Abstract
Voltage-gated L-type calcium channels (VLCC) are distributed widely throughout the brain. Among the genes involved in schizophrenia (SCZ), genes encoding VLCC subunits have attracted widespread attention. Among the four subunits comprising the VLCC (α − 1, α −2/δ, β and γ), the γ subunit that comprises an eight-member protein family is the least well understood. In our study, to further investigate the risk susceptibility by the γ subunit gene family to SCZ, we conducted a large-scale association study in Han Chinese individuals. The SNP rs17645023 located in the intergenic region of CACNG4 and CACNG5 was identified to be significantly associated with SCZ (OR = 0.856, P = 5.43 × 10−5). Similar results were obtained in the meta-analysis with the current SCZ PGC data (OR = 0.8853). We also identified a two-SNP haplotype (rs10420331-rs11084307, P = 1.4 × 10−6) covering the intronic region of CACNG8 to be significantly associated with SCZ. Epistasis analyses were conducted and significant statistical interaction (OR = 0.622, P = 2.93 × 10−6, Pperm < 0.001) was observed between rs192808 (CACNG6) and rs2048137 (CACNG5). Our results indicate that CACNG4, CACNG5, CACNG6 and CACNG8 may contribute to the risk of SCZ. The statistical epistasis identified between CACNG5 and CACNG6 suggests that there may be an underlying biological interaction between the two genes.
Similar content being viewed by others
Introduction
Schizophrenia (SCZ) has a lifetime risk of approximately 1% and is a complex psychiatric disorder characterized by delusions, hallucinations, altered cognition, emotional reactivity and disorganized behavior1,2,3. Many epidemiological studies and recent genetic studies of association mapping have suggested that genetic and gene–environment interactions account for over 80% of the susceptibility of developing SCZ4,5. These studies have been accelerated by the widely used application of genome-wide association studies (GWASs) in SCZ6,7,8,9,10. To date, the Psychiatric Genomics Consortium (PGC) has published three mega-analyses on SCZ11,12,13. A recent multi-stage GWAS of SCZ by PGC demonstrated that 108 schizophrenia-associated loci met genome-wide significance in a sample of 36,989 cases and 113,075 controls, 83 of which have not been previously reported13. However, current replicable GWAS results account for only a small percentage of the estimated heritability14 and their systematic biological interpretation is lacking. Focusing on a specific GWAS-based variant might be misleading because the results of a GWAS contains many false positives15,16 and genetic heterogeneity of SCZ exists in different populations17. Thus, follow-up studies are essential to confirm the GWAS findings and extend them.
Voltage-gated L-type calcium channels (VLCC) are distributed widely throughout the brain and are critical for mediating intracellular Ca2 + influx in response to action potentials at the synapse. VLCCs have an important role in N-methyl-d-aspartate (NMDA) receptor-independent synaptic plasticity processes18. Among the genes involved in SCZ, genes encoding VLCC subunits have attracted widespread attention. Recent GWASs have indicated that the CACNA1C gene, encoding an alpha-1 subunit of VLCC, is a susceptibility gene for SCZ in individuals of European ancestry11,12,13. Our previous study has confirmed the association of CACNA1C with SCZ in a Han Chinese population19 and some neuroimaging studies have also documented the effects of CACNA1C variants on a range of structural and functional brain phenotypes, including the circuitry involved in emotion processing, executive function, attention and memory20,21,22. However, among the four subunits comprising the mammalian VLCC (alpha-1, alpha-2/delta, beta and gamma), the gamma subunit is the least well understood. The calcium channel gamma subunits comprise an eight-member protein family that shares a common topology consisting of four transmembrane domains and intracellular N- and C-termini. Some phylogenetic, bioinformatic and functional studies have indicated that they are a functionally diverse protein family and that CACNG1-CACNG8 are all co-expressed in adult brains as an important mechanism for regulating Ca2+ channel function23,24. CACNG2 has been reported to be a susceptibility locus for SCZ in some populations25,26. However, given that some studies suggest an involvement of calcium-dependent regulatory processes in prefrontal-hippocampal network plasticity27,28 and the pathophysiology of SCZ20,21, it is still necessary to explore the potential association between common variants in the CACNG genes and SCZ. In the present study, to further investigate the risk of susceptibility to SCZ due to CACNG1-CACNG8, we conducted a large-sample association analysis in Han Chinese individuals.
Materials and Methods
Subjects
A total of 10,039 individuals, including 7521 healthy controls (3,903 females and 3,618 males; range of age 18–51 years; mean age 37.8 ± 9.6) and 2518 patients with SCZ (1,307 females and 1,211 males; range of age 18–51 years; mean age 36.9 ± 9.3), were recruited for this study. All patients were recruited from the inpatient and outpatient clinical services of a psychiatric unit at the Xi’an Mental Health Center and were diagnosed by at least two experienced psychiatrists based on the Diagnostic and Statistical Manual of Mental Disorders (DSM-IV) criteria for SCZ. All unrelated healthy controls were selected from a combination of local volunteers and blood transfusion donors; those with a personal family history of mental illness in the previous three generations and with current or past evidence of psychoses were ruled out from the present study. All subjects enrolled from the city of Xi’an in Shaanxi Province were of Han descent and we excluded anyone not born locally or whose immediate family members from the previous three generations were not born locally. This research was performed in accordance with the ethical guidelines of the Declaration of Helsinki (version 2002) and was approved by the Medical Ethics Committee of Xi’an Jiaotong University. All participants completed written informed consent forms.
Genotyping
Because CACNG1, CACNG4 and CACNG5 are located next to each other on chromosome 17, we searched for all SNPs in the region covering the three genes with minor allele frequencies (MAF) > 0.05 and heterozygosity (>0.2) in the HapMap Han Chinese in Beijing (CHB) population (Release 28). SNPs with pair-wise tagging and r2 ≥ 0.5 were used as the criteria when selecting tag SNPs in the LD blocks. The same criteria were used to select tag SNPs within CACNG6-8 on chromosome 19, CACNG2 on chromosome 22 and CACNG3 on chromosome 16. Given that the lengths of CACNG2 and CACNG3 are both more than 100 kb, we only selected the tag SNPs (tagging number > 3) in the LD blocks. Based on above criteria, a custom set of 35 tag SNPs were finally selected in our study and the distributions of these SNPs is shown in Supplementary Table S1. DNA was extracted from whole blood according to the standard protocol of the DNA Isolation Kit for Mammalian Blood (Tiangen Biotech Co., Ltd, Beijing, China). The DNA was stored at −80°C for genotyping. The genotyping was conducted for all of the SNPs using the Sequenom MassARRAY matrix-assisted laser desorption ionization-time of flight mass spectrometry platform (Sequenom, San Diego, CA, USA) on the genomic DNA isolated from the peripheral leukocytes. The final data were released using Typer Analyzer software. The reliability of the subsequent statistical analysis was ensured, as the final genotype call rate of each SNP was greater than 99.3% and the overall genotyping call rate was 99.5%. Additionally, 5% of random samples were repeated and the results were 100% concordant.
Statistical analysis
Power analysis and quality control
We conducted a comprehensive power analysis using Genetic Power Calculator29 based on our sample size and genotyped markers to estimate the statistical power. The results are summarized in Supplemental Table S2. As shown in the table, for our sample size (10,039), we were able to achieve a statistical power greater than 80% to detect a common risk genetic variant that had an effect range from 1.2 to 1.3. Genetic markers that had an MAF greater than 0.05 and that passed the Hardy-Weinberg equilibrium test were included for further analysis.
Association Analysis
We performed both single-marker-based and haplotype-based analyses to detect the potential association signals between our genotyped SNPs and the SCZ disorder status. We conducted single-marker-based analysis by fitting a logistic model and the age and sex of the individuals were included in the model as two covariates to account for potential confounding effects. The SNPs were coded in three different models: additive, dominant and recessive. In the additive model, the individual’s genotype is coded as 0, 1 and 2 when there are 0, 1 or 2 minor alleles. In the dominant model, the individual’s genotype is coded as 0 when it is a homozygote of a major allele and 1 when it is a heterozygote or homozygote of a minor allele. In the recessive model, the individual’s genotype is coded as 1 when it is a homozygote of a minor allele and 0 otherwise. The LD blocks were constructed using the default algorithm taken from Gabriel et al.30 We generated 95% confidence bounds on D’ and each comparison was called “strong LD” when the confidence bounds had an upper bound ≥ 0.98 and a lower bound ≥ 0.7. A block was created if 95% of the informative comparisons were “strong LD”. Plink31 was utilized for logistic model fitting, linkage disequilibrium (LD) blocks construction and haplotype-based association. A Bonferroni correction was performed to address the potential multiple comparison problems.
Epistasis Analysis
We conducted two-way epistasis analysis to detect the potential gene-by-gene interactions among the eight CACNG genes. Logistic models were fitted as shown in the following functions,

where Y is the SCZ disorder status. The test for interaction is based on the coefficient b3. Two covariates, sex and age, were added in the logistic model similar to the single-marker-based analysis. The SNP pairs were tested using the logistic model in an exhaustive way. Although SNP pairs located on the same chromosome were removed during analysis, this method still produces a large number of statistical tests (432 tests in total). To reduce the inflated type I errors generated by multiple comparison, we performed 1,000 permutations by shuffling the case-control labels of our samples and the null distribution was created by using the most significant P values in each permutation cycle. The corrected P values for each SNP pair were calculated using this newly created null distribution to account for potential problems of multiple comparison. We utilized statistical computing software R32 to conduct this epistasis analysis.
Imputation and Meta-Analysis
We conducted imputation for a 1 Mb region around SNP rs17645023. We utilized genetic software IMPUTE233 for the imputation and SNPTEST34 for the follow-up dosage data association test. The CHD + CHB HapMap3 dataset served as the reference data. The info metric was used as a method to remove poorly imputed SNPs from our association results. There is no universal cutoff value for post-imputation SNP filtering so we assessed different info thresholds by checking whether they produced sensible Q-Q plots (Supplemental Figure S1). We chose 0.4 as our info metric threshold by the Q-Q plots. In addition to the imputation analysis, we also conducted Meta-analysis for SNP rs17645023 using five GWAS datasets from the PGC. The R package meta35 was utilized for summarizing and virtualization of the meta-analysis.
Bioinformatics analysis
We examined the targeted SNPs and/or genes using several bioinformatics tools and databases. We utilized the protein-protein interaction database STRING (http://string-db.org/) to explore the potential interactions of our targeted genes. The Regulome DB (http://regulomedb.org/) was used to predict the potential functional consequences of the identified risk SNPs. This database is a web-based bioinformatics tool integrated with multiple types of data (including ChIP-seq, DNase-seq and eQTLs) from the Encyclopedia of DNA Elements (ENCODE) project36.
Results
Association analysis
All of our genotyped SNPs passed our quality control criteria (Supplemental Table S3). The P value threshold for single-marker-based analysis was 0.001 (≈0.05/35). The SNP rs17645023 located on the intergenic region of CACNG5 and CACNG4 was identified to be significant in both the additive and dominant models (Table 1). The genotype AA (or allele A) was significantly overrepresented in SCZ individuals. We also identified a significant protective effect of minor allele T in both the additive (OR = 0.86, P = 5.43 × 10−5) and dominant (OR = 0.85, P = 3.46 × 10−4) models. The full results of single marker based association analysis are summarized in Supplemental Table S4. Four LD blocks were constructed using our dataset. In the haplotype-based analysis, we found a two-SNP haplotype (rs10420331-rs11084307, P = 1.4 × 10−6) to be significantly associated with the SCZ disease status. This SNP pair covers the intronic region of gene CACNG8. The full results of haplotype analysis are summarized in Table 2.
Epistasis analysis
The two-way epistasis analysis included 432 SNP pairs and the full results are summarized in Supplemental Table S5. We identified significant statistical interaction (OR = 0.622, P = 2.93 × 10−6, Pperm < 0.001) between rs192808 (CACNG6) and rs2048137 (CACNG5) (Fig. 1). Both double homozygote GG-TT and double heterozygote GA-GT (rs192808-rs2048137) were overrepresented in control samples.
Epistasis analysis results for rs192808 and rs2048137.
(a) Average percentage of SCZ patients for different genotypes of SNP rs2048137. (b) Average percentage of SCZ patients for different genotypes of SNP rs192808. c. Average percentage of SCZ patients for the combination genotype of rs192808 and rs2048137. The average percentage of SCZ patients in all samples is indicated with the dotted line. The significant P values based on the additive model are shown in each plot.
Imputation and Meta-Analysis
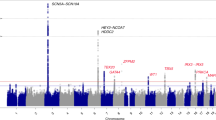
A total of 73 imputed SNPs passed our quality control criteria (info > 0.4). The full results of imputation association are summarized in Supplemental Table S6. No new significant SNPs were identified with imputation analysis. The regional association plot based on both imputed and genotyped SNPs is shown in Fig. 2. The results of the meta-analysis are presented in Fig. 3 and the summarized result of the meta-analysis focused on rs17645023 was still significant with OR = 0.8853. The detailed information on the datasets used in the meta-analysis is summarized in Supplemental Table S7.
Bioinformatics Analysis
We explored the protein-protein interaction for the two genes with significant statistical epistasis evidence (CACNG5 and CACNG6) (Fig. 4). No direct evidence was found to support the protein-protein interaction between the two genes. The most related calcium channel voltage-dependent gamma subunits for CACNG6 are CACNG7 and CACNG8. Conversely, for CACNG5, the most related protein is encoded by CACNG1. We investigated the potential functional consequences of rs17645023 in Regulome DB. This SNP has a Regulome DB score of 4, which means the potential biological function of this SNP is supported by both transcription factor (TF) binding and DNase peak data. The ChIP-seq data indicates that this SNP is located within the protein binding region of the transcription factor CEBPB in all three cell types (A549, HeLa-S3 and IMR90). In addition, rs17645023 is also located within the DNase hypersensitive region in all eight cell types (T47d, Sknmc, A549, Hvmf, Hpde6e6e7, Fibrop, Fibropag20443 and Fibrobl).
Discussion
One major finding in our study is the identification of a significant association between an intergenic SNP, rs17645023 and SCZ disease status. The potential association of this SNP with SCZ was first reported by Curtis et al. in 2011 using 523 SCZ cases and 505 controls with European ancestry (P < 10−6)37. The T allele at SNP rs17645023 is protective in both our study and the study of Curtis et al. However, the follow-up GWAS with the largest sample sizes has not replicated this result13. Our study is the first conducted with individuals of Chinese Han ancestry to replicate the significant association of rs17645023 and SCZ. The finding in this study of the association between rs17645023 and SCZ has generated more questions than answers. As an intergenic SNP, it is difficult to map to a specific gene. In the study conducted by Curtis et al., rs17645023 was originally mapped to CACNG5, which is located 35 kb upstream from rs17645023. However, rs17645023 was also shown to be located approximately 44 kb upstream of CACNG4. It should not be assumed that this SNP is mapped to CACNG5 simply because it is located 9 kb closer than CACNG4. Fortunately, we now have more efficient bioinformatics tools and large-scale datasets (such as ENCODE). A bioinformatics analyses indicates that rs17645023 may affect the binding of the transcription factor CEBPB, suggesting that it is more reasonable to map this SNP to CACNG4, which is located downstream of the SNP.
Our haplotype-based analyses provided some clues to a potential association between CACNG8 and SCZ. To our knowledge, our study is the first to identify such a link. Although more studies are still needed to illustrate the specific effects of the CACNG8 gene on SCZ, a general discussion based on previous literature might be helpful to provide some clues. Rouach et al. studied the role of the Cacng8 protein in mice38. The authors found that mice deficient in the Cacng8 protein showed a differential regulation of functional α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptor (AMPA receptor) in the extrasynaptic and synaptic pools. These findings indicate that the Cacng8 protein may act as a critical protein for AMPA receptor expression and distribution in the hippocampus of mice. Interestingly, multiple studies have demonstrated that SCZ is related to brain region- and subunit-specific abnormalities in the expression of the AMPA receptor subunits39,40. In this sense, the CACNG8 gene may affect the onset and development of SCZ through its effect on the expression and distribution of AMPA receptors.
Numerous studies have indicated that the epistasis effects can account for a large portion of the genetic effects for a complex disorder41,42,43. Early studies have provided statistical evidence for the epistasis between DISC1, CIT and NDEL1 and risk for SCZ44. Our two-way interaction analyses identified a significant SNP pair (rs192808- rs2048137, CACNG6-CACNG5). Despite the problems associated with multiple comparisons, the result of the permutation analysis (Pperm < 0.001) indicates that this signal cannot be attributed to chance. Nevertheless, the identification of a statistical epistasis among susceptible genes does not illustrate their underlying biological interactions. As discussed above, CACNG5 was reported to be associated with SCZ in the study of Curtis et al.37, although it is unlikely the SNP rs17645023 maps to CACNG5. CACNG6 was previously reported to be associated with aspirin-intolerance in asthmatics in a Korean population45 and to date, no evidence has shown a potential link between CACNG6 and SCZ. Based on sequence homology and chromosomal linkage, the calcium channel γ subunits can be divided into three clusters: (γ1,γ6), (γ5,γ7) and (γ2,γ3,γ4,γ8)24. Therefore, the proteins encoded by CACNG5 and CACNG6 belong to different clusters of calcium channel γ subunits. Our bioinformatics analysis using STRING found no direct evidence for the protein-protein interaction between the two different proteins encoded by CACNG5 and CACNG6. More studies are still needed to clarify the underlying biological interactions between the two proteins.
A major strength of our study is the candidate gene-based study design. Although a GWAS has a much larger genomic coverage, its results are susceptible to many type I errors. Additionally, applying the Bonferroni correction for the inflated type I errors will result in the loss of several signals with moderate effects. Our candidate gene-based design avoids this disadvantage by focusing on a couple of functionally related genes and controlling the number of genotyped markers to a reasonable level. Another strength of our study is that we have a much larger sample size (~10,000) in comparison to other similar studies. A power analysis showed that this sample size enables us to detect moderate genetic effects with ORs as small as 1.15. There are also several limitations of our study. The coverage of our genotyped SNPs is a major disadvantage of this study. In the present study, we genotyped only 35 SNPs (~4 SNPs per gene) within the calcium channel γ subunit gene family (CACNG1-CACNG8). Therefore, it is doubtful that our study is complete because potential significant signals may be missed due to the low coverage of our genetic markers. Although this limitation can be partly compensated through imputation analyses, our results should be confirmed in other cohorts and populations using high-density marker panels and large-scale genotyping to provide more confident evidence for the association of the SNPs with SCZ. In addition to the low coverage of genetic markers, the population stratification might be another limitation for our study. Unlike large-scale genetic studies, which enable researchers using high-density marker data to apply some statistical intensity methods (e.g., genomic control, PCA) to correct the underlying population stratifications, we cannot perform any direct controls in our study because of the limited number of genotyped markers. Another limitation of this study is that in order to reduce the genotyping costs and maximize the genetic information, all SNPs genotyped in our study are tag SNPs in the LD blocks and very few of them can be constructed into LD blocks again. As a result of this strategy, it is difficult to carry out haplotype-based analysis for the SNP of rs17645023 (due to the difficulties of constructing LD blocks) and we lost the chance to further validate our significant signal from a single-marker analysis in the haplotype-based analysis. In addition, although several quality control processes have been applied during the sample collection process to ensure the uniformity of the genetic background of our samples, this does not guarantee the complete removal of the confounding effects of different sub-populations especially considering the genetic diversity of the Han Chinese population. Therefore, our findings should be considered preliminary and additional follow-up studies are required, including high-density mapping and targeted deep sequencing to provide additional information beyond studies focused on common variants and to undercover fundamental characteristics of any potential associations with SCZ. In addition, in addition to the calcium channel γ gene family, recent studies have also identified several susceptible genes for SCZ within the α and β gene families. Systematic studies focusing on all the calcium channel gene families are needed to illustrate the biomedical and the biological significance of these genes on SCZ.
In summary, our results indicated that CACNG4, CACNG5, CACNG6 and CACNG8 might be genes that contribute to the risk of SCZ. The statistical epistasis identified between CACNG5 and CACNG6 suggests that there is an underlying biological interaction between the two genes. However, more research is still needed to confirm our findings and to clarify the pathological mechanisms of the functional role of the calcium channel γ subunits gene family in SCZ and to eventually confer changes in clinical practice.
Additional Information
How to cite this article: Guan, F. et al. Evaluation of voltage-dependent calcium channel γ gene families identified several novel potential susceptible genes to schizophrenia. Sci. Rep. 6, 24914; doi: 10.1038/srep24914 (2016).
References
Sullivan, P. F., Daly, M. J. & O’Donovan, M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat Rev Genet 13, 537–551 (2012).
Owen, M. J. Implications of genetic findings for understanding schizophrenia. Schizophrenia Bull 38, 904–907 (2012).
Chen, G., Guan, F., Lin, H., Li, L. & Fu, D. Genetic analysis of common variants in the HDAC2 gene with schizophrenia susceptibility in Han Chinese. J Hum Genet 60, 479–484 (2015).
Gejman, P. V., Sanders, A. R. & Kendler, K. S. Genetics of schizophrenia: new findings and challenges. Annu Rev Genomics Hum Genet 12, 121–144 (2011).
McGrath, J. J., Mortensen, P. B., Visscher, P. M. & Wray, N. R. Where GWAS and epidemiology meet: opportunities for the simultaneous study of genetic and environmental risk factors in schizophrenia. Schizophrenia Bull 39, 955–959 (2013).
O’donovan, M. C. et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat Genet 40, 1053–1055 (2008).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Shi, J. et al. Common variants on chromosome 6p22. 1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Yue, W. H. et al. Genome-wide association study identifies a susceptibility locus for schizophrenia in Han Chinese at 11p11. 2. Nat Genet 43, 1228–1231 (2011).
Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat Genet 43, 969–976 (2011).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 45, 1150–1159 (2013).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Ozomaro, U., Wahlestedt, C. & Nemeroff, C. B. Personalized medicine in psychiatry: problems and promises. BMC Med 11, 132 (2013).
Burmeister, M., McInnis, M. G. & Zöllner, S. Psychiatric genetics: progress amid controversy. Nat Rev Genet 9, 527–540 (2008).
Johnson, A. D. & O’Donnell, C. J. An open access database of genome-wide association results. BMC Med Genet 10, 6 (2009).
Guan, F. et al. Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese. Sci Rep 5 (2015).
Moosmang, S. et al. Role of hippocampal Cav1. 2 Ca2+ channels in NMDA receptor-independent synaptic plasticity and spatial memory. J Neurosci 25, 9883–9892 (2005).
Guan, F. et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr Res 152, 97–104 (2014).
Bigos, K. L. et al. Genetic variation in CACNA1C affects brain circuitries related to mental illness. Arch Gen Psychiat 67, 939–945 (2010).
Erk, S. et al. Brain function in carriers of a genome-wide supported bipolar disorder variant. Arch Gen Psychiat 67, 803–811 (2010).
Thimm, M. et al. Effects of a CACNA1C genotype on attention networks in healthy individuals. Psychol Med 41, 1551–1561 (2011).
Burgess, D. L., Gefrides, L. A., Foreman, P. J. & Noebels, J. L. A cluster of three novel Ca2+ channel γ subunit genes on chromosome 19q13. 4: evolution and expression profile of the γ subunit gene family. Genomics 71, 339–350 (2001).
Chen, R. S., Deng, T.-C., Garcia, T., Sellers, Z. M. & Best, P. M. Calcium channel γ subunits: a functionally diverse protein family. Cell Biochem Biophys 47, 178–186 (2007).
Wilson, G. M. et al. DNA copy-number analysis in bipolar disorder and schizophrenia reveals aberrations in genes involved in glutamate signaling. Hum Mol Genet 15, 743–749 (2006).
Liu, Y. L. et al. RASD2, MYH9 and CACNG2 genes at chromosome 22q12 associated with the subgroup of schizophrenia with non-deficit in sustained attention and executive function. Biol Psychiatry 64, 789–796 (2008).
Spitzer, N. C. Electrical activity in early neuronal development. Nature 444, 707–712 (2006).
Zoladz, P. R. et al. Differential expression of molecular markers of synaptic plasticity in the hippocampus, prefrontal cortex and amygdala in response to spatial learning, predator exposure and stress-nduced amnesia. Hippocampus 22, 577–589 (2012).
Purcell, S., Cherny, S. S. & Sham, P. C. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19, 149–150 (2003).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296(5576), 2225–2229 (2002).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575 (2007).
Ripley, B. D. The R project in statistical computing. MSOR Connections 1, 23–25 (2001).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat Genet 44, 955–959 (2012).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39, 906–913 (2007).
Schwarzer, G. meta: General Package for Meta-Analysis. R package version 4.3-1. http://CRAN.R-project.org/package=meta (2015).
The ENCODE (ENCyclopedia of DNA elements) project. The ENCODE (ENCyclopedia of DNA elements) project. Science 306, 636–640 (2004).
Curtis, D. et al. Case-case genome wide association analysis reveals markers differentially associated with schizophrenia and bipolar disorder and implicates calcium channel genes. Psychiat Genet 21, 1 (2011).
Rouach, N. et al. TARP γ-8 controls hippocampal AMPA receptor number, distribution and synaptic plasticity. Nat Neurosci 8, 1525–1533 (2005).
Tucholski, J. et al. Abnormal N-linked glycosylation of cortical AMPA receptor subunits in schizophrenia. Schizophr Res 146, 177–183 (2013).
Drummond, J. B., Tucholski, J., Haroutunian, V. & Meador-Woodruff, J. H. Transmembrane AMPA receptor regulatory protein (TARP) dysregulation in anterior cingulate cortex in schizophrenia. Schizophr Res 147, 32–38 (2013).
Bell, J. T. et al. Genome-ide Association Scan Allowing for Epistasis in Type 2 Diabetes. Ann Hum Genet 75, 10–19 (2011).
Yi, N. et al. Characterization of epistasis influencing complex spontaneous obesity in the BSB model. Genetics 167, 399–409 (2004).
Wei, W. H., Hemani, G. & Haley, C. S. Detecting epistasis in human complex traits. Nat Rev Genet 15, 722–733 (2014).
Nicodemus, K. K. et al. Evidence of statistical epistasis between DISC1, CIT and NDEL1 impacting risk for schizophrenia: biological validation with functional neuroimaging. Hum Genet 127, 441–452 (2010).
Lee, J. S. et al. Association of CACNG6 polymorphisms with aspirin-intolerance asthmatics in a Korean population. BMC Med Genet 11, 138 (2010).
Acknowledgements
This research was supported by and National Natural Science Foundation of China (No. 81401563, 81373253 and 81273351), Shaanxi Province Natural Science Foundation (No. 2015JQ8306) and Ph.D. Programs Foundation of Ministry of Education of China (No. 2013021120078) and Shaanxi Key Project on Science and Technology (2014K11-02-03-05). The funding sources had no role in the design of this study, the collection, analysis and interpretation of data, the writing of the report, or the decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
Authors F.G. and Li.T. conceived and designed the study. Li.L and G.C. carried out the computational analyses and candidate SNPs selection. X.L., W.H. and H.L. conducted subject screening. F.G. and T.Z. performed the statistical analysis of the genotype and haplotype data. G.F. and T.Z. wrote the first draft of the paper and T.L. revised the current paper. W.H. and H.L. recruited, diagnosed and gathered patients. L.L and G.C. contributed to the collection and preparation of control DNA samples.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Guan, F., Zhang, T., Liu, X. et al. Evaluation of voltage-dependent calcium channel γ gene families identified several novel potential susceptible genes to schizophrenia. Sci Rep 6, 24914 (2016). https://doi.org/10.1038/srep24914
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24914
This article is cited by
-
Patient-specific identification of genome-wide DNA-methylation differences between intracranial and extracranial melanoma metastases
Scientific Reports (2023)
-
SNP rs10420324 in the AMPA receptor auxiliary subunit TARP γ-8 regulates the susceptibility to antisocial personality disorder
Scientific Reports (2021)
-
Genome-wide association studies for production, respiratory disease, and immune-related traits in Landrace pigs
Scientific Reports (2021)
-
Evaluation of genetic susceptibility between systemic lupus erythematosus and GRB2 gene
Scientific Reports (2019)
-
Relationship of common variants in MPP7, TIMP2 and CASP8 genes with the risk of chronic achilles tendinopathy
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.