Abstract
Proprotein convertase subtilisin/kexin type 2 (PCSK2) is a prohormone processing enzyme involved in insulin and glucagon biosynthesis. We previously found the genetic polymorphism of PCSK2 on chromosome 20 was responsible for the linkage peak of several glucose homeostasis parameters. The aim of this study is to investigate the association between genetic variants of PCSK2 and glucose homeostasis parameters and incident diabetes. Total 1142 Chinese participants were recruited from the Stanford Asia-Pacific Program for Hypertension and Insulin Resistance (SAPPHIRe) family study and 759 participants were followed up for 5 years. Ten SNPs of the PCSK2 gene were genotyped. Variants of rs6044695 and rs2284912 were associated with fasting plasma glucose and variants of rs2269023 were associated with fasting plasma glucose and 1-hour plasma glucose during OGTT. Haplotypes of rs4814605/rs1078199 were associated with fasting plasma insulin levels and HOMA-IR. Haplotypes of rs890609/rs2269023 were also associated with fasting plasma glucose, fasting insulin and HOMA-IR. In the longitudinal study, we found individuals carrying TA/AA genotypes of rs6044695 or TC/CC genotypes of rs2284912 had lower incidence of diabetes during the 5-year follow-up. Our results indicated that PCSK2 gene polymorphisms are associated with pleiotropic effects on various traits of glucose homeostasis and incident diabetes.
Similar content being viewed by others
Introduction
Development of type 2 diabetes (T2DM) is characterized by tissue resistance to insulin action and failure of pancreatic beta cells to secrete insulin to maintain glucose homeostasis1. Both genetic and environmental factors influence susceptibility to T2DM2. However, identification of the genes responsible for T2DM is complicated by the high degree of genetic heterogeneity, the involvement of multiple genes and a small to moderate risk conferred by each of the genes. Searching for quantitative trait loci (QTLs) that explain the variation in the “intermediate” phenotypes of T2DM has therefore been considered a plausible way to tease out the genetic factors involved in biological pathways that lead to development of diabetes3. So far, a number of genome-wide linkage scans have been carried out to identify the QTLs for the related intermediate phenotypes of T2DM in Pima Indian4, Mexican Americans5,6, Han Chinese7,8, Japanese Americans9, Caucasians10, European Americans and African Americans11 and multiethnic population12. A QTL for the age of onset of T2DM was identified in a set of French families13. However, the putative genetic variants for most of the reported QTLs are largely unknown, including ours7.
Calpain 10 (CAPN10)14, ectonucleotide pyrophosphatase phosphodiesterase 1 (ENPP1)15, hepatocyte nuclear factor 4α (HNF4A)16,17, adiponectin (ADIPOQ)18 and transcription factor 7-like 2 (TCF7L2)19 were previously identified in T2DM-linked chromosomal regions. Using a genome-wide association approach, more than 60 genetic loci to date have been identified for T2DM20,21. However, the overall fraction of these newly identified T2DM genes remains small, accounting for 5–10% of the heritability of T2DM22.
We previously identified a QTL located at 37 cM on chromosome 20 for the fasting insulin and insulin resistance index by homeostasis model assessment (HOMA-IR) in 1,365 non-diabetic Chinese subjects from 411 nuclear families7. Following subsequent fine mapping, we found the genetic polymorphism of proprotein convertase subtilisin/kexin type 2 (PCSK2) was responsible for the linkage peak of fasting insulin, HOMA-IR and glucose levels at 1 hr after 75 gm oral glucose loading (data not shown). Therefore, in this study we focused on PCSK2.
The protein coded by this gene, PCSK2, is a prohormone processing enzyme that plays a key role in regulating insulin and glucagon biosynthesis. PCSK2 is expressed in the brain and the pancreatic islets. Variants of the PCSK1 and PCSK2 genes previously have been linked to T2DM and obesity23,24,25,26,27. The aim of the current study is to investigate the association between genetic variants of PCSK2 and fasting insulin and glucose concentration, the homeostasis model assessment of beta cell function (HOMA-beta) and HOMA-IR, various parameters during the 75 g oral glucose tolerance test (OGTT) and clinical progression from normal glucose tolerance to diabetes after a follow-up of 5 years.
Results
Characteristics of the SAPPHIRe Chinese participants at baseline and after 5-yr follow- up
A total of 1144 Chinese participants were recruited from the SAPPHIRe study at baseline and 759 of them received 5-year follow-up examinations. The anthropometric characteristics, plasma glucose and insulin concentration during OGTT, HOMA-IR and HOMA-beta at baseline and after a 5-year follow-up are shown in Table 1.
Association of PCSK2 genetic variants with various traits of glucose homeostasis
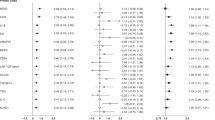
The SNPs and their location in the PCSK2 gene, their position in the physical map and minor allele frequency are shown in Supplementary Table S1. All the following traits of glucose homeostasis were adjusted for age, gender and body mass index (BMI) and analyzed by family-based association test (FBAT). Genetic variants of rs6044695 and rs2284912 were negatively associated with fasting plasma glucose (FPG) concentration. Genetic variants of rs2269023 were positively associated with FPG and 1-hour plasma glucose concentration (1 h-PG) during OGTT (Table 2). All associations with a q value < 0.05 were considered statistically significant.
Association of SNP haplotypes of PCSK2 with various traits of glucose homeostasis
After analysis with Haploview 4.1, the 10 tag SNPs of PCSK2 were divided into 2 haplotype blocks (block 1: rs4814605/rs1078199, block 2: rs890609/rs2269023, Fig. 1). Haplotypes of rs4814605/rs1078199 (block 1) were associated with fasting plasma insulin concentration (FINS) and HOMA-IR. Haplotypes of rs890609/rs2269023 (block 2) were associated with FPG, FINS and HOMA-IR (Table 3). Both the haplotype-specific P value and global P value were derived from permutation testing 10,000 times. A null hypothesis was rejected if the permuted global P value was <0.05.
Haploview LD graph of the PCSK2 gene (10 genotyped SNPs in this study).
Pairwise LD coefficients D′ × 100 are shown in each cell (D′ values of 1.0 are not shown). The standard color scheme of Haploview was used for the LD color display (logarithm of likelihood odds ratio [LOD] (a measure of confidence in the value of D′) ≥2 and D′ = 1, shown in bright red; LOD ≥ 2 and D′ < 1 shown in blue; LOD < 2 and D′ = 1 shown in pink; LOD < 2 and D′ < 1 shown in white).
Specific SNPs of PCSK2 were associated with progression from normoglycemia to diabetes during a 5-year follow-up
We further used the proportional hazard model to analyze whether the presence or absence of specific SNPs was associated with the progression from normoglycemia to diabetes during a 5-year follow-up. All the p values were adjusted for age, sex, center, drug, environmental factors (including smoking, drinking and sedentary lifestyle) and BMI. The individuals with TA or AA genotypes of rs6044695, or TC or CC genotypes of rs2284912 had a significantly lower incidence of diabetes (Table 4). As shown in Table 5, rs4814597, rs1609659, rs2208203 and rs2021785 were also associated with type 2 diabetes or glucose homeostatic traits according to GWAS database28. Since we did not have genotype data of the four SNPs in this study, we reexamined these additional established loci from GWAS in Table 5 by imputing their genotypes using MACH imputation package29,30 based on 1000 Genomes data. The results are presented in Supplementary Table S2 with a serial number starting with A. None of the imputed SNPs showed evidence of association with incident diabetes. The Haploview linkage disequilibrium (LD) graph of the PCSK2 gene (10 genotyped SNPs in this study and 4 imputed SNPs: rs4814597, rs1609659, rs2208203 and rs2021785) was shown in Supplementary Fig S1.
Discussion
In our study, significant associations between some SNPs as well as haplotypes of PCSK2 and various traits of glucose homeostasis, including FPG, 1 h-PG, FINS and HOMA-IR, were found. Furthermore, individuals with some specific SNPs of PCSK2 were also associated with progression to diabetes during a 5-year follow-up. In our previous study31, we reported the potential pleiotropy of the locus at 37 cM on chromosome 20 on each pair of traits, such as fasting insulin/HOMA-beta and HOMA-IR/HOMA-beta, which supports our present findings that PCSK2 gene polymorphisms are associated with pleiotropic effects on these metabolic variables.
PCSK2 is a type II proinsulin-processing enzyme and it cleaves the proinsulin molecule on the COOH-terminal side of dibasic peptide, Lys64-Arg65, which joins the C-peptide and A-chain domains32. Defects affecting the catalytic activity of the prohormone-processing enzymes have been found to be associated with obesity and other metabolic disorders33,34. The etiology of hyperproinsulinemia is thought to be pancreatic β cell dysfunction, which is manifested in part by inadequate cleavage of proinsulin. Previous studies have shown that increased concentrations of proinsulin are a significant predictor of the development of T2DM in several ethnic groups35,36,37,38. Furuta et al.39 reported that increased levels of proinsulin and split proinsulin were detected in pancreatic islet cells isolated from homozygous pcsk2 null mice.
There have been several studies reporting that genetic polymorphisms of PCSK2 were associated with either T2DM or various glucose homeostasis parameters (Table 5). A significant difference in the allele frequency distribution of a simple CA tandem-repeat DNA polymorphism (STRP) in intron 2 of PCSK2 has been reported in a case-control study of T2DM patients and normal controls in a Japanese population26 (Table 5). Jonssan et al. recently reported that the C allele of PCSK2 rs2208203 in intron 2 was associated with reduced insulin secretion measured as the corrected insulin response as well as disposition index40. The variant was also associated with lower fasting glucagon levels in non-diabetic individuals with FPG over 5.5 mmol/l40 (Table 5). The above microsatellite and rs2208203 in intron 2 were not examined in this study. According to imputation analysis based on 1000 Genomes data, rs2208203 was not associated with incident T2DM.
A more recent genome-wide association study (GWAS) on T2DM in African American families also showed linkage to chromosome 20p in a subset with a later age at diagnosis. The PCSK2 gene is within the 1-logarithm of odds (LOD) interval of this linkage peak. Association with T2DM was observed among 4 SNPs: rs2021785, rs1609659, rs4814597 and rs226902325 (Table 5). A recent report showed that an association of the risk allele of rs2021785 at PCSK2 with T2DM also existed in a Han Chinese population27 (Table 5). Rs2021785, rs1609659 and rs4814597 were not genotyped in this study. According to imputation analysis based on 1000 Genomes data, the above three imputed SNPs were not associated with incident T2DM. Consistently, in this study, rs2269023 was associated with FPG and 1-hour PG during OGTT in a non-diabetic Han Chinese population (Table 2). Therefore, rs2269023 may play an important role in the regulation of glucose homeostasis in different ethnic groups.
We further searched the open GWAS Central database28 for associations between the 10 genetic variations of PCSK2 investigated in this study and related metabolic phenotypes in Caucasian populations. Significant associations were found between rs2206447 and T2DM (P = 0.008, FUSION Study) and between rs6080705 and HOMA-beta (P = 0.008588), HOMA-IR (P = 0.02582) and fasting insulin (P = 0.01508) (https://www.gwascentral.org/, searched on 6.10.2014) (Table 5). However, these associations could not be replicated in a Han Chinese population in this study. Furthermore, genetic variants of rs6044695 and rs2284912 were associated with both baseline FPG and progression of T2DM during the 5-year follow-up in this study. Therefore, the association at baseline was also replicated in the longitudinal follow-up study. To the best of our knowledge, this is the first study to report that the genetic variants of PCSK2 were associated with incident T2DM.
This study has several strengths. First, our study used a family-based design, which is a systemic approach to capture all common genetic variations, to control for population stratification. Second, we adopted q-values as our measure of significance in order to reduce false-positive results derived from multiple tests. The q-value is an false-discovery rate (FDR)-based measure of significance used in genome-wide studies. Most importantly, a systematic use of q-values in genome-wide tests of significance will yield a clear balance of false-positive results to true-positive results and provide a standard measure of significance that can be universally interpreted41. Third, this study examined SNPs associated with incidence of diabetes rather than prevalence. The limited number of diabetes incidences would be the limitation of this study though.
In conclusion, several genetic variants and haplotypes of PCSK2 were associated with various traits of glucose homeostasis and progression to diabetes. These findings, together with several earlier observations in different ethnic groups, support an involvement of the PCSK2 gene in the pathogenesis of T2DM.
Methods
Study population of the SAPPHIRe study cohort
The Stanford Asia-Pacific Program for Hypertension and Insulin Resistance (SAPPHIRe) was a collaborative study that was part of the Family Blood Pressure Program of the National Heart, Lung and Blood Institute of the National Institutes of Health meant to investigate the genetic determinants of hypertension and insulin resistance in Chinese and Japanese. The study collected over 1,300 sib pairs that were either concordant or discordant for high blood pressure. Detailed descriptions of the study cohort were published in our previous work42,43. In brief, subjects were aged between 35 and 60 years and of Chinese or Japanese ancestry. Hypertension was defined as systolic blood pressure >160 mm Hg, diastolic blood pressure >95 mm Hg, or use of 2 medications for high blood pressure (stage II hypertension). Also, the subjects could be taking one medication for high blood pressure with a systolic blood pressure >140 mm Hg or a diastolic blood pressure >90 mm Hg. Low-normal blood pressure was defined as blood pressure in the bottom 30% of the age- and sex-adjusted blood pressure distribution. Individuals with chronic illnesses like diabetes, cancer, or diseases of the heart, liver, or kidney were excluded. In this study, 1142 Chinese participants were recruited from the SAPPHIRe study and 759 participants received a 5-year follow-up. The institutional review board of each participating site (National Taiwan University Hospital, Taipei Veterans General Hospital, Taichung Veterans General Hospital and Tri-Serve General Hospital) approved all the experiments in this study. Informed consent was obtained from all subjects.
Phenotyping
The participants underwent anthropometric measurements at 8 A.M. after an 8–10 h overnight fast. Each subject was subjected to a 75-g OGTT after the anthropometric measurements. Fasting blood samples were collected for the measurement of plasma glucose and insulin. Then, 75 g glucose monohydrate (in 300 ml water) was administered to the subject to drink within 5 minutes. Blood samples were taken for plasma glucose and insulin 1 and 2 hours after glucose loading. The patients were not allowed to eat or drink until the end of the test7. Plasma glucose and insulin levels were measured as described previously7. HOMA-IR and HOMA-beta derived from the homeostasis model were identical to the previous study7.
Selection of tagSNPs and genotyping
To identify common tagSNPs, we selected tagSNPs from the HapMap CHB (Han Chinese in Beijing) database (phase 1&2, build 35) (http://www.hapmap.org)44 using the Tagger program implemented in Haploview version 4 (http://www.broad.mit.edu/mpg/haploview/)45. Ten SNPs were selected with minor allele frequencies of more than 10% at r2 = 0.7 and that captured 80% of alleles of PCSK2. SNP genotyping was performed using the GenomeLab SNPstream genotyping platform (Beckman Coulter, Fullerton, CA) and its accompanying SNPstream software suite. ASPEX software was applied to examine Mendelian inconsistencies. When an error was found, the marker data were converted to missing; less than 1% of the marker data were converted to missing in this study. All the methods were carried out in accordance with the approved guidelines. All experimental protocols were approved by committee of National Taiwan University Hospital, Taipei Veterans General Hospital, Taichung Veterans General Hospital and Tri-Serve General Hospital.
Statistical analysis
All data were summarized as mean values ± S.D. unless otherwise specified. Pairwise linkage disequilibrium (LD) measures D′ and r2 were estimated to assess LD between SNPs in the PCSK2 gene. The structure of the haplotype block was evaluated using the confidence interval method developed by Gabriel et al. and implemented in the Haploview program45. The association of PCSK2 SNP and haplotypes with metabolic phenotypes was analyzed using the family-based association test (FBAT)46. The trait residuals were obtained based on the generalized linear models adjusted for age, gender, center, drug, environmental factors (i.e., smoking, drinking and sedentary lifestyle) and BMI, then imported into FBAT for association analysis. For each association, we derived a q-value41 that was calculated using the statistical package SAS version 9.1. The q-value has been proposed as a FDR-based measure of significance for multiple testing41. FDR is the expected proportion of Type I errors among the rejected hypotheses. Q-value is defined as an analog of the p-value that incorporates FDR-based multiple testing correction41. Namely, q-value is the minimum FDR that can be attained to reach significance (i.e., expected proportion of false positives incurred for significance). A p-value of 0.05 implies that 5% of all tests will result in false positives, while an FDR adjusted p-value (or q-value) of 0.05 implies that 5% of significant tests will result in false positives.
We also used the proportional hazard model to analyze whether the presence or absence of specific SNPs was associated with the progression from normoglycemia to diabetes during a 5-year follow-up. A null hypothesis was rejected if the q-value was <0.05. We presented the hazard ratio of the allelic effect from the major allele (A) for each SNP based on Cox regression models. A Cox regression model is a regression-based method for exploring the associations between survival data and explanatory variables. It provides an estimate of the hazard ratio and its confidence interval between two groups. In the present study, the survival data is the person-years for diabetes incidence during the 5-year follow-up period and the explanatory variable of interest is individual SNPs. Proportional hazards regression assumes the hazard ratio is constant over time. Therefore, we conducted Schoenfeld’s residuals test47 to check the proportional hazard assumption for each SNP. None of the proportional hazard assumption was rejected suggesting the assumption is legitimate for all the SNPs in the Cox regression analysis (Supplementary Table S3).
We obtained haplotype-specific and whole marker P-value by a permutation test. Ten-thousand times were permuted when analyzing family-based association test of PCSK2 haplotypes with various traits of glucose homeostasis. To calculate permutation-based P values, the phenotype labels are randomly shuffled and all the multiple tests are recalculated emperically on the reshuffled data set, with the smallest P value of these multiple tests. The procedure is repeated for 10,000 times to construct an empirical frequency distribution of the smallest P values. If the P value calculated for the actual data set is smaller than r of the 10,000 smallest P value from the permuted data sets, then an empirical adjusted P value (P*) is given by P* = (r + 1)/(n + 1), where n is the number of replicate samples that have been simulated and r is the number of these replicates that produce a test statistic greater than or equal to that calculated for the actual data. A null hypothesis was rejected if the permuted P value was <0.0548.
Additional Information
How to cite this article: Chang, T.-J. et al. Genetic polymorphisms of PCSK2 are associated with glucose homeostasis and progression to type 2 diabetes in a Chinese population. Sci. Rep. 5, 14380; doi: 10.1038/srep14380 (2015).
References
Saad, M. F. et al. A two-step model for development of non-insulin-dependent diabetes. Am J Med 90, 229–235 (1991).
Gerich, J. E. The genetic basis of type 2 diabetes mellitus: impaired insulin secretion versus impaired insulin sensitivity. Endocr Rev 19, 491–503 (1998).
Menzel, S. Genetic and molecular analyses of complex metabolic disorders: genetic linkage. Ann N Y Acad Sci 967, 249–257 (2002).
Pratley, R. E. et al. An autosomal genomic scan for loci linked to prediabetic phenotypes in Pima Indians. J Clin Invest 101, 1757–1764 (1998).
Mitchell, B. D. et al. Linkage of serum insulin concentrations to chromosome 3p in Mexican Americans. Diabetes 49, 513–516 (2000).
Cai, G. et al. Genome-wide scans reveal quantitative trait Loci on 8p and 13q related to insulin action and glucose metabolism: the San Antonio Family Heart Study. Diabetes 53, 1369–1374 (2004).
Chiu, Y. F. et al. An autosomal genome-wide scan for loci linked to pre-diabetic phenotypes in nondiabetic Chinese subjects from the Stanford Asia-Pacific Program of Hypertension and Insulin Resistance Family Study. Diabetes 54, 1200–1206 (2005).
Ng, M. C. et al. Genome-wide scan for metabolic syndrome and related quantitative traits in Hong Kong Chinese and confirmation of a susceptibility locus on chromosome 1q21-q25. Diabetes 53, 2676–2683 (2004).
Kovac, I. P. et al. Linkage and association analyses of type 2 diabetes/impaired glucose metabolism and adiponectin serum levels in Japanese Americans from Hawaii. Diabetes 56, 537–540 (2007).
Meigs, J. B., Panhuysen, C. I., Myers, R. H., Wilson, P. W. & Cupples, L. A. A genome-wide scan for loci linked to plasma levels of glucose and HbA(1c) in a community-based sample of Caucasian pedigrees: The Framingham Offspring Study. Diabetes 51, 833–840 (2002).
Freedman, B. I. et al. Genome-wide scans for heritability of fasting serum insulin and glucose concentrations in hypertensive families. Diabetologia 48, 661–668 (2005).
An, P. et al. Genome-wide linkage scans for fasting glucose, insulin and insulin resistance in the National Heart, Lung and Blood Institute Family Blood Pressure Program: evidence of linkages to chromosome 7q36 and 19q13 from meta-analysis. Diabetes 54, 909–914 (2005).
Cheyssac, C. et al. EIF4A2 is a positional candidate gene at the 3q27 locus linked to type 2 diabetes in French families. Diabetes 55, 1171–1176 (2006).
Horikawa, Y. et al. Genetic variation in the gene encoding calpain-10 is associated with type 2 diabetes mellitus. Nat Genet 26, 163–175 (2000).
Meyre, D. et al. Variants of ENPP1 are associated with childhood and adult obesity and increase the risk of glucose intolerance and type 2 diabetes. Nat Genet, 37, 863–867 (2005).
Love-Gregory, L. D. et al. A common polymorphism in the upstream promoter region of the hepatocyte nuclear factor-4 alpha gene on chromosome 20q is associated with type 2 diabetes and appears to contribute to the evidence for linkage in an Ashkenazi Jewish population. Diabetes 53, 1134–1140 (2004).
Silander, K. et al. Genetic variation near the hepatocyte nuclear factor-4 alpha gene predicts susceptibility to type 2 diabetes. Diabetes 53, 1141–1149 (2004).
Vasseur, F. et al. Single-nucleotide polymorphism haplotypes in the both proximal promoter and exon 3 of the APM1 gene modulate adipocyte-secreted adiponectin hormone levels and contribute to the genetic risk for type 2 diabetes in French Caucasians. Hum Mol Genet 11, 2607–2614 (2002).
Grant, S. F. et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet, 38, 320–323 (2006).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scalereplication identifies additional susceptibility loci for type 2 diabetes. Nat Genet 40, 638–645 (2008).
Voight, B. F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat Genet 42, 579–589 (2010).
Florez, J. C. The genetics of type 2 diabetes: a realistic appraisal circa 2008. J Clin Endocrin Metab 93, 4633–4642 (2008).
Benzinou M. et al. (2008) Common nonsynonymous variants in PCSK1 confer risk of obesity. Nat Genet 40, 943–945 (2008).
Heni, M. et al. Association of obesity risk SNPs in PCSK1 with insulin sensitivity and proinsulin conversion. BMC Med Genet 11, 86 (2010).
Leak, T. S. et al. Association of the proprotein convertase subtilisin/kexin-type 2 (PCSK2) gene with type 2 diabetes in an African American population. Mol Genet Metab 92, 145–150 (2007).
Yoshida, H. et al. Association of the prohormone convertase 2 gene (PCSK2) on chromosome 20 with NIDDM in Japanese subjects. Diabetes 44, 389–393 (1995).
Zheng, X. et al. Association of type 2 diabetes susceptibility genes (TCF7L2, SLC30A8, PCSK1 and PCSK2) and proinsulin conversion in a Chinese population. Mol Biol Rep 39, 17–23 (2012).
Thorisson, G. A. et al. HGVbaseG2P: a central genetic association database. Nucleic Acids Res 37, D797–D802 (2009).
Li, Y., Willer, C. J., Ding, J., Scheet, P. & Abecasis, G. R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 34, 816–834 (2010).
Li, Y., Willer, C. J., Sanna, S. & Abecasis, G. R. Genotype Imputation. Annu Rev Genomics Hum Genet 10, 387–406 (2009).
Chiu Y. F. et al. Bivariate genome-wide scan for metabolic phenotypes in non-diabetic Chinese individuals from the Stanford, Asia and Pacific Program of Hypertension and Insulin Resistance Family Study. Diabetologia 50, 1631–1640 (2007).
Davidson, H. W., Rhodes, C. J. & Hutton, J. C. Intraorganellar calcium and pH control proinsulin cleavage in the pancreatic beta cell via two distinct site-specific endopeptidases. Nature 333, 93–96 (1988).
Naggert, J. K. et al. Hyperproinsulinaemia in obese fat/fat mice associated with a carboxypeptidase E mutation which reduces enzyme activity. Nat Genet 10, 135–142 (1995).
Jackson, R. S. et al. Obesity and impaired prohormone processing associated with mutations in the human prohormone convertase 1 gene. Nat Genet 16, 303–306 (1997).
Kahn, S. E. et al. Proinsulin as a marker for the development of NIDDM in Japanese-American men. Diabetes 44, 173–179 (1995).
Mykkanen, L., Zaccaro, D. J., Hales, C. N., Festa, A. & Haffner, S. M. The relation of proinsulin and insulin to insulin sensitivity and acute insulin response in subjects with newly diagnosed type II diabetes: the Insulin Resistance Atherosclerosis Study. Diabetologia 42, 1060–1066 (1999).
Hanley, A. J. et al. Increased proinsulin levels and decreased acute insulin response independently predict the incidence of type 2 diabetes in the insulin resistance atherosclerosis study. Diabetes 51, 1263–1270 (2002).
Wareham, N. J., Byrne, C. D., Williams, R., Day, N. E. & Hales, C. N. Fasting proinsulin concentrations predict the development of type 2 diabetes. Diabet Care 22, 262–270 (1999).
Furuta, M. et al. Defective prohormone processing and altered pancreatic islet morphology in mice lacking active SPC2. Proc Natl Acad Sci USA 94, 6646–6651 (1997).
Jonsson, A. et al. Effect of a common variant of the PCSK2 gene on reduced insulin secretion. Diabetologia 55, 3245–3251 (2012).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100, 9440–9445 (2003).
Ranade, K. et al. Genetic variation in aldosterone synthase predicts plasma glucose levels. Proc Natl Acad Sci USA 98, 13219–13224 (2001).
Olivier, M. et al. Single nucleotide polymorphisms in protein tyrosine phosphatase 1beta (PTPN1) are associated with essential hypertension and obesity. Hum Mol Genet 13, 1885–1892 (2004).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Horvath, S., Xu, X. & Laird, N. M. The family based association test method: strategies for studying general genotype-phenotype associations. Eur J Hum Genet 9, 301–306 (2001).
Grambsch, P. M. & Therneau, T. M. Proportional hazards tests and diagnostics based on weighted residuals. Biometrika 81, 515–526 (1994).
North, B. V., Curtis, D. & Sham, P. C. A note on the calculation of empirical P values from Monte Carlo procedures. Am J Hum Genet 71, 439–441 (2002).
Acknowledgements
The authors wish to thank all participants in this study. We also thank Ms. Kuan-Ching Lee for her excellent technical support. The authors are also very grateful to Ms. Su-Mei Wang for her computing help. We also thank the staff of the Eighth Core Lab, Department of Medical Research, National Taiwan University Hospital, for technical support during the study. This work was supported by grants (NSC94-3112-B-002-019; NSC95-3112-B-002-002; NSC96-3112-B-002-002) from the National Science Council, Executive Yuan, Taiwan.
Author information
Authors and Affiliations
Contributions
Y.F.C., Y.S.J. and L.M.C. participated in concept/design. W.H.-H.S., K.C.S., C.M.H., T.Q. and S.S.K. participated in the collection of clinical and laboratory data. T.J.C. participated in data analysis/interpretation and the drafting of the paper. Y.F.C. participated in data analysis and critical revision of the paper. Y.C.C. participated in data analysis and interpretation. L.M.C. participated in critical revision and approval of paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chang, TJ., Chiu, YF., Sheu, WH. et al. Genetic polymorphisms of PCSK2 are associated with glucose homeostasis and progression to type 2 diabetes in a Chinese population. Sci Rep 5, 14380 (2015). https://doi.org/10.1038/srep14380
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14380
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.