not recombine with LrAp in 160 RILs indicating one of these might be carrying a putative candidate gene for LrAp. Delineating LrAp introgression will now allows this hypothesis to be tested by mutagenising IL pau16061 to obtain leaf rust susceptible mutants and sequencing these by MutRenSeq to clone LrAp18,28,29. Telomeric region of the wheat chromosome 6B was observed to be a hot spot of disease resistance genes as almost 52 disease resistance genes have been annotated in this region in wheat RefSeqv1.1 (https://urgi.versailles.inra.fr/).
To date, only two other leaf rust resistance genes have been transferred from Ae. peregrina to wheat, namely Lr59 into chromosome arm 1AL30 and LrP linked with a stripe rust resistance gene, YrP, on wheat chromosome arm 5DS31 by our group. The original Lr59-containing translocation has group 6 chromosome homoeology at the distal end 6BS telomere (T1AS·1LP-6SP-6BS)32. The gene LrAp targeted in the present study is clearly different and new since no other published leaf rust resistance gene has been transferred from 6UpL of Ae. peregrina onto chromosome 6B of wheat. Only two Lr genes have been reported on the long arm of 6B viz. Lr3 from T. aestivum33 and Lr9 from the U genome of Ae. umbellulata34,35. The relationship, if any, between LrAp and Lr9 will require further studies.
The identification of a co-segregating markers for LrAp in the present study will facilitate its marker-assisted selection and pyramiding with other major genes such as Yr15, Yr5, Lr76/Yr70, Lr57/Yr40, Lr37/Yr17/Sr38, Yr36 and minor genes like Lr34/Yr18/Sr57 to increase the potential for durable leaf rust resistance.
Methods
Plant materials
IL pau16061 was derived from the cross T. aestivum cv. Chinese Spring PhI/Ae. peregrina pau3519//2*WL711(NN). Leaf and stripe rust susceptible recipient parent WL711 (NN), has the kr (crossability) alleles and is a near-isogenic line of T. aestivum cv. ‘WL711’. The strategy for developing introgression lines has been previously described in detail21. To study the inheritance pattern of rust resistance, IL pau16061 was crossed with WL711 (NN). The F1 was self-pollinated and a population of 160 F5:6 RILs was developed for genetic mapping of leaf rust resistance.
Leaf rust screening
IL pau16061 and WL711 was screened against 10 P. triticina (Pt) pathotypes prevalent in the Indian subcontinent. Seedlings of RILs along with parental lines were screened against the Pt77-536 in a glasshouse in the School of Agricultural Biotechnology, Punjab Agricultural University, Ludhiana, Punjab, India. The pathotype 77-5 is highly virulent, having overcome a number of leaf rust resistance genes in common wheat cultivars37. Seven-day-old seedlings were inoculated with urediniospores mixed with talc, incubated in humidified chambers for 16 h in the dark and then moved to a glasshouse maintained at 18-20 °C. Seedling reactions were recorded 14 days after inoculation and were classified according to the ‘0–4’ infection type (IT) scale38, with ITs ranging from ‘0’ to ‘2’ considered as resistant and ITs ranging from 3 to 3+ considered as susceptible. After recording the ITs, the seedlings were transplanted in the field in 1-meter rows to assess leaf rust severity at the adult plant stage. The susceptible cultivar WL711 was planted all around the experimental area. To maintain rust pressure, a mixture of urediniospores of leaf rust pathotypes 77-2, 77-5 and 104-2 was sprayed over the WL711 infector lines. Disease severity was recorded as the percentage of leaf area covered by rust following a modified Cobb’s scale39.
Cytogenetic characterisation of IL pau16061 and homozygous resistant and homozygous susceptible lines
Genomic in situ hybridisation (GISH) was performed on IL pau16061 following the procedure of Zhang et al. (2001)40. Total genomic DNA from Ae. umbellulata (University of Sydney Cytogenetics collection C64.108) and Ae. speltoides (C64.69) were both labeled with Biotin-16-dUTP (Roche Diagnostic Australia, Castle Hill, NSW, Australia) using nick translation. Unlabeled total genomic DNA of wheat cv. Chinese Spring was used as a blocker. The probe to blocker ratio was ~1:80 and 1:120 for Ae. umbellulata and Ae. speltoides, respectively. Signals were detected with Fluorescein Avidin DN (Vector Laboratories, Burlingame, CA). Chromosomes were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) (Invitrogen Life Science, Carlsbad, CA) and pseudo colored red. Slides were analyzed with a Zeiss Axio Imager epifluorescence microscope. Images were captured with a Retiga EXi CCD (charge-coupled device) camera (QImaging, Surrey, BC, Canada) operated with Image-Pro Plus version 7.0 software (Media Cybernetics Inc., Bethesda, MD) and processed with Photoshop version CS6 software (Adobe Systems, San Jose, CA).
Sequential Fluorescence in situ hybridisation (FISH) was performed on the same metaphase spreads using FITC- and Rhodamine- labelled probes, Oligo-pSc119.2-2 and Oligo-pTa535-2, respectively, for the identification of chromosomes according to Zhang et al. (2019)41.
DNA extraction and quantification
Genomic DNA from young leaves of IL pau16061 (resistant parent), WL711 (susceptible parent) and the RILs was extracted using the CTAB method42 and quantified with Nanodrop and the Quant-iT PicoGreen dsDNA Assay kit (Life Technologies, Carlsbad, CA, USA). Two bulks were produced; resistant bulk (RB) and susceptible bulk (SB), comprising an equimolar concentration of DNA from the 20 most resistant RILs and the 20 most susceptible RILs, respectively.
Illumina library preparation, enrichment and sequencing
A multiplexed NLR capture experiment was conducted on DNA of the resistant parent, susceptible parent, and RB and SB RILs. Short read libraries were prepared on these samples using the NEBNext Ultra DNA Library Prep Kit for Illumina (New England Biolabs, Inc., Ipswich, MA, USA) following the manufacturer’s instructions. Three to four samples were pooled prior to capture. Target NLR capture was carried out using a custom MYcroarray MYbaits kit v2.0 (MI, USA)18. Approximately 500 ng of the prepared libraries were hybridised in hybridisation buffer (10x SSPE, 10x Denhardt’s solution, 10 mM EDTA, 0.2% SDS) to the biotinylated RNA baits for 16-24 h at 65 °C. After hybridisation, bound DNA was recovered using magnetic streptavidin-coated beads and amplified through 8 cycles of PCR. Libraries were purified with AMPure XP beads and quantitative PCR was performed to establish the level of enrichment in NLRs relative to other non-target genes. The enriched libraries were paired-end sequenced on the Illumina HiSeq. 2500 platform at the Earlham Institute (previously The Genome Analysis Center, Norwich Research Park, UK). The raw data is available from NCBI, Study Numbers SRR8485686 (IL pau16061), SRR7867988 (WL711), SRR8369790 (resistant bulk) and SRR8382591 (susceptible bulk). The resistant parent was de novo assembled using CLC assembly cell (http://www.clcbio.com/products/clc-assembly-cell/) and default parameters. All contigs were run through NLR-parser43 to select those harboring NLR sequences.
Genetic mapping of LrAp and chromosome assignment
Quality controlled reads of resistant parent, susceptible parent, RB and SB were mapped onto the assembled resistant parent contigs using Bowtie Wheeler’s Algorithm (BWA) with default settings44. Mapping data were visualised in the Savant Genome Browser (www.genomesavant.com). To identify linked polymorphisms, we looked into all NLR contigs for SNPs between the resistant and susceptible parents. Positive contigs with an alternate allele frequency of >70–80% in the susceptible parent and SB reads with an average coverage above 50 were identified. Regions with an average coverage less than 10% of the median overall coverage were considered as a presence/absence variation. Positive contigs were subsequently used in BLASTn searches (>80% identity) against the International Wheat Genome Sequencing Consortium (IWGSC) RefSeq v.1.0 (IWGSC 2018), to identify the most likely position in the wheat genome.
Primer design
Primers were designed with Primer3 version 4.1.045. The allele specific primers were designed in such a way that they encompass the parental polymorphism in the 3′ end. The reverse primers were selected 40-50 bp downstream specific to the required genome. For each SNP, two allele specific forward oligos with standard FAM or HEX compatible tails (FAM tail: 5′ GAAGGTGACCAAGTTCATGCT 3′; HEX tail: 5′ GAAGGTCGGAGTCAACGGATT 3′) and one common reverse primer were ordered. Primer mix was prepared by mixing 46 µL ddH2O, 30 µL common primer (100 µM) and 12 µL of each allele specific primer (100 µM)46. An assay of 5.07 µl volume included 2.5 µl of 2x KASP master mix (LGC, UK), 0.07 µl of primer mix and 30 ng of gDNA. PCR was performed as described by Ramirez et al. (2015)24. Optically clear plates (384-well) were read on a Tecan Safire plate reader. Fluorescence was observed at 36 °C for 30 s. For allelic discrimination KlusterCaller software v3.4.1.36 (LGC, UK) was used and three genotyping groups, FAM (homozygote), HEX (homozygote) and mixed (heterozygote), were observed.
Statistical analysis and map construction
Chi-squared tests were performed to establish goodness of fit of observed segregation with the expected genetic ratio in the F5:6 RIL population and to detect marker-trait linkages. The positioning of markers was done manually based on recombination between genotypic and phenotypic data and on the basis of recombination frequency. Genetic distances were computed in centiMorgan (cM).
Data availability
The datasets generated during and/or analysed during the current study are available at the NCBI, as under SRR8485686 (IL pau16061), [https://www.ncbi.nlm.nih.gov/sra/?term=SRR8485686], SRR7867988 (WL711), [https://www.ncbi.nlm.nih.gov/sra/?term=SRR7867988] SRR8369790 (resistant bulk), [https://www.ncbi.nlm.nih.gov/sra/?term=SRR8369790] and SRR8382591 (susceptible bulk), [https://www.ncbi.nlm.nih.gov/sra/?term=SRR8382591].
References
Mondal, S. et al. Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front Plant Sci. 7, 991, https://doi.org/10.3389/fpls.2016.00991 (2016).
Wang, C., Hu, S., Gardner, C. & Lübberstedt, T. Emerging Avenues for Utilization of Exotic Germplasm. Trends Plant Sci. 22(7), 624–637 (2017).
Hovmøller, M. S., Walter, S. & Justesen, A. F. Escalating threat of wheat rusts. Science. 329(5990), 369, https://doi.org/10.1126/science.1194925 (2010).
Huerta-Espino, J. et al. Global status of wheat leaf rust caused by Puccinia triticina. Euphytica. 179, 143–160 (2011).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat Ecol Evol. 3(3), 430–439, https://doi.org/10.1038/s41559-018-0793-y (2019).
Germán, S. et al. The situation of common wheat rusts in the Southern Cone of America and perspectives for control. Aust J Agri Res. 58, 620–630 (2007).
Herrera-Foessel, S. A. et al. New slow rusting leaf rust and stripe rust resistance genes Lr67 and Yr46 are pleiotropic or closely linked. Theor Appl Genet. 122, 239–249 (2011).
Bhardwaj, S. C., Gangwar, O. P., Pramod, P. & Khan, H. I. C. A. R. Indian Inst. of Wheat and Barley Research, Regional Station, Flowerdale, Shimla, India. Mehtaensis newsletter 35, 1–16 (2015).
Kiran, K. et al. Draft genome of the wheat rust pathogen (Puccinia triticina) unravels genome-wide structural variations during evolution. Genome Biol Evol. 8, 2702–2721 (2016).
Friebe, B., Jiang, J., Raupp, W. J., McIntosh, R. A. & Gill, B. S. Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica. 91, 59–87 (1996).
Chhuneja, P. et al Introgression and exploitation of biotic stress tolerance from related wild species in wheat cultivars. In: Molecular Breeding for Sustainable Crop Improvement, Sustainable Development and Biodiversity, 11th edn. Sustainable Development and Biodiversity, Springer International Publishing, Switzerland (eds. Rajpal V. R. et al.) 269–324 (2016).
McIntosh, R. A. et al Catalogue of gene symbols for wheat. In: KOMUGI - Wheat Genetic Resources Database at, https://www.shigen.nig.ac.jp/wheat/komugi/genes/download.jsp (2017).
Friebe, B. et al. Identification of alien chromatin specifying resistance to wheat streak mosaic virus and greenbug in wheat germplasm by C-banding and in situ hybridization. Theor Appl Genet. 81, 381–389 (1991).
Lukaszewski, A. J., Lapinski, B. & Rybka, K. Limitations of in situ hybridization with total genomic DNA in routine screening for alien introgressions in wheat. Cytogenet Genome Res. 109, 373–377 (2005).
Mullan, D. J. et al. EST-derived SSR markers from defined regions of the wheat genome to identify Lophopyrum elongatum specific loci. Genome 48, 811–822 (2006).
Dangl, J. L., Horvath, D. M. & Staskawicz, B. J. Pivoting the plant immune system from dissection to deployment. Science 341, 746–751 (2013).
Kourelis, J. & Van der Hoorn, R. A. L. Defended to the nines: 25 years of resistance gene cloning identifies nine mechanisms for R protein function. Plant Cell. 30, 285–299, https://doi.org/10.1105/tpc.17.00579 (2018).
Marchal, C. et al. BED-domain containing immune receptors confer diverse resistance spectra to yellow rust. Nat Plants. 4, 662–668 (2018).
Jupe, F. et al. Resistance gene enrichment sequencing (RenSeq) enables reannotation of the NB-LRR gene family from sequenced plant genomes and rapid mapping of resistance loci in segregating populations. Plant J. 76, 530–544 (2013).
Andolfo, G. et al. Defining the full tomato NB-LRR resistance gene repertoire using genomic and cDNA RenSeq. BMC Plant Biol. 14, 120, https://doi.org/10.1186/1471-2229-14-120 (2014).
Narang, D., Kaur, S., Saini, J. & Chhuneja, P. Development and molecular characterization of wheat-Aegilops peregrina introgression lines with resistance to leaf rust and stripe rust. J Crop Imp. 32, 59–70, https://doi.org/10.1080/15427528.2017.1398117 (2018).
Narang, D. Fine Mapping and identification of candidate gene(s) for leaf rust and stripe rust resistance introgressed in Triticum aestivum L. from Aegilops peregrina. PhD thesis, Punjab Agricultural University, Ludhiana, Punjab, India (2017).
Michelmore, R. W., Paran, I. & Kesseli, R. V. Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc. Natl. Acad. Sci. USA 88, 9828–9832 (1991).
Ramírez-González, R. H. et al. RNA-Seq bulked segregant analysis enables the identification of high-resolution genetic markers for breeding in hexaploid wheat. Plant Biotechnol J. 13, 613–624 (2015).
Meyers, B. C., Kozik, A., Griego, A., Kuang, H. & Michelmore, R. W. Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell. 15, 809–834, https://doi.org/10.1105/tpc.009308 (2003).
Steuernagel, B. et al. Physical and transcriptional organisation of the bread wheat intracellular immune receptor repertoire. Preprint at https://www.biorxiv.org/content/10.1101/339424v1 (2018).
Witek, K. et al. Accelerated cloning of a potato late blight–resistance gene using RenSeq and SMRT sequencing. Nat Biotechnol. 34, 656–660, https://doi.org/10.1038/nbt.3540 (2016).
Steuernagel, B. et al. Rapid cloning of disease resistance genes in plants using mutagenesis and sequence capture. Nat Biotechnol. 34, 652–655 (2016).
Xing, L. et al. Pm21 from Haynaldia villosa encodes a CC-NBS-LRR protein conferring powdery mildew resistance in wheat. Mol Plant. 11(6), 874–878, https://doi.org/10.1016/j.molp.2018.02.013 (2018).
Marais, G. F., McCallum, B. & Marais, A. S. Wheat leaf rust resistance gene Lr59 derived from Aegilops peregrina. Plant Breed. 127, 340–45 (2008).
Narang, D. et al. Fine mapping of Aegilops peregrina cosegregating leaf and stripe rust resistance genes to distal-most end of 5DS. Theor Appl Genet. https://doi.org/10.1007/s00122-019-03293-5 (2019).
Pirseyedi, S. M. et al. Characterization of recombinants of the Aegilops peregrina-derived Lr59 translocation of common wheat. Theor Appl Genet 128, 2403–2414 (2015).
McIntosh, R. A. et al. Catalogue of gene symbols for wheat. Proceedings of the 10th International Wheat Genetics Symposium, Paestum, Italy (2003)
Sears E. R. The transfer of leaf rust resistance from Aegilops umbellulata to wheat. Brookhaven Symp Biol. No. 9, Genetics in Plant Breeding, pp 1–22 (1956).
Sears, E. R. Identification of the wheat chromosome carrying leaf rust resistance from Aegilops umbellulata. Wheat Inf Serv. 12, 12–13 (1961).
Bansal, M. et al. Mapping of Aegilops umbellulata - derived leaf rust and stripe loci in wheat. Plant Pathol. 66, 38–44, https://doi.org/10.1111/ppa.12549 (2017).
Saini, R. G., Kaur, L. & Kaur, M. Adult plant leaf rust Puccinia recondita resistance of known Lr genes against three virulence variants of race 77 from Indian subcontinent. Indian J Agri Sci 68, 776–779 (1998).
McIntosh, R. A., Wellings, C. R. & Park, R. F. Wheat Rusts: an Atlas of Resistance Genes. CSIRO publications, East Melbourne, Australia (1995).
Peterson, R. F., Campbell, A. B. & Hannah, A. E. A diagnostic scale for estimating rust severity on leaves and stem of cereals. Can J Res Sect C Bot Sci. 26, 496–500 (1948).
Zhang, P., Friebe, B., Lukaszewski, A. J. & Gill, B. S. The centromere structure in Robertsonian wheat–rye translocation chromosomes indicates that centric breakage-fusion can occur at different positions within the primary constriction. Chromosoma 110, 335–344 (2001).
Zhang, J. et al. A strategy for identifying markers linked with stem rust resistance in wheat harbouring an alien chromosome introgression from a non-sequenced genome. Theor Appl Genet 132, 125–135 (2019).
Saghai-Maroof, M. A., Biyashev, R. M., Yang, G. P., Zhang, Q. & Allard, R. W. Extraordinarily polymorphic microsatellite DNA in barley: species diversity, chromosomal locations and population dynamics. Proc Natl Acad Sci USA 91, 5466–5470 (1994).
Steuernagel, B., Jupe, F., Witek, K., Jones, J. D. G. & Wulff, B. B. H. NLR-parser: rapid annotation of plant NLR complements. Bioinformatics. 31, 1665–1667 (2015).
Li, H. & Durban, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Untergasser, A. et al. Primer3–new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
LGC Genomics. http://www.lgcgroup.com/services/genotyping (2013).
Tang, Z., Yang, Z. & Fu, S. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J Appl Genet 55, 313–318 (2014).
Acknowledgements
This project was funded by the Sustainable Crop Production Research for International Development (SCPRID) and the Crop Genomics and Technologies (CGAT) programmes from the Department of Biotechnology, Ministry of Science and Technology, Government of India and the Biotechnology and Biological Sciences Research Council, UK grant BT/IN/UK/08/PC/2012 (BB/J012017/1) to PC and CU, grant BT/IN/IndoUK/CGAT/14/PC/2014-15 (BBS/E/J/000CA572) to PC and BBHW and the BBSRC Designing Future Wheat programme (BB/P016855/1). The provision of rust cultures by the Directorate of Wheat Research Regional Research Station, Shimla is greatfully acknowledged. We thank Shorinola Oluwaseyi (John Innes Centre) for help in designing KASP markers, Guotai Yu (John Innes Centre) and Kamil Witek (The Sainsbury Lab) for help with RenSeq library preparation.
Author information
Authors and Affiliations
Contributions
D.N. - Acquisition of data, analysis and interpretation of data, Illumina library preparation, statistical analysis, drafting of manuscript; S.K. - Rust screening and data analysis, Illumina library preparation; B.S. and S.G. - Illumina sequence analysis; U.K.B. – Marker analysis and review of the manuscript; P.Z. and J.L. - Genomic and fluorescence in situ hybridisation, review of the manuscript; S.B. – multi-pathotype screening of the introgression line, review of the manuscript; C.U.- study concept and design, review of the manuscript; B.B.H.W. - study concept and design, study supervision, drafting of manuscript; P.C. - Development of plant material, study concept and design, study supervision, drafting of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Narang, D., Kaur, S., Steuernagel, B. et al. Discovery and characterisation of a new leaf rust resistance gene introgressed in wheat from wild wheat Aegilops peregrina. Sci Rep 10, 7573 (2020). https://doi.org/10.1038/s41598-020-64166-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-64166-2
This article is cited by
-
Broadening the horizon of crop research: a decade of advancements in plant molecular genetics to divulge phenotype governing genes
Planta (2022)
-
Stem and leaf rust–induced miRNAome in bread wheat near-isogenic lines and their comparative analysis
Applied Microbiology and Biotechnology (2022)
-
Multiplex PCR assay for the detection of Lr24 and Lr68 in salt tolerant wheat genotypes
Cereal Research Communications (2022)
-
The grain quality of wheat wild relatives in the evolutionary context
Theoretical and Applied Genetics (2022)
-
An approach for high-resolution genetic mapping of distant wild relatives of bread wheat: example of fine mapping of Lr57 and Yr40 genes
Theoretical and Applied Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.