Abstract
Cancer stem cells are responsible for tumor progression, metastasis, therapy resistance and cancer recurrence, doing their identification and isolation of special relevance. Here we show that low adherent breast and colon cancer cells subpopulations have stem-like properties. Our results demonstrate that trypsin-sensitive (TS) breast and colon cancer cells subpopulations show increased ALDH activity, higher ability to exclude Hoechst 33342, enlarged proportion of cells with a cancer stem-like cell phenotype and are enriched in sphere- and colony-forming cells in vitro. Further studies in MDA-MB-231 breast cancer cells reveal that TS subpopulation expresses higher levels of SLUG, SNAIL, VIMENTIN and N-CADHERIN while show a lack of expression of E-CADHERIN and CLAUDIN, being this profile characteristic of the epithelial-to-mesenchymal transition (EMT). The TS subpopulation shows CXCL10, BMI-1 and OCT4 upregulation, differing also in the expression of several miRNAs involved in EMT and/or cell self-renewal such as miR-34a-5p, miR-34c-5p, miR-21-5p, miR-93-5p and miR-100-5p. Furthermore, in vivo studies in immunocompromised mice demonstrate that MDA-MB-231 TS cells form more and bigger xenograft tumors with shorter latency and have higher metastatic potential. In conclusion, this work presents a new, non-aggressive, easy, inexpensive and reproducible methodology to isolate prospectively cancer stem-like cells for subsequent biological and preclinical studies.
Similar content being viewed by others
Introduction
The cancer stem cell (CSC) model posits that tumors are maintained by a subpopulation of cells that self-renew and yield heterogeneous progeny with reduced proliferative potential1,2. Different molecular mechanisms activated in normal stem cells are also involved in CSC self-renewal, including the expression of certain embryonic stem cells-transcription factors (ES-TFs)3 or the similar regulation of several signaling pathways4,5. Additionally, short non-coding miRNAs are also able to modulate gene expression programs to maintain self-renewal in normal and CSCs6.
It is important to note that CSCs also underlie drug resistance, tumor recurrence and metastasis1,2,7. Drug resistance and tumor recurrence showed by CSCs are mainly explained by the overexpression of multidrug resistance (MDR) membrane proteins and the enzyme aldehyde dehydrogenase (ALDH), or their ability to maintain a quiescent state8. On the other hand, metastasis is one of the most crucial steps in cancer progression and the main cause of mortality1. During local invasion and distant metastasis, the associated cancer cells typically develop alterations in their shape as well as in their attachment to other cells and to the extracellular matrix9. Metastatic cancer cells are characterized for suffering an epithelial-to-mesenchymal transition (EMT), a process by which cancer cells lose their attachment to the epithelial niche and acquire a mesenchymal phenotype10. These cells then are transported through the vasculature and are disseminated to anatomically distant organ sites where are able to establish new neoplastic growths11,12.
The relevance of this cancer cell subpopulation has yield to develop methodologies for their identification and isolation. Breast CSCs are characterized by the phenotype CD44+/CD24low/− 13,14, while the expression of the cell surface proteins CD133, CD44 and/or CD326 is related with colon CSCs properties15,16,17. Other CSCs characteristics that have been extensively used for their identification and isolation are their high ALDH activity and ability to exclude Hoechst 33342, which is used to determine the side population (SP) phenotype1,18,19,20,21. However, the isolation of CSCs based on all of these properties requires the fluorescence-activated cell sorting (FACS) method, an expensive and aggressive technique.
As it has been reported previously, cancer cells that undergo EMT acquire characteristics of CSCs22,23. On the other hand, it is known that during EMT process cells lose their adhesion ability with adjacent epithelial cells9. This property has been extensively used to remove mesenchymal cells from primary epithelial cell cultures following the protocol known as “differential trypsinization” developed by Owens et al. in 197424. Here we report that the application of this technique in cancer cell cultures shows that cells selected by differential trypsinization differ in phenotypical and functional CSCs properties, including ALDH activity, SP proportion, xenograft tumor formation ability and metastatic potential, among others. As expected, trypsin-sensitive (TS) cancer cells subpopulations show increased CSCs properties when compared with the total population (TP) and/or the trypsin-resistant (TR) subpopulation.
Materials and Methods
Cell lines and cell culture
Breast (MCF7 and MDA-MB-231) and colon (HT-29 and T84) cancer cell lines were obtained from American Type Culture Collection (ATCC) and cultured following ATCC recommendations.
Differential trypsinization
Cells at 60–80% of confluence were slowly washed with phosphate buffered saline (PBS) without directly flow falls on the cells. Then, 0.05% trypsin was added and incubated 2 minutes at 37 °C. Detached cells were collected in centrifuge tubes and were named as Trypsin-Sensitive 1 (TS1). The remaining attached cells were washed twice and, then, 0.25% trypsin was added. These cells were named as Trypsin-Resistant 1 (TR1). On the other hand, TS1 cells were again plated for 24 hours. After that, 0.05% trypsin was added and incubated for 2 minutes at 37 °C. These detached cells were named as Trypsin-Sensitive 2 (TS2) (Fig. 1A).
Differential trypsinization protocols.
(A) Isolation of TS1, TS2 and TR1 subpopulations, used for Aldefluor assays, Hoechst 33342, surface markers, spheres and colonies formation ability. (B) Improved protocol to isolate highly attached cells (TR2) used for Aldefluor, qRT-PCR and in vivo assays. Red balls represent cells with higher CSC-like properties and green balls cells with a non CSC-like phenotype.
In order to separate more strictly cells with stem-like and no stem-like properties, cells were grown until 60–80% of confluence and washed slowly with PBS. After obtaining the TS1 subpopulation, remaining cells attached to the dishes were washed twice with PBS and incubated with 0.05% trypsin for 4 minutes at 37 °C. Cells detached from this trypsinization were discarded. Dishes with remaining trypsin-resistant cells were named as TR2 (Fig. 1B). Total population (TP) was used as control cells.
Flow cytometry analyses
ALDEFLUOR assays (Stem Cell Technologies) to detect ALDH1 activity were performed according to manufacturer’s instructions. Diethylaminobenzaldehyde (DEAB) was used as an ALDH1 inhibitor to set ALDH1 gates. Cell surface levels were determined with anti-human antibodies CD44-PE, CD24-APC, CD133-APC, CD326-FITC, CK18-FITC and CK20-FITC (Miltenyi Biotec). All samples were analyzed on a FACS CANTO II (BD Biosciences) using the FACS DIVA software.
Side population assays
Hoechst 33342 exclusion (Side Population) assays were carried out as previously described21,25.
Sphere-forming assay
3 × 103 cells were washed with PBS and resuspended in spheres culture medium (DMEM:F12, 1% penicilin/streptomycin, B27, 10 μg/mL ITS, 1 μg/mL Hydrocortisone, 4 ng/mL Heparin, 10 ng/mL EGF, 20 ng/mL FGF) in ultra-low adherence 24-well plates (Corning). Spheres >75 μM diameter were counted after 6 days by light microscopy.
For the secondary sphere-forming assay, cells from primary spheres were collected by centrifugation, then dissociated with trypsin-EDTA and mechanically disrupted with a pipette. 103 single cells were plated and resuspended in spheres culture medium in ultra-low adherence 24-well plates. Spheres >75 μM diameter were counted after 6 days by light microscopy.
Soft agar assay
104 cells were seeded in 0.4% cell agar base layer, which was on top of 0.8% base agar layer in 6-well culture plates. Cells were then incubated for a further 21 days at 37 °C and 5% CO2. Cell colony formation was then examined under a light microscope after staining with 0.01% Crystal Violet (Sigma-Aldrich).
qRT-PCR
Total RNA extraction and qRT-PCR assays were done following standard protocols. See Supplementary Material and Methods for more details. Primer sequences are listed in Supplementary Table 1.
Proliferation assay
Different subpopulations (TP, TS1 and TR2) were seeded in 24-well plates in medium supplemented with 10% FBS for 6 days. Cells were incubated with MTT every two days and the fluorescence was measured at 570 nm.
Lentivirus transfections
293T cells were co-transfected with a lentiviral vector encoding for the firefly luciferase and the red fluorescent protein td-Tomato (L2T)26, a packaging vector (psPAX2) and an envelope vector (pCMV-VSVG) using lipofectamine transfection reagent (Life Technologies). Viral particles produced from 293T cells were collected and used to infect MDA-MB-231 cells. Briefly, supernatant recovered after 48h from 293T transfected cells was filtered by a 0.45 μm pore membrane and added to MDA-MB-231 plated cells supplemented with 4 μg/mL polybrene (Sigma-Aldrich). After viral infection, td-Tomato+ cells were selected for stable integration of the transfects by FACS.
Mouse models
Tumor-initiation experiments and experimental lung metastasis assays are detailed in Supplementary Materials and Methods. All animal studies were performed according to protocols reviewed and approved by the Institutional Animal Care and Use Committee at the University of Granada.
Histological and immunological analysis
Described in detail in Supplementary Materials and Methods.
Statistical analysis
All graphed data are presented as mean ± SD from at least three experiments. two-tailed Student’s t-tests and two-way ANOVA were used to determine statistical differences for in vitro and in vivo experiments respectively. P-values <0.05 were considered statistically significant in all cases.
Results
Trypsin-sensitive subpopulation displays CSC-like phenotypic properties
To test whether TS subpopulations are enriched in CSC-like properties, we determined the ALDH activity, the rate of SP and specific breast and colon CSC-related cell surface markers (Fig. 2). Regarding to ALDH activity, both TS1 and TS2 subpopulations presented a significantly higher proportion of ALDH positive cells than TP. In contrast, TR1 subpopulation displayed a lesser ALDH activity, although this change was not significant when compared to the TP (Fig. 2A, Supplementary Fig. S1A and Supplementary Table 2).
Phenotypic properties of breast and colon cancer cell subpopulations isolated by differential tripsinization.
Percentage of ALDH+ (A) and SP (B) cells determined in subpopulations of MCF7, MDA-MB-231, HT-29 and T84 cancer cell lines. (C,D) Percentage of CD44+/CD24− cells in TR1, TP, TS1 and TS2 isolated from MCF7 (C) and MDA-MB-231 (D) breast cancer cells measured by flow cytometry. (E) Co-expression of CD44, CD133 and CD326 in HT-29 cells isolated by differential trypsinization and in the TP. (F) Flow cytometry for CD44 and CD326 expression, alone or together, in T84 cells. Data are graphed as mean ± SD from at least two different experiments carried-out by triplicates (**P < 0.01; *P < 0.05). See also Supplementary Fig. S1.
Accordingly, the rate of SP in TS subpopulations was significantly higher than in both TP and TR1 cells for all cell lines tested. Thus, TS1 cells showed values ranged from 48.6% up to 70.5% and TS2 from 82.3% up to 89.9%. However, TP cells displayed percentages comprising between 3.9% and 13.2% (Fig. 2B, Supplementary Fig. S1B and Supplementary Table 3).
Moreover, we determined CD44/CD24 expression for breast and CD44/CD133/CD326 expression for colon cancer cell lines to study the proportion of cancer cells with a CSC phenotype in the different subpopulations. Flow cytometry analysis indicated that in the MCF7 cell line, TS1 and TS2 subpopulations showed a significantly higher CD44+/CD24− proportion than TP and TR1 (Fig. 2C). However, no significant differences were found for these markers in the different subpopulations of the MDA-MB-231 breast cancer cell line, probably due to the high proportion of cells with this phenotype in the TP (Fig. 2D). In HT-29 colon cancer cells, the expression of CD44+/CD133+/CD326+ was significantly higher for TS1 and TS2 in comparison to TR1, although only TS2 showed a significant increase in the proportion of cells with this phenotype when compared to the TP (Fig. 2E). It is important to note that HT-29 cells showed two different subpopulations based on CD326 expression and that CD326high and CD326low HT-29 cells were separated prospectively by differential trypsinization, being the TS subpopulation composed for CD326high cells (Supplementary Fig. S1C). In regards to T84 colon cancer cells, TS1 and TS2 showed a higher enrichment in cells with a CD44+/CD326+ phenotype, even when their expression was studied together or separate (Fig. 2F). Differences in the expression of CD133 were not significant; however, this cell surface marker was more expressed in TS subpopulations (Supplementary Fig. S1D).
Trypsin-sensitive subpopulation possesses higher self-renewal ability and clonogenicity
We next tested in vitro functional characteristics of these cell subpopulations isolated by differential trypsinization. We studied the anchorage-independent growth of TS and TR subpopulations under free-serum conditions to determine their self-renewal ability; and also their clonogenic capacity by colony-formation assay in soft agar. As it is shown in Fig. 3A,B, tumorspheres were formed more efficiently from TS1 and TS2 than TR1 and TP cells in all cell lines; therefore TS subpopulations have a higher capacity of self-renewal. Moreover, TR1 subpopulation formed significantly less spheres than TP for MCF7, MDA-MB-231 and T84 (Fig. 3A,B).
Tumorsphere- and colony-forming ability of breast and colon cancer cell subpopulations isolated by differential trypsinization.
(A) Representative images of mammospheres formed from different MCF7 subpopulations. (B) Number of spheres formed by different subpopulations of each cell line. (C) Secondary sphere formation assay performed in MDA-MB-231 and T84 cells selected by differential trypsinization and compared to the TP. Data shown as mean ± SD (**P < 0.01; *P < 0.05). (D) Number of colonies formed by TP and TS1 of each cell line. Data are represented as mean ± SD (**P < 0.01; *P < 0.05).
We also performed a secondary sphere-formation assay by dissociating primary spheres formed by TS1, TR1 and TP cells, as indicated in the Materials and Methods section. As it is shown in Fig. 3C for the breast cancer MDA-MB-231 and colon cancer T84 cell lines, the number of cells with the ability to form secondary spheres was higher in primary spheres formed by the TS1 subpopulation, while those formed by TR1 were poorly enriched in sphere-forming cells. In addition, TS subpopulations showed a significant higher capacity to form colonies in soft agar than TP for all cell lines studied (Fig. 3D). These data are in concordance with our previous results obtained from the primary sphere formation assay and support our prior conclusion about the higher self-renewal ability of the TS subpopulation.
All together, these results indicate that cells isolated by differential trypsinization differ in their enrichment in cells with functional CSC-like properties. Furthermore, the rate of these cells in the subpopulations isolated is well maintained when cells are cultured in suspension in FBS-free media.
Isolation of cancer cells with stem- and no stem-like properties based on their adhesion capacity
As we shown above, we obtained a cell subpopulation enriched for CSCs features based in their lower adhesion capacity to the cell culture surface. The amount of cells detached after the first trypsinization was very low and the rest of attached cells was very similar to the total population. For this reason, to demonstrate that it is possible to separate CSCs and cancer cells no stem based on their sensitivity to trypsin digestion, we developed another protocol. After the first trypsinization we recovered the TS1 subpopulation and remaining attached cells were trypsinized again with the same trypsin dilution indicated above, but in this case for 4 minutes. Cells that were detached by this last trypsinization were discarded and cells still attached were recovered and named as TR2, as described in the Material and Methods section (Fig. 1B). As it is shown in Fig. 4A and Supplementary Fig. S2, TP and subpopulations TS1 and TR2 differed in ALDH activity. In concrete, TS1 subpopulation was enriched in cells with a high ALDH activity, with purity higher than 90% and TR2 showed a lack of ALDH activity in both MDA-MB-231 and MCF7 (Fig. 4A and Supplementary Fig. S2). As it is shown the purity of ALDH+ cells obtained by differential trypsinization from MCF7 cells was similar to the enrichment obtained by FACS (Supplementary Fig. S2).
ALDH activity, mRNA and miRNA expression in enriched subpopulations of MDA-MB-231 cells isolated by their different adhesion capacity.
(A) Aldefluor assay performed in TR2, TP and TS1 MDA-MB-231 cells. See also Supplementary Fig. S2. (B) qRT-PCR analysis for the expression of CSCs and EMT-related genes in different subpopulations of MDA-MB-231 cells. Data are normalized to 1 for TP using GAPDH as internal control and graphed as mean ± SEM (n = 3) (**P < 0,01; *P < 0,05). (C) Differential expression of miRNAs related to CSC phenotype and/or EMT in MDA-MB-231 cells isolated by differential trypsinization. Data are normalized to 1 for TP using miR-24c-3p as internal control and graphed as mean ± SEM (n = 3) (**P < 0.01; *P < 0.05).
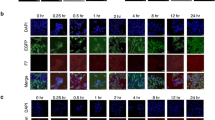
On the other hand, we also studied the expression levels of both the luminal cytokeratin 18 and the intestinal epithelium differentiation cytokeratin 20 for breast and colon cancer cells, respectively, after differentiation of TS1 cells in media containing FBS for 7 days when grown attached to the cell culture surface. As it is shown in Supplementary Fig. S3, TS1 cells cultured under these conditions recovered the expression levels of these cytokeratins at similar values than those observed in the TP. These results are an indicator of the ability of the TS1 cells to recapitulate the heterogeneity of the parental cell line.
Trypsin-sensitive subpopulation expresses higher levels of EMT- and pluripotency- associated genes
The enrichment of cancer cells with stem cell-like properties in cells which show low adherence to plastic surface may be due to an EMT process, which is associated with metastasis. To understand the molecular changes underlying the different characteristics of these cell subpopulations, we analyzed the expression of genes related with the EMT process and/or pluripotency on MDA-MB-231 cells.
qRT-PCR studies demonstrated that all genes assayed were expressed differentially in the TS1 and the TR2 cell subpopulations. SNAIL, SLUG, VIMENTIN, N-CADHERIN, CXCL10, OCT4 and BMI1 were overexpressed in the TS1 subpopulation when compared to the TP and underexpressed in the TR2 subpopulation (Fig. 4B). In contrast, expression levels of CLAUDIN1 and E-CADHERIN were higher in TR2 cells and much lower in the TS1 subpopulation (Fig. 4B).
On the other hand, the expression patterns of several miRNAs related with EMT and/or pluripotency also differ between these two cell subpopulations isolated by differential trypsinization. The TR2 subpopulation showed a significant increase in the expression levels of miR-93-5p, miR-34a-5p, miR-100-5p and miR-34c-5p (Fig. 4C). However, all of these miRNAs were downregulated in the TS1 subpopulation when compared to the TP, although not all of them were statistically significant (Fig. 4C). In contrast, the expression of miR-21-5p was higher in TS1 and lower in TR2 cells when compared to the TP (Fig. 4C).
Moreover, the proliferation assay demonstrated that both TR2 and TP subpopulations grow faster that TS1 for all tumor cell lines (Fig. 5), showing that the TS1 subpopulation has a lower proliferation rate in vitro.
Trypsin-sensitive subpopulation is enriched in tumor-initiating cells
To confirm the results observed in vitro, TS1 and TR2 subpopulations were isolated from MDA-MB-231 cells, injected subcutaneously in NSG mice and compared with the TP in a limiting dilution in vivo assay. The overall survival of mice injected with TS1 cells was lower than for the other groups (Fig. 6A). When 5 × 103 TS1, TP or TR2 cells were injected, the overall survival of mice was 63 and 77 days, for TS1 and TP respectively, while 2 out of 6 mice injected with the TR2 subpopulation were still alive at day 81, the end-point of the experiment (Fig. 6A). Furthermore, tumor volume was higher after injection of TS1 cells, while tumors formed by TR2 cells were smaller (Fig. 6B), being these results not explained by a global mitogenic effect since TR2 cells have higher proliferation rate than TS1 cells (Fig. 5). Similar results were observed when 1.5 × 104 cells were injected, however the overall survival was shorter and the tumor size bigger for all groups, as it was expected (Fig. 6A,B). All together, these data demonstrate that TS cells are enriched in CSCs with a high tumor-initiating capacity while the TR subpopulation is composed mainly by non CSC-like cells.
Overall survival and subcutaneous xenograft tumor formation
(A) Kaplan-Meier curve for overall survival of NSG mice injected with 5 × 103 cells (left) and 1.5 × 104 cells (right) from TS1, TR2 or TP. (B) Tumor volume (mm3) of subcutaneous xenograft tumors formed by 5 × 103 and 1.5 × 104 cells from TS1, TR2 and TP in NSG mice. Data are shown as mean ± SD (**P < 0.01; *P < 0.05).
Trypsin-sensitive subpopulation generates more metastasis than trypsin-resistant subpopulation after tail vein injection
To further demonstrate the higher aggressiveness of the TS subpopulation, we compared the metastatic ability of both cell subpopulations isolated by differential trypsinization (TS1 and TR2) with the TP using MDA-MB-231 L2T cells. When 2.5 × 105 TS1 cells were injected into the tail vein of NSG mice, 5 of 6 animals developed lung metastasis, however only 4 and 2 of 6 animals developed metastasis when TP or TR2 cells, respectively, were injected (IVIS images from week 3 are shown in Fig. 7A). In addition, the TS1 subpopulation, that showed the lowest proliferation rate in vitro (Fig. 5), caused a more rapid increase in tumor bioluminescence, demonstrating their higher metastatic ability (Fig. 7A). After 4 weeks, animals were sacrificed and lungs harvested for further analysis (Fig. 7B). Lungs from mice injected with TS1 cells showed more pulmonary metastatic nodules than these from TP or TR2 cells (representative images, Fig. 7C). Furthermore, histological and immunohistochemical analysis of lungs demonstrated that metastatic nodules formed by TS1 cells were bigger when compared to the TP and smaller when TR2 cells were injected (Fig. 7D,E), being these differences in size independent of a global mitogenic effect (Fig. 5). This further demonstrates the higher metastatic ability of the TS subpopulation and that the TR subpopulation is composed by non CSC-like cells.
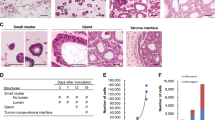
Experimental lung metastasis assay.
(A) IVIS images of lung metastasis formed after tail vein injection of TP, TR2 and TS1 MDA-MB-231 L2T cells (up). Measure of photon flux is graphed as mean ± SEM (down). (B) Fluorescence of metastatic lungs ex vivo. (C) Representative optical images of lung metastasis for a healthy mice (control) and lungs extracted from NSG mice injected with different subpopulations of MDA-MB-231 cells obtained by differential trypsinization. (D,E) Histological (D) and immunohistochemical (E) images of lungs obtained from healthy mice (control) and mice injected with TR2, TP and TS1 cells (left to right).
Discussion
In this work, we demonstrate that cultured cancer cells possess low adherent cancer cell subpopulations with stemness properties. Therefore, differential trypsinization, an extended methodology for the establishment of primary epithelial cell cultures24,27, can be applied to isolate breast and colon cancer cells with CSCs properties in vitro and in vivo and likewise for other cancer types with an epithelial origin. In this work, we demonstrate that two distinct subpopulations of cancer cells can be isolated by this protocol: i) a subpopulation with CSCs properties and lower adherence ability (TS subpopulation); ii) and a highly adherent subpopulation enriched in non CSC-like cells (TR subpopulations).
Breast and colon cancer TS subpopulation is enriched in cells with high ALDH activity and greater SP rate, being these characteristics associated with CSCs properties. The isolation of a subpopulation with high ALDH activity has been demonstrated to show higher tumorigenic potential in both breast and colon cancers18,20. Regarding to Hoechst 33342 dye efflux, different studies have shown that the SP phenotype is enriched in cancer stem-like cells in different tumors, including breast and colon cancers19,21. Moreover, the ability to exclude Hoechst 33342 is associated with an upregulation of ATP-binding cassette transporters that confer multidrug resistance, a characteristic of CSCs28.
However, the characterization of CSCs by the expression of specific cell surface markers is the most extended method to identify and isolate this cancer cell subpopulation. We have demonstrated that MCF7 breast cancer cells from the TS subpopulation are enriched in cells with a CD44+/CD24− phenotype, which is characteristic of breast CSCs13. On the other hand, HT-29 and T84 TS colon cancer cells are enriched in CD44+, CD133+ and/or CD326+ cells. Colon CSCs were firstly identified based on the expression of CD133, being determined that the ability to develop tumors in immunodeficient NOD/SCID mice was restricted to CD133+ cells15,16. In a latter study, Dalerba et al. demonstrated that the ability to engraft in immunodeficient mice was restricted to a minority subpopulation CD326high/CD44+. Furthermore, tumors originated from CD326high/CD44+ cells maintained a differentiated phenotype and reproduced the full morphologic and phenotypic heterogeneity of their parental lesions17. Regarding to MDA-MB-231 TS cells, we did not see a significant increase in the CD44+/CD24− subpopulation, however the percentage in all groups was very high, around 80%, which is in concordance with the highly invasive/metastatic skills of this triple negative breast cancer cell line29.
Breast and colon cancer cells isolated by differential trypsinization also differ in their self-renewal ability. CSC self-renewal can be assayed in vitro by tumor spheres and holoclone colonies formation assays30. The TS subpopulation obtained from both breast and colon cancer cell lines generated more colonies in soft agar and spheres in low attachment plates under serum deprivation. It has been demonstrated that colon CSCs enriched in CD133+ cells form more colonies in soft agar used as indicator of their higher tumor-initiating ability31. In addition, it has been shown that MDA-MB-231 breast cancer cells enriched in cancer-initiating stem cells have higher colony-formation ability32. On the other hand, CSCs have the ability to form spheres in low-attachment culture surfaces under free serum conditions and this kind of culture conditions increases the percentage of cells with a CSC phenotype based on surface markers expression25,33. In contrast, the number of colonies and spheres formed from TR cells was significantly lower than in the TP, indicating the lower self-renewal ability and therefore, the non CSC-like properties of this cancer cell subpopulation selected by a higher adhesion capability to the cell culture surface.
All these in vitro studies for phenotypical and functional properties of CSCs were notably increased in the TS subpopulations for all cell lines, except for CD44+/CD24− expression in MDA-MB-231 breast cancer cells that is basally high in TP cells. For this reason, we selected this cell line to perform further studies. qRT-PCR analysis of different mRNA and miRNA related with EMT, self-renewal and/or aggressiveness demonstrated differences in their expression between TR and TS subpopulations. The gene profile of MDA-MB-231 TS cells was more associated with a mesenchymal phenotype while the TR subpopulation was more related with an epithelial phenotype. During the EMT process, cancer cells lose their attachment to the epithelial niche and acquire a mesenchymal phenotype which is characteristic of metastatic cancer cells34. Moreover, the triple-negative MDA-MB 231 breast cancer cell line, which overexpresses EGFR35, has potent and reproducible migratory responses36. MDA-MB-231 TS cells showed a lack of expression of E-CADHERIN and CLAUDIN. The loss by carcinoma cells of E-cadherin, a key cell-to-cell adhesion molecule which establish adherent junctions with adjacent epithelial cells, is the best characterized alteration during the EMT process9,34. In regard to claudin expression, a phenotype claudin-low is associated with enrichment for EMT markers, immune response genes, cancer stem cell-like features and poor outcome in breast cancer37. In addition, the TS subpopulation also shows a higher expression of several transcription factors that modulates the EMT, such as SNAIL or SLUG and other mesenchymal markers including N-CADHERIN and VIMENTIN38. In contrast, the expression of these genes was contrary regulated in MDA-MB-231 TR cells, indicating their more epithelial features. In fact, we have recently demonstrated important role of Vimentin/Slug EMT markers in tumor growth of primary breast cancer patients and in circulating tumor cells (CTCs) EGFR+ from cytokeratin negative non-metastatic breast cancer patients39.
Certain ES-TFs, such as Oct4, Sox2 and Nanog, which are master regulators of ES self-renewal and pluripotency, are also upregulated in CSCs40. OCT4 expression was measured in MDA-MB-231 cells, demonstrating that it is overexpressed in the TS subpopulation. Interestingly, a recent study showed that Oct4 is associated with poor prognosis and that positively regulates the EMT process, contributing to breast cancer metastasis38. Furthermore, CSCs share similarities with normal stem cells in the regulation of the Wnt, Notch and Hedgehog signaling pathways and the expression of several genes of the polycomb family, including BMI-1, or the HOX family among others which are involved in stem cells self-renewal4. BMI-1, which is overexpressed in TS cells, has been also related with the EMT process and drug resistance41. Moreover, CXCL10 is expressed in basal tumors as compared with ER+ tumors, being associated with a higher tumor grade and a poor prognosis of breast cancer42.
On the other hand, miRNAs play critical regulatory roles in a wide range of biologic and pathologic processes and miRNA expression profiling reveals substantial differences between cancer and normal tissue and among different cancer types43. We studied the expression patterns of different miRNAs that have been previously shown to be related with CSCs properties, EMT, metastasis and poor outcome. For example, the TR subpopulation overexpresses miR-93, which inhibits cell proliferation, the capacity to form colonies44 and promotes mesenchymal-to-epithelial transition (MET)45. miR-34a-5p and miR-34c-5p are downregulated in TS cells and upregulated in the TR subpopulation. miR-34 family are reportedly as tumor-suppressor miRNAs implicated in reduced CSC properties and increased sensitivity to drug treatment by directly targeting NOTCH146. miR-100 is also upregulated in the TR subpopulation and its expression levels relate to the cellular differentiation state, with lowest expression in cells displaying stem cell markers47. In addition, the expression of miR-21, a well known miRNA which is dysregulated in several types of cancer and plays a key role in carcinogenesis, recurrence and metastasis48, is increased in the TS subpopulation and decreased in TR cells.
All together, these data demonstrate that differential trypsinization of breast and colon cancer cell cultures allows the isolation of two subpopulations (TR and TS) that differ in their CSC-like cells properties. TS cells show increased ALDH activity, SP proportion and CSC-like phenotype. This subpopulation is enriched in cells that have followed an EMT process and possess high self-renewal ability. Moreover, TS cells show higher tumorsphere- and colony-formation ability in vitro; and tumorigenic and metastatic potential in vivo. In contrast, the TR subpopulation is mainly composed by non CSC-like cells with a lower tumorigenic capacity both in vitro and in vivo. In a previous study carried-out by Walia and Elbe, it was shown that the TS subpopulation, obtained by differential trypsinization from transformed human mammary epithelial cells, was enriched in stem cell properties49. In a more recent study, this methodology was used to isolate a subpopulation of cells with a mesenchymal phenotype from an oncogenic transformed human mammary epithelial cell line50. However, these studies were limited to determine EMT markers expression, mammosphere-forming ability, drug resistance and CD44/CD24 expression and were carried-out in a well differentiated oncogenic transformed human mammary epithelial cell line.
Conclusion
In summary, our results evidence that cultured cancer cells possess low adherent cell subpopulations with stemness properties and with high tumorigenic and metastatic abilities. We show for the first time that differential trypsinization, a new, non-aggressive, easy, inexpensive and reproducible methodology to isolate prospectively breast and colon cancer stem-like cells for subsequent biological studies, high throughput screening therapeutic strategies targeting CSCs and preclinical assays.
Additional Information
How to cite this article: Morata-Tarifa, C. et al. Low adherent cancer cell subpopulations are enriched in tumorigenic and metastatic epithelial-to-mesenchymal transition-induced cancer stem-like cells. Sci. Rep. 6, 18772; doi: 10.1038/srep18772 (2016).
References
Tirino, V. et al. Cancer stem cells in solid tumors: an overview and new approaches for their isolation and characterization. FASEB J 27, 13–24, 10.1096/fj.12-218222 (2013).
O’Brien, C. A., Kreso, A. & Dick, J. E. Cancer stem cells in solid tumors: an overview. Semin Radiat Oncol 19, 71–77, 10.1016/j.semradonc.2008.11.001 (2009).
Zeineddine, D., Hammoud, A. A., Mortada, M. & Boeuf, H. The Oct4 protein: more than a magic stemness marker. Am J Stem Cells 3, 74–82 (2014).
Liu, S. et al. Hedgehog signaling and Bmi-1 regulate self-renewal of normal and malignant human mammary stem cells. Cancer Res 66, 6063–6071, 10.1158/0008-5472.CAN-06-0054 (2006).
O’Brien, C. A., Kreso, A. & Jamieson, C. H. Cancer stem cells and self-renewal. Clin Cancer Res 16, 3113–3120, 10.1158/1078-0432.CCR-09-2824 (2010).
Chhabra, R. & Saini, N. MicroRNAs in cancer stem cells: current status and future directions. Tumour Biol 35, 8395–8405, 10.1007/s13277-014-2264-7 (2014).
Visvader, J. E. & Lindeman, G. J. Cancer stem cells: current status and evolving complexities. Cell Stem Cell 10, 717–728, 10.1016/j.stem.2012.05.007 (2012).
Raha, D. et al. The cancer stem cell marker aldehyde dehydrogenase is required to maintain a drug-tolerant tumor cell subpopulation. Cancer Res 74, 3579–3590, 10.1158/0008-5472.CAN-13-3456 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674, 10.1016/j.cell.2011.02.013 (2011).
Ombrato, L. & Malanchi, I. The EMT universe: space between cancer cell dissemination and metastasis initiation. Crit Rev Oncog 19, 349–361, 10.1615/CritRevOncog.2014011802 (2014).
Talmadge, J. E. & Fidler, I. J. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res 70, 5649–5669, 10.1158/0008-5472.CAN-10-1040 (2010).
Valastyan, S. & Weinberg, R. A. Tumor metastasis: molecular insights and evolving paradigms. Cell 147, 275–292, 10.1016/j.cell.2011.09.024 (2011).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci U S A 100, 3983–3988, 10.1073/pnas.0530291100 (2003).
Azzam, D. J. et al. Triple negative breast cancer initiating cell subsets differ in functional and molecular characteristics and in gamma-secretase inhibitor drug responses. EMBO Mol Med 5, 1502–1522, 10.1002/emmm.201302558 (2013).
O’Brien, C. A., Pollett, A., Gallinger, S. & Dick, J. E. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445, 106–110, 10.1038/nature05372 (2007).
Ricci-Vitiani, L. et al. Identification and expansion of human colon-cancer-initiating cells. Nature 445, 111–115, 10.1038/nature05384 (2007).
Dalerba, P. et al. Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci U S A 104, 10158–10163, 10.1073/pnas.0703478104 (2007).
Ginestier, C. et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 1, 555–567, 10.1016/j.stem.2007.08.014 (2007).
Haraguchi, N. et al. Cancer stem cells in human gastrointestinal cancers. Hum Cell 19, 24–29, 10.1111/j.1749-0774.2005.00004.x (2006).
Huang, E. H. et al. Aldehyde dehydrogenase 1 is a marker for normal and malignant human colonic stem cells (SC) and tracks SC overpopulation during colon tumorigenesis. Cancer Res 69, 3382–3389, 10.1158/0008-5472.CAN-08-4418 (2009).
Britton, K. M., Kirby, J. A., Lennard, T. W. & Meeson, A. P. Cancer stem cells and side population cells in breast cancer and metastasis. Cancers (Basel) 3, 2106–2130, 10.3390/cancers3022106 (2011).
Liu, X. & Fan, D. The epithelial-mesenchymal transition and cancer stem cells: functional and mechanistic links. Curr Pharm Des, 21, 1279–1291, 10.2174/1381612821666141211115611 (2015).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715, 10.1016/j.cell.2008.03.027 (2008).
Owens, R. B., Smith, H. S. & Hackett, A. J. Epithelial cell cultures from normal glandular tissue of mice. J Natl Cancer Inst 53, 261–269 (1974).
Kanwar, S. S., Yu, Y., Nautiyal, J., Patel, B. B. & Majumdar, A. P. The Wnt/beta-catenin pathway regulates growth and maintenance of colonospheres. Mol Cancer 9, 212, 10.1186/1476-4598-9-212 (2010).
Liu, H. et al. Cancer stem cells from human breast tumors are involved in spontaneous metastases in orthotopic mouse models. Proc Natl Acad Sci USA 107, 18115–18120, 10.1073/pnas.1006732107 (2010).
Garcia-Posadas, L., Arranz-Valsero, I., Lopez-Garcia, A., Soriano-Romani, L. & Diebold, Y. A new human primary epithelial cell culture model to study conjunctival inflammation. Invest Ophthalmol Vis Sci 54, 7143–7152, 10.1167/iovs.13-12866 (2013).
Natarajan, K., Xie, Y., Baer, M. R. & Ross, D. D. Role of breast cancer resistance protein (BCRP/ABCG2) in cancer drug resistance. Biochem Pharmacol 83, 1084–1103, 10.1016/j.bcp.2012.01.002 (2012).
Xu, J., Prosperi, J. R., Choudhury, N., Olopade, O. I. & Goss, K. H. Beta-catenin is required for the tumorigenic behavior of triple-negative breast cancer cells. PLoS One 10, e0117097, 10.1371/journal.pone.0117097 (2015).
Shigeishi, H. et al. Maintenance of stem cell self-renewal in head and neck cancers requires actions of GSK3beta influenced by CD44 and RHAMM. Stem Cells 31, 2073–2083, 10.1002/stem.1418 (2013).
Puglisi, M. A. et al. Isolation and characterization of CD133+ cell population within human primary and metastatic colon cancer. Eur Rev Med Pharmacol Sci 13 Suppl 1, 55–62 (2009).
Zhao, D. et al. VEGF drives cancer-initiating stem cells through VEGFR-2/Stat3 signaling to upregulate Myc and Sox2. Oncogene, 10.1038/onc.2014.257 (2014).
Rappa, G. et al. Growth of cancer cell lines under stem cell-like conditions has the potential to unveil therapeutic targets. Exp Cell Res 314, 2110–2122, 10.1016/j.yexcr.2008.03.008 (2008).
Cervantes-Arias, A., Pang, L. Y. & Argyle, D. J. Epithelial-mesenchymal transition as a fundamental mechanism underlying the cancer phenotype. Vet Comp Oncol 11, 169–184, 10.1111/j.1476-5829.2011.00313.x (2013).
Corkery, B., Crown, J., Clynes, M. & O’Donovan, N. Epidermal growth factor receptor as a potential therapeutic target in triple-negative breast cancer. Ann Oncol 20, 862–867, 10.1093/annonc/mdn710 (2009).
Price, J. T., Tiganis, T., Agarwal, A., Djakiew, D. & Thompson, E. W. Epidermal growth factor promotes MDA-MB-231 breast cancer cell migration through a phosphatidylinositol 3’-kinase and phospholipase C-dependent mechanism. Cancer Res 59, 5475–5478 (1999).
Sabatier, R. et al. Claudin-low breast cancers: clinical, pathological, molecular and prognostic characterization. Mol Cancer 13, 228, 10.1186/1476-4598-13-228 (2014).
Wang, D. et al. Oct-4 and Nanog promote the epithelial-mesenchymal transition of breast cancer stem cells and are associated with poor prognosis in breast cancer patients. Oncotarget 5, 10803–10815 (2014).
Serrano, M. J. et al. EMT and EGFR in CTCs cytokeratin negative non-metastatic breast cancer. Oncotarget 5, 7486–7497 (2014).
Liu, A., Yu, X. & Liu, S. Pluripotency transcription factors and cancer stem cells: small genes make a big difference. Chin J Cancer 32, 483–487, 10.5732/cjc.012.10282 (2013).
Paranjape, A. N. et al. Bmi1 regulates self-renewal and epithelial to mesenchymal transition in breast cancer cells through Nanog. BMC Cancer 14, 785, 10.1186/1471-2407-14-785 (2014).
Mulligan, A. M. et al. Tumoral lymphocytic infiltration and expression of the chemokine CXCL10 in breast cancers from the Ontario Familial Breast Cancer Registry. Clin Cancer Res 19, 336–346, 10.1158/1078-0432.CCR-11-3314 (2013).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838, 10.1038/nature03702 (2005).
Leal, J. A. & Lleonart, M. E. MicroRNAs and cancer stem cells: therapeutic approaches and future perspectives. Cancer Lett 338, 174–183, 10.1016/j.canlet.2012.04.020 (2013).
Liu, S., Clouthier, S. G. & Wicha, M. S. Role of microRNAs in the regulation of breast cancer stem cells. J Mammary Gland Biol Neoplasia 17, 15–21, 10.1007/s10911-012-9242-8 (2012).
Park, E. Y. et al. Targeting of miR34a-NOTCH1 axis reduced breast cancer stemness and chemoresistance. Cancer Res 74, 7573–7582, 10.1158/0008-5472.CAN-14-1140 (2014).
Deng, L. et al. MicroRNA100 inhibits self-renewal of breast cancer stem-like cells and breast tumor development. Cancer Res 74, 6648–6660, 10.1158/0008-5472.CAN-13-3710 (2014).
Liu, J. et al. MicroRNA-21 is a novel promising target in cancer radiation therapy. Tumour Biol 35, 3975–3979, 10.1007/s13277-014-1623-8 (2014).
Walia, V. & Elble, R. C. Enrichment for breast cancer cells with stem/progenitor properties by differential adhesion. Stem Cells Dev 19, 1175–1182, 10.1089/scd.2009.0430 (2010).
Tam, W. L. et al. Protein kinase C alpha is a central signaling node and therapeutic target for breast cancer stem cells. Cancer Cell 24, 347–364, 10.1016/j.ccr.2013.08.005 (2013).
Acknowledgements
We thanks Dr Gambhir’s laboratory from the Standford University for providing L2T plasmid. This work was supported in part by grants from the Ministry of Economy and Competitiveness, Instituto de Salud Carlos III (FEDER funds, projects n°. PI10/02295 and PI10/02149), the Fundación Progreso y Salud, Consejería de Salud, Junta de Andalucía (Ministry of Health, Government of Andalucia, project number PI-0533-2014) and by Consejería Economía, Innovación, Ciencia y Empleo, Junta de Andalucia (Ministry of Economy, Innovation, Science and Employment, Government of Andalucia, excellence project number CTS-6568) providing a fellowship granted to GJ.
Author information
Authors and Affiliations
Contributions
C.M.-T. Conception and design, collection and/or assembly of data, data analysis and interpretation, manuscript writing; G.J., J.M.E., C.G.-L. Collection and/or assembly of data, data analysis and interpretation. M.A.G. Data analysis and interpretation, manuscript writing; M.A. Financial support, provision of study material; M.P.-R. Conception and design, collection and/or assembly of data, data analysis and interpretation, manuscript writing and final approval of manuscript; J.A.M. Conception and design, financial support, administrative support, provision of study material, data analysis and interpretation, manuscript writing and final approval of manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Morata-Tarifa, C., Jiménez, G., García, M. et al. Low adherent cancer cell subpopulations are enriched in tumorigenic and metastatic epithelial-to-mesenchymal transition-induced cancer stem-like cells. Sci Rep 6, 18772 (2016). https://doi.org/10.1038/srep18772
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18772
This article is cited by
-
Biophysics in tumor growth and progression: from single mechano-sensitive molecules to mechanomedicine
Oncogene (2023)
-
A reliable mouse model of liver and lung metastasis by injecting esophageal cancer stem cells (CSCs) through tail-vein injection
Molecular Biology Reports (2023)
-
Sijunzi Decoction Inhibits Stemness by Suppressing β-Catenin Transcriptional Activity in Gastric Cancer Cells
Chinese Journal of Integrative Medicine (2022)
-
Identification of PARP-1 in cancer stem cells of gastrointestinal cancers: A preliminary study
Journal of Biosciences (2021)
-
Glucose-limiting conditions induce an invasive population of MDA-MB-231 breast cancer cells with increased connexin 43 expression and membrane localization
Journal of Cell Communication and Signaling (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.