Abstract
CD4+FOXP3+ regulatory T (Treg) cells are essential for maintaining immunological self-tolerance. Treg cell development and function depend on the transcription factor FOXP3, which is present in several distinct isoforms due to alternative splicing. Despite the importance of FOXP3 in the proper maintenance of Treg cells, the regulation and functional consequences of FOXP3 isoform expression remains poorly understood. Here, we show that in human Treg cells IL-1β promotes excision of FOXP3 exon 7. FOXP3 is not only expressed by Treg cells but is also transiently expressed when naïve T cells differentiate into Th17 cells. Forced splicing of FOXP3 into FOXP3Δ2Δ7 strongly favored Th17 differentiation in vitro. We also found that patients with Crohn’s disease express increased levels of FOXP3 transcripts lacking exon 7, which correlate with disease severity and IL-17 production. Our results demonstrate that alternative splicing of FOXP3 modulates T cell differentiation. These results highlight the importance of characterizing FOXP3 expression on an isoform basis and suggest that immune responses may be manipulated by modulating the expression of FOXP3 isoforms, which has broad implications for the treatment of autoimmune diseases.
Similar content being viewed by others
Introduction
CD4+FOXP3+ regulatory T (Treg) cells suppress immune activation in a dominant manner and are essential for maintenance of immunological tolerance1. Treg cells depend on the forkhead/winged-helix transcription factor FOXP3 to be able to exert their function2,3,4. The importance of FOXP3 for efficient regulation of the immune system is best illustrated by the development of lethal lymphoproliferative disease in both mice and humans with genetic deficiencies in FOXP3, known respectively as scurfy and immune dysregulation, polyendocrinopathy, enteropathy, X-linked (IPEX) syndrome5,6,7. Deficiency in Treg cell function has also been suggested as an underlying cause for disease conditions ranging from autoimmune diseases to infectious diseases8.
The FOXP3 gene encodes a transcription factor that contains three functional domains including a proline-rich N-terminal domain encoded by exons 2–4, a zinc finger and leucine zipper domain encoded by exons 5–7 and a fork-head domain encoded by exons 9–119,10,11,12. The proline-rich N-terminal domain of FOXP3 enables the protein to interact with transcriptional repressors and activators, which consequently alter gene expression and regulate the suppressive ability of Treg cells13. The zinc finger and leucine zipper domain of FOXP3 is necessary and sufficient to undergo homo-oligomerization and hetero-association with other transcription factors, such as FOXP19,11,14,15. Lastly, FOXP3 has a high specificity for gene regulation, conferred by the C terminal fork-head domain, which mediates DNA binding of FOXP312,16.
Alternative splicing is a strictly regulated process wherein particular exons of a pre-mRNA are either included in or excluded from the mature mRNA. Alternative splicing consequently allows a single gene to give rise to multiple proteins that can have different or even opposing functions. The two most abundant FOXP3 isoforms, full-length FOXP3 (FOXP3fl) and FOXP3 lacking exon 2 (FOXP3Δ2), confer suppressive ability to Treg cells17,18. In contrast, FOXP3 lacking exons 2 and 7 (FOXP3Δ2Δ7) has been reported to inhibit other FOXP3 isoforms in a dominant negative manner19. A recent study has also demonstrated that exon 7 of FOXP3 is required for proper Treg cell function, as two different point mutations located near the intron 7 splice donor site result in excision of FOXP3 exon 7 and IPEX syndrome20. Despite the importance of FOXP3 in Treg cells, the regulation and functional consequences of FOXP3 isoform expression remains poorly understood.
Results
Alternative splicing of FOXP3 in patients suffering from Crohn’s disease
Based on the suggested counter-suppressive activity of FOXP3Δ2Δ7, we hypothesized that lack of FOXP3 exon 7 could contribute to the pathogenesis of chronic inflammatory diseases. To determine the role of exon 7 loss in the pathogenesis of chronic inflammatory disease, we examined the expression of FOXP3 splice variants in patients suffering from Crohn’s disease by real time PCR using primers targeting exon/exon boundaries of exon 2 and exon 7. This allowed us to distinguish between FOXP3 mRNA containing exon 2 (FOXP3ex1/2; i.e. FOXP3fl and FOXP3Δ7), FOXP3 mRNA lacking exon 2 (FOXP3ex1/3; i.e. FOXP3Δ2 and FOXP3Δ2Δ7) and FOXP3 mRNA lacking exon 7 (FOXP3ex6/8; i.e. FOXP3Δ7 and FOXP3Δ2Δ7). According to this nomenclature the primer sets can specifically amplify mRNA molecules when the listed exons are adjacent to each other. To adequately compare the abundance of the different splice variants, we determined and compensated for the efficiency of the different primer sets (Supplementary Fig. 1). The total amount of FOXP3 mRNA was calculated as the sum of FOXP3ex1/2 and FOXP3ex1/3 mRNA, which allowed us to determine the relative proportion of a specific splicing event.
The total amount of FOXP3 mRNA was 0.11 ± 0.026 arbitrary units (mean ± SEM, n = 11) in Crohn’s disease patients, which did not significantly differ (two-tailed unpaired Student’s t test) from the levels found, 0.095 ± 0.018 (mean ± SEM, n = 10), in healthy donors. We found that the absolute amount (Fig. 1b) and relative abundance of FOXP3ex6/8 mRNA (Fig. 1c), but not FOXP3ex1/2 or FOXP3ex1/3 was greater in Crohn’s disease patients than in healthy donors. This increased expression of FOXP3ex6/8 mRNA correlated with increased disease severity (Fig. 1d). Furthermore, patients with Crohn’s disease displayed slightly decreased proportions of FOXP3ex6/8 when successfully treated with anti-TNF-α antibodies (Fig. 1e).
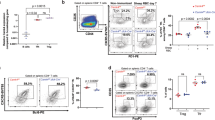
Increased alternative splicing of FOXP3 exon 7 in Crohn’s disease.
(a) Schematic overview of FOXP3 depicting alternatively spliced exons, epitopes of antibody clones and binding sites for primers used for detection of each splice variant. (b) Real-time PCR quantification of FOXP3 transcripts expressing exon 2 (FOXP3ex1/2), lacking exon 2 (FOXPex1/3) and lacking exon 7 (FOXP3ex6/8) in PBMCs obtained from Crohn’s disease patients (n = 8) or healthy donors (n = 10). (c) Percentage of FOXP3ex6/8 transcripts in relation to total FOXP3 transcripts in PBMCs (n = 11) of Crohn’s disease patients and healthy donors (n = 10). (d) Crohn’s disease colon biopsies (n = 29) stratified into 50% lowest and highest FOXP3ex6/8 expression samples plotted versus clinical score. (e) Percentage of FOXP3ex6/8 transcripts in relation to total FOXP3 in intestinal biopsies obtained from Crohn’s disease patients (n = 7) before and after successful anti-TNF-α treatment. (b–e) Data represent one pooled experiment with n biological replicates and are presented as (b,d,e) mean ± SD, (c) median ± IQR. P < 0.05 was considered significant ((b,d) two-tailed unpaired Student’s t test, (c) Kruskal-Wallis ANOVA and Dunn’s post hoc test, (e) two-tailed paired Student’s t test).
IL-1β promotes excision of FOXP3 exon 7
Having determined that patients suffering from Crohn’s disease had an increased frequency of exon 7 splicing in FOXP3 mRNA, we went on to identify factors that modulate the alternative splicing of FOXP3 in human Treg cells. To examine whether activation of Treg cells altered the balance of FOXP3 isoforms we compared the expression of FOXP3 transcripts between freshly isolated Treg cells and Treg cells activated with anti-CD3 and IL-2. We observed that FOXP3ex1/2 and FOXP3ex1/3 mRNA, but not FOXP3ex6/8 mRNA, were upregulated upon activation (Fig. 2a). We reasoned that signals promoting immune responses might alter the splicing of FOXP3 mRNA molecules to modulate their proinflammatory actions. Therefore, we addressed whether proinflammatory cytokines could modify the expression of FOXP3 splice variants in Treg cells. The expression of amount of total FOXP3, FOXP3ex1/2 and FOXP3ex1/3 mRNA were unchanged in response to TCR stimulation regardless of the presence of IL-1β, IL-6 or TNF-α (Fig. 2b–d, data not shown). Interestingly, although FOXP3ex6/8 mRNA was unchanged in response to TCR stimulation combined with IL-6 or TNF-α (Fig. 2c–d), it was increased in response to TCR stimulation supplemented with IL-1β (Fig. 2b).
Activation and IL-1β regulate alternative splicing of FOXP3.
Quantitative PCR was used to analyze: (a) fold induction of FOXP3 transcripts in CD4+CD25hiCD127low cells activated with plate bound α-CD3 and soluble α-CD28 and IL-2 for 18 hours relative to freshly isolated cells. (b-d) Fold induction of FOXP3 transcripts in CD4+CD25hiCD127low cells activated as above in the presence of 10 ng/ml IL-1β, IL-6 or TNF-α relative to cells activated without cytokines. (e) FOXP3ex1/2 (dark gray), FOXP3ex1/3 (gray) and FOXP3ex6/8 (white) transcripts in CD4+ T cells sorted for low, intermediate (int) and high (hi) expression of CD25. (a,b) Data are representative of four independent experiments and presented as mean ± SD. P < 0.05 was considered significant (two-tailed unpaired Student’s t test). (c) Data (n = 3 technical replicates) are presented as mean ± SD.
It is also possible that subsets of Treg cells differ in their expression of FOXP3 splice variants. To address this possibility we isolated CD4+ T cells based on their expression of CD25, a component of the high affinity receptor for IL-2 and analyzed the expression of FOXP3 splice variants using qPCR. We found that the total amount of FOXP3 mRNA correlated with the degree of CD25 expression, but no difference in FOXP3 splice variant distribution was apparent (Fig. 2e). Taken together, these results indicate that environmental cues regulate alternative splicing of FOXP3 and consequently the function of FOXP3.
Increased splicing frequency of FOXP3 exon 7 promotes Th17 differentiation
Previous studies have demonstrated that FOXP3Δ2Δ7 is incapable of conferring suppressive ability to T cells19. However, FOXP3 is not only expressed by Treg cells but it is also transiently expressed during Th17 differentiation. Because IL-1β promotes both alternative splicing of FOXP3 and Th17 differentiation, we next assessed the ability of FOXP3 isoforms to modulate Th17 differentiation. We purposely did not use overexpression of FOXP3 isoforms as T cell activation induces relatively high levels of endogenous FOXP3fl and FOXP3Δ2. Instead, we altered the splicing pattern of FOXP3 using morpholino antisense oligonucleotides (MAO) that prevent splice-directing small nuclear ribonucleoproteins from binding the exon/exon boundaries of FOXP3 pre-mRNA21. We demonstrated that it is possible to remove FOXP3 exon 2 in Treg cells with great efficiency (Fig. 3a) but the MAO targeting exon 7 was less efficient and could only partially remove FOXP3 exon 7 in Treg cells as determined by qPCR (data not shown). On the other hand MAO targeting efficiently removed FOXP3 exon 7 in naïve T cells upon differentiating into Th17 cells, presumably because MAO transfection precedes the induction of FOXP3 mRNA expression and the total amount of FOXP3 mRNA is much lower in these cells (Fig 3b). MAO-mediated splice shifting was observed as early as 24 hours post-transfection and persisted for at least 1 week (data not shown). Importantly, in these cells we found that expression of IL-2 (Fig. 3c) and IL-17A (Fig. 3d), but not IFN-γ (data not shown), is modulated by alternative splicing of FOXP3. Indeed, the combined removal of exon 2 and exon 7 mediated by MAO treatment strongly enhanced expression of IL-2 and IL-17A (Fig. 3c,d).
FOXP3Δ2Δ7 promote IL-17A production in naïve T cells.
(a) Density plots of FOXP3 expression (total and exon 2) in enriched CD25+CD4+ T cells (n = 10), that had been transfected with control MAO (control), or with splice-redirecting MAO specific for FOXP3 exon 2 (MAO Δ2) and/or MAO specific for FOXP3 exon 7 (MAO Δ7). (b) Real-time PCR quantification of FOXP3ex1/2 (black), FOXP3ex1/3 (gray) and FOXP3ex6/8 (white) transcripts of naïve T cells transfected with control MAO or MAO Δ2 and MAO Δ7 (n = 10). Expression was normalized to HPRT-1 and transcription ratio was calculated relative to total FOXP3. (c,d) MAO transfected CD4+ Naïve T cells were differentiated towards the Th17 lineage for 5 days and (c) IL-2 (n = 7) and (d) IL-17A (n = 10) cytokine expression were assessed by flow cytometry. Data are presented as mean ± SD, P<0.05 was considered significant,two-tailed paired Student’s t test.
Increased splicing frequency of FOXP3 exon 7 correlate with IL-17A expression in vivo
Based on research with murine cells indicating that exon 2 of Foxp3 binds to and antagonizes ROR-γt22,23, it has been suggested that FOXP3Δ2 may be linked to increased levels of IL-17 production in humans. However, a recent study with patients suffering from inflammatory bowel disease could not verify such an association24. Here, we analyzed biopsies obtained from Crohn’s disease patients and determined the levels of FOXP3 mRNA splice variants and cytokine mRNA expression. In accordance with previous studies we found no correlation between FOXP3ex1/3 and IL-17A expression. However, we observed a significant positive correlation between the levels of FOXP3ex6/8 mRNA and IL-17A mRNA (Fig. 4). Furthermore, no correlations were evident between FOXP3 splice variants and IFN-γ (data not shown), suggesting that alternative splicing of FOXP3 specifically affects Th17 differentiation. Altogether, these results suggest that FOXP3Δ2Δ7 favors the differentiation of naïve T cells towards Th17 cells in the setting of Crohn’s disease.
Alternative splicing of FOXP3 exon 7 correlates with IL-17A expression.
Quantitative PCR was used to measure FOXP3 and IL-17A transcript levels in colon biopsies from patients suffering from Crohn’s disease and normalized to GAPDH expression (n = 26). Data are presented as mean of n = 3 technical replicates. P < 0.05 was considered significant, Spearman rank correlation test; r = correlation coefficient).
Discussion
In this study, we demonstrate that patients suffering from Crohn’s disease display an irregular pattern of FOXP3 splicing with an increased proportion of FOXP3 transcripts lacking exon 7. We also found that the excision of FOXP3 exon 7 is controlled by the proinflammatory cytokine IL-1β and show that this FOXP3 splicing event promotes the differentiation of naïve T cells into Th17 cells.
Alternative splicing is the process by which exons of RNA are reconnected in multiple ways during RNA splicing25. Its is a major contributor to transcriptome and proteome diversity and recent studies indicate that up to 95% of human pre-mRNAs that contain more than one exon are processed to yield multiple mRNAs26. There are now several prominent examples of regulation of immune responses through alternative splicing27. For example, MyD88 an adaptor protein involved in Toll like receptor signaling, is found in two different isoforms with opposing function, where the MyD88L isoform activates the innate immune responses and the MyD88S isoform that lacks exon 2 inhibits immune responses28,29. In a similar manner activated T cells produce a soluble form of the common γ-chain, where exon 6 have been excised from the mature mRNA transcript, resulting in a new 9-amino-acid epitope followed by a stop codon. This soluble form of the common γ-chain inhibits cytokine signaling and consequently opposes the function of the full-length common γ-chain30,31. These and other examples of isoforms having opposing functions may illustrate that alternative splicing is a fast approach for turning off a biological response, as a dominant negative form can interfere with the function of already translated proteins that otherwise would continue to function.
Many questions remain regarding the function of FOXP3Δ2Δ7, however it is becoming increasingly clear that this isoform is unable to confer suppressive ability to Treg cells. Post-translational modifications, such as acetylation of the lysine-rich region in exon 7, have been proposed to stabilize FOXP3. However, when analyzing MAO-treated Treg cells treated with the proteasome inhibitor MG132 it appeared that FOXP3Δ2Δ7 is as stable as the other FOXP3 isoforms (data not shown). FOXP3Δ2Δ7 does however maintain the ability to bind DNA (data not shown) and presumably acts in a dominant negative manner by displacing FOXP3fl and FOXP3Δ2. The arguably most important aspect of this study is that we were able to identify that IL-1β promotes the excision of exon 7 of FOXP3. This gives us a valuable tool when we in the future want to study the molecular mechanisms of FOXP3 splicing. It also illustrates a novel mechanism as how a proinflammatory environment can tune the function of Treg cells through alternative splicing of FOXP3. A previous study has shown increased mRNA level of FOXP3Δ2Δ7 in patients suffering from rheumatoid arthritis37. Thus it would appear that altered splicing of FOXP3 mRNA is shared among several distinct inflammatory disorders, which is expected considering that IL-1β expression is a prominent feature during inflammation. However before we are able to fully elucidate the function of different FOXP3 isoforms, we need a new set of tools that will allow us to quantify FOXP3 isoforms on a protein level. While this additional layer of analysis could initially prove cumbersome, it may also resolve controversies where expression of FOXP3 does not correlate with an expected anti-inflammatory phenotype.
The intestinal immune system has to provide an effective immune response against pathogenic bacteria while maintaining tolerance towards food and commensal flora32,33 and T helper cells control the type of immune response that is mounted. Therefore it is not surprising that inflammatory bowel diseases are characterized by the excessive activation of certain T helper subsets such as Th1 and Th17. Recent genome-wide association studies have demonstrated that a number of genes involved in Th17 differentiation/function (IL23R, IL12B, JAK2, STAT3, CCR6 and TNFSF15) are associated with susceptibility to Crohn’s disease34,35,36,37,38. Treg cells are found in increased numbers in the intestinal mucosa of patients suffering from Crohn’s disease39, but appear unable to break the chronic inflammatory state. This could in part be due to increased expression of FOXP3∆2∆7, which is unable to confer a suppressive phenotype to Treg cell in vitro19. While several studies demonstrated that Treg cells from inflammatory bowel disease patients are functional40,41,42, there are also suggestions that these cells can alter their lineage commitment in response to the extracellular environment. In fact, patients with inflammatory bowel disease exhibit higher prevalence of circulating IL-17 and FOPX3 double positive CD4+ T cells43. In this study we did not assess the impact of FOXP3∆2∆7 on Treg cell plasticity, however, our study could potentially provide a molecular explanation for such Treg cell plasticity. Previous studies have demonstrated a key role of IL-1β as Helios-FOXP3+ Treg cells downregulate their suppressive functions in response to IL-1β44. In addition, Treg cells exposed to both IL-1β and IL-2 differentiate into proinflammatory Th17 cells45. It is undoubtedly a field that merits further investigations in order to establish a functional link between inflammation and autoimmunity.
TGF-β regulates the differentiation of both Treg cells and Th17 cells by inducing transient expression of both FOXP3 and RORγt. Foxp3 directly binds to and antagonizes ROR-γt22,23. The binding and inhibition of ROR-γt were to a large extent dependent on exon 2 of Foxp3. In addition, Zhou et al. also noted that knockdown of Foxp3 during Th17 differentiation resulted in an increase in Th17 cells23. Here, we determined that preferential expression of the FOXP3∆2∆7 isoform facilitated Th17 differentiation, which agrees with the latter finding, as FOXP3∆2∆7 inhibits the function of FOXP3fl and FOXP3∆2 in a dominant-negative manner. Our finding that FOXP3∆2∆7 facilitates Th17 differentiation is further supported by the correlation between FOXP3∆2∆7 expression and IL-17 (but not IFN-γ) expression in patients suffering from Crohn’s disease. By contrast, a recent study by Lord et al. suggests that no such correlation exists between FOXP3∆2 expression and IL-17 production in patients suffering from inflammatory bowel disease24. Thus, FOXP3 isoforms may regulate Th17 differentiation in two distinct ways: (a) by FOXP3fl directly binding to and antagonizing ROR-γt or (b) by general inhibition of FOXP3 function through upregulation of the dominant negative isoform FOXP3∆2∆7. The ability of IL-1β to promote Th17 differentiation is only partially conserved across species. Chung et al. have demonstrated that IL-1 receptor 1 expression in T cells, which is induced by IL-6 signaling, is necessary for early Th17 cell differentiation in vivo in mice46. Taking into account that alternative splicing of FOXP3 does not occur in mice, the latter study strongly supports the hypothesis that IL-1β modulates Th17 differentiation in more ways than simply inducing alternative splicing of FOXP3.
In summary, our study highlights the importance of characterizing FOXP3 expression on an isoform basis, as different FOXP3 isoforms are differentially regulated and exhibit distinct functional characteristics. Splicing of FOXP3 may be an important physiological regulator of Treg cell function and T-cell lineage commitment. Importantly, we found that the proinflammatory cytokine IL-1β promotes excision of exon 7 of FOXP3 by alternative splicing resulting in increased Th17 polarization. As such, our results shed new light on the mechanisms underlying chronic inflammatory diseases, such as Crohn’s disease, providing new pathways that could be targeted for treatment of these diseases.
Material and Methods
Patient Samples
Peripheral blood mononuclear cells (PBMCs) and biopsies from affected areas of the rectum and sigmoid colon were obtained from patients with Crohn’s disease. Disease activity was graded according to the simplified endoscopic activity score for Crohn’s disease as previously described47. A subset of patients was treated with the anti-TNF-α antibodies, infliximab or adalimumab (Remicade® or Humira®, respectively). Patients’ response to treatment was assessed using the Harvey–Bradshaw Index48. Patients with a clinical index activity decrease by ≥3 points were regarded as responders. The choice of treatment for individual patient was based on clinical evaluation without any intervention by the study. Anti-TNF-α treatment was administered either as infusions of 5 mg/kg infliximab at weeks 0, 2 and 6 or as subcutaneous injections of 80 mg adalimumab at week 0 followed by 40 mg every other week.
Ethical considerations
The study was approved by the Ethical Committee of Northern Stockholm and written informed consent was obtained from all participants. All experiments were performed in accordance with relevant guidelines and regulations.
Antibodies
The antibody clones used throughout this study are listed in Supplementary Table I.
Isolation of T cells
PBMCs were isolated from buffy coats using Ficoll-Paque Plus gradient centrifugation (GE Healthcare). CD4+ T cells were enriched from PBMCs using positive selection on an AutoMACS Separator with human CD4 micro-beads (Miltenyi Biotec). The enriched CD4+ T cells were then stained with antibodies recognizing CD4, CD25, CD127 (Supplementary Table I). CD4+CD25hiCD127low Treg cells were sorted using a FACSJazz instrument (BD Biosciences). Untouched naive CD4+ T cells were isolated from PBMCs by depleting non-T helper cells and memory CD4+ T cells using naïve CD4+ T cell Isolation Kit II according to the manufacturer’s instructions (Miltenyi Biotec). The enriched CD4+ T cells were then stained with antibodies recognizing CD4, CD25, CD45RA and CD62L (Supplementary Table I). Highly purified naïve T cell populations (>95%) were obtained by sorting for CD4+CD25−CD45RA+CD62L− cells using a FACSJazz instrument.
Cell culture and Th17 differentiation in vitro
T cells were activated and grown in X-VIVO 15 Medium supplemented with 1% Penicillin-Streptomycin (both from Lonza), 5 μg/ml plate-bound α-CD3, 1 μg/ml soluble α-CD28 (both from Biolegend), 300 U/ml IL-2 (Preprotech) and 6% CO2 at 37 °C. Naïve T cells were differentiated towards the Th17 lineage by culturing them with 10 ng/ml TGF-β, 10 ng/ml IL-1β, 25 ng/ml IL-6 and 10 ng/ml IL-23 (all from Peprotech) for 6 days. Four hours prior to harvesting, cells were restimulated with 50 ng/ml PMA (Sigma) and 1 μg/ml ionomycin (Life Technologies) and treated with GolgiBlocker (BD Biosciences).
Splice-shifting
Enhanced FOXP3 exon splicing was achieved using Fluorescein-labelled Morpholino Antisense Oligonucleotides (MAO) with the following sequences: 5′-TGCCCATTCACCGTCCATACCTGGT-3′ for FOXP3ex1/3, 5′-AGCTGTGAAATGGCACAAACATGAG-3′ for FOXP3ex6/8 and 5′-CCTCTTACCTCAGTTACAATTTATA-3′ as a control (GeneTools). Prior to activation, T cells were transfected with 15 μM MAO using the P3 Primary Cell Nucleofector Kit and a Nucleotransfector device (Lonza) according to the manufacturer’s instructions.
FACS
Single-cell suspensions were stained with LIVE/DEAD Fixable Dead Cell Stain Kit (Life Technologies) to identify dead cells. Intracellular staining was performed using eBioscience’s FOXP3 staining kit according to the manufacturer’s instructions. Data was acquired on a LSRFortessa flow cytometer (BD Biosciences) and analyzed with FlowJo Version 7.6.4 software for Mac (TreeStar).
Quantitative PCR
Total RNA from T cells was isolated with Trizol (Life Technologies) and cDNA was generated using Vilo cDNA Synthesis Kit (Life Technologies). Amplification was performed with iQ SYBR Green Supermix (Bio-Rad) with the following protocol: 2 min 95 °C, (15 sec 95 °C, 45 sec 58 °C, 30 sec 68 °C) × 39. Gene specific primer pairs are listed in Supplementary Table II. Threshold cycles (Ct) calculated by Bio-Rad CFX software were normalized to the expression of GAPDH (ex vivo samples) or HPRT1 (cell culture samples). Relative amounts were calculated with [c] = 2ΔCt, changes in expression levels with [fold] = 2−ΔΔCt. Primer specificity in all samples was confirmed by single peak performances of PCR products in melt curve analysis. Expression of FOXP3 splice variants was calculated with respect to their individual primer pair efficiency (Supplementary Fig. 1). Total FOXP3 mRNA expression was calculated as the sum of FOXP3ex1/2 and FOXP3ex1/3 and splice variant percentage was calculated as (FOXP3 variant/total FOXP3)*100.
Additional Information
How to cite this article: Mailer, R. K. W. et al. IL-1b promotes Th17 differentiation by inducing alternative splicing of FOXP3. Sci. Rep. 5, 14674; doi: 10.1038/srep14674 (2015).
References
Josefowicz, S. Z., Lu, L. F. & Rudensky, A. Y. Regulatory t cells: Mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of cd4+cd25+ regulatory t cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory t cell development by the transcription factor foxp3. Science . 299, 1057–1061 (2003).
Khattri, R., Cox, T., Yasayko, S. A. & Ramsdell, F. An essential role for scurfin in cd4+cd25+ t regulatory cells. Nat. Immunol. 4, 337–342 (2003).
Brunkow, M. E. et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 27, 68–73 (2001).
Bennett, C. L. et al. The immune dysregulation, polyendocrinopathy, enteropathy, x-linked syndrome (ipex) is caused by mutations of foxp3. Nat. Genet. 27, 20–21 (2001).
Wildin, R. S. et al. X-linked neonatal diabetes mellitus, enteropathy and endocrinopathy syndrome is the human equivalent of mouse scurfy. Nat. Genet. 27, 18–20 (2001).
Taams, L. S. et al. Regulatory t cells in human disease and their potential for therapeutic manipulation. Immunology 118, 1–9 (2006).
Chae, W. J., Henegariu, O., Lee, S. K. & Bothwell, A. L. The mutant leucine-zipper domain impairs both dimerization and suppressive function of foxp3 in t cells. P. Natl. Acad. Sci. USA . 103, 9631–9636 (2006).
Bettelli, E., Dastrange, M. & Oukka, M. Foxp3 interacts with nuclear factor of activated t cells and nf-kappa b to repress cytokine gene expression and effector functions of t helper cells. P. Natl. Acad. Sci. USA . 102, 5138–5143 (2005).
Lopes, J. E. et al. Analysis of foxp3 reveals multiple domains required for its function as a transcriptional repressor. J. Immunol. 177, 3133–3142 (2006).
Wu, Y. et al. Foxp3 controls regulatory t cell function through cooperation with nfat. Cell. 126, 375–387 (2006).
Deng, G. et al. Molecular and biological role of the foxp3 n-terminal domain in immune regulation by t regulatory/suppressor cells. Exp. Mol. Pathol. 93, 334–338 (2012).
Li, B. et al. Foxp3 is a homo-oligomer and a component of a supramolecular regulatory complex disabled in the human xlaad/ipex autoimmune disease. Int. Immunol. 19, 825–835 (2007).
Song, X. et al. Structural and biological features of foxp3 dimerization relevant to regulatory t cell function. Cell Rep . 1, 665–675 (2012).
Bandukwala, H. S. et al. Structure of a domain-swapped foxp3 dimer on DNA and its function in regulatory t cells. Immunity 34, 479–491 (2011).
Aarts-Riemens, T., Emmelot, M. E., Verdonck, L. F. & Mutis, T. Forced overexpression of either of the two common human foxp3 isoforms can induce regulatory t cells from cd4(+)cd25(–) cells. Eur. J. Immunol. 38, 1381–1390 (2008).
Allan, S. E. et al. The role of 2 foxp3 isoforms in the generation of human cd4+ tregs. J. Clin Invest. 115, 3276–3284 (2005).
Mailer, R. K., Falk, K. & Rotzschke, O. Absence of leucine zipper in the natural foxp3delta2delta7 isoform does not affect dimerization but abrogates suppressive capacity. PloS One 4, e6104 (2009).
Harbuz, R. et al. Identification of new foxp3 mutations and prenatal diagnosis of ipex syndrome. Prenatal Diag. 30, 1072–1078 (2010).
Kole, R., Krainer, A. R. & Altman, S. Rna therapeutics: Beyond rna interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Ichiyama, K. et al. Foxp3 inhibits rorgammat-mediated il-17a mrna transcription through direct interaction with rorgammat. J. Biol. Chem. 283, 17003–17008 (2008).
Zhou, L. et al. Tgf-beta-induced foxp3 inhibits t(h)17 cell differentiation by antagonizing rorgammat function. Nature 453, 236–240 (2008).
Lord, J. D., Valliant-Saunders, K., Hahn, H., Thirlby, R. C. & Ziegler, S. F. Paradoxically increased foxp3+ t cells in ibd do not preferentially express the isoform of foxp3 lacking exon 2. Digest. Dis. Sci. 57, 2846–2855 (2012).
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by rna-binding proteins. Nat. Rev. Genet. 15, 689–701 (2014).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Martinez, N. M. & Lynch, K. W. Control of alternative splicing in immune responses: Many regulators, many predictions, much still to learn. Immunol. Rev. 253, 216–236 (2013).
Burns, K. et al. Inhibition of interleukin 1 receptor/toll-like receptor signaling through the alternatively spliced, short form of myd88 is due to its failure to recruit irak-4. J. Exp. Med. 197, 263–268 (2003).
De Arras, L. & Alper, S. Limiting of the innate immune response by sf3a-dependent control of myd88 alternative mrna splicing. PLoS Genet. 9, e1003855 (2013).
Meissner, U., Blum, H., Schnare, M., Rollinghoff, M. & Gessner, A. A soluble form of the murine common gamma chain is present at high concentrations in vivo and suppresses cytokine signaling. Blood. 97, 183–191 (2001).
Hong, C. et al. Activated t cells secrete an alternatively spliced form of common gamma-chain that inhibits cytokine signaling and exacerbates inflammation. Immunity 40, 910–923 (2014).
Geremia, A., Biancheri, P., Allan, P., Corazza, G. R. & Di Sabatino, A. Innate and adaptive immunity in inflammatory bowel disease. Autoimmun. Rev. 13, 3–10 (2014)
Monteleone, I., Sarra, M., Pallone, F. & Monteleone, G. Th17-related cytokines in inflammatory bowel diseases: Friends or foes? Cur. Mol. Med. 12, 592–597 (2012).
Duerr, R. H. et al. A genome-wide association study identifies il23r as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Cargill, M. et al. A large-scale genetic association study confirms il12b and leads to the identification of il23r as psoriasis-risk genes. Am. J. Hum Genet. 80, 273–290 (2007).
Wellcome Trust Case Control C, et al. Association scan of 14,500 nonsynonymous snps in four diseases identifies autoimmunity variants. Nat. Genet. 39, 1329–1337 (2007).
Barrett, J. C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for crohn’s disease. Nat. Genet. 40, 955–962 (2008).
Fisher, S. A. et al. Genetic determinants of ulcerative colitis include the ecm1 locus and five loci implicated in crohn’s disease. Nat. Genet. 40, 710–712 (2008).
Chamouard, P. et al. Diminution of circulating cd4+cd25 high t cells in naive crohn’s disease. Digest. Dis. Sci. 54, 2084–2093 (2009).
Maul, J. et al. Peripheral and intestinal regulatory cd4+ cd25(high) t cells in inflammatory bowel disease. Gastroenterology 128, 1868–1878 (2005).
Saruta, M. et al. Characterization of foxp3+cd4+ regulatory t cells in crohn’s disease. Clin. Immunol. 125, 281/290 (2007).
Eastaff-Leung, N., Mabarrack, N., Barbour, A., Cummins, A. & Barry, S. Foxp3+ regulatory t cells, th17 effector cells and cytokine environment in inflammatory bowel disease. J. Clin. Immunol. 30, 80–89 (2010)
Ueno, A. et al. Increased prevalence of circulating novel il-17 secreting foxp3 expressing cd4+ t cells and defective suppressive function of circulating foxp3+ regulatory cells support plasticity between th17 and regulatory t cells in inflammatory bowel disease patients. Inflamm. Bowel Dis. 19, 2522–2534 (2013).
Raffin, C. et al. Human memory helios- foxp3+ regulatory t cells (tregs) encompass induced tregs that express aiolos and respond to il-1beta by downregulating their suppressor functions. J. Immunol. 191, 4619–4627 (2013).
Deknuydt, F., Bioley, G., Valmori, D. & Ayyoub, M. Il-1beta and il-2 convert human treg into t(h)17 cells. Clin. Immunol. 131, 298–307 (2009).
Chung, Y. et al. Critical regulation of early th17 cell differentiation by interleukin-1 signaling. Immunity 30, 576–587 (2009).
Daperno, M. et al. Development and validation of a new, simplified endoscopic activity score for crohn’s disease: The ses-cd. Gastrointest. Endosc. 60, 505–512 (2004).
Harvey, R. F. & Bradshaw, J. M. A simple index of crohn’s-disease activity. Lancet 1, 514 (1980).
Acknowledgements
We thank Ludvig Linton and the staff at the Gastroenterology Clinic at Danderyd’s hospital for help with patient samples. This work was supported by the Swedish Research Council (projects: 2008–3055 and 2012-1851), the Swedish Cancer Foundation, the Swedish Heart-Lung Foundation, the Swedish Society of Medicine and Sigurd and Elsa Goljes minne.
Author information
Authors and Affiliations
Contributions
R.K.W.M., A.-L.J., S.L., S.E., J.T. and J.A. designed the research; R.K.W.M., A.-L.J., S.L. and S.E. performed experiments; J.T. and J.A. supervised the study; R.K.W.M. and J.A. wrote the manuscript and all authors discussed the results and their implications and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mailer, R., Joly, AL., Liu, S. et al. IL-1β promotes Th17 differentiation by inducing alternative splicing of FOXP3. Sci Rep 5, 14674 (2015). https://doi.org/10.1038/srep14674
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14674
This article is cited by
-
Pyroptosis: the potential eye of the storm in adult-onset Still’s disease
Inflammopharmacology (2023)
-
RNA metabolism and links to inflammatory regulation and disease
Cellular and Molecular Life Sciences (2022)
-
Single-cell differential splicing analysis reveals high heterogeneity of liver tumor-infiltrating T cells
Scientific Reports (2021)
-
Recent advances in the role of Th17/Treg cells in tumor immunity and tumor therapy
Immunologic Research (2021)
-
Cytokine “fine tuning” of enthesis tissue homeostasis as a pointer to spondyloarthritis pathogenesis with a focus on relevant TNF and IL-17 targeted therapies
Seminars in Immunopathology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.