Abstract
Bone homeostasis is maintained by the balance of osteoblasts (OBs) and osteoclasts (OCs). Increased activity of OCs not only contributes to pathological bone resorption, such as osteoporosis and periodontitis, but also is responsible for physiological conditions like orthodontic tooth movement (OTM). However, the detailed mechanism by which orthodontic force promotes the formation of OCs is still poorly understood. In this study, we confirmed that static force promoted the differentiation of human cord monocytes (HMNCs) into OCs depending on loading time and magnitude. Protein expression profiles among HMNCs, HMNCs subjected to static force and mature OCs were established via 2-DE and MALDI-TOF-MS analyses. Total respective protein spot numbers of 549 ± 13, 612 ± 19 and 634 ± 16 were detected in each of the gels by image analysis. The five proteins identified were plasminogen activator inhibitor 2 (PAI-2, Spot 1), peroxiredoxin-6 (PRD-6, Spot 3), manganese superoxide dismutase (SOD2, Spot 6), Rho GDP-dissociation inhibitor 2 (Rho-GDI2, Spot 11) and L-lactate dehydrogenase B chain (L-LDH, Spot 15). More importantly, we revealed that SOD2 was required to maintain monocyte differentiation into functional OCs and may become a potential target for regulating the efficiency of OTM in the future.
Similar content being viewed by others
Introduction
Remodeling processes regulate bone mass homeostasis using the interplay of bone-forming osteoblasts (OBs) and bone-resorbing osteoclasts (OCs)1, which are multinucleated giant cells that originate from the hematopoietic stem cell monocyte/macrophage lineage. OCs are responsible for bone erosion not only in diverse pathological states, such as osteoporosis and periodontitis2,3, but also in physiological conditions like orthodontic tooth movement (OTM)4.
Alveolar bone has the strongest remodeling capacity in the body, which is considerably impacted by the application of mechanical loading5. In orthodontics, the mechanical force exerted on alveolar bone induces alterations in the cellular and molecular processes underlying bone homeostasis and eventually evokes tooth movement. Furthermore, the forces applied generate mechanical stress in the microenvironment of the alveolar extracellular matrix, as well as in the outer membranes, the cytoskeleton, the nuclear protein matrix and the genome of resident cells. These local homeostatic changes lead to synthesis and release of various key molecules that affect the cellular response of different cell types6. As we know, the important factor to determine the efficiency of OTM is the activity of resident OCs, which is also modulated by the mechanical stress5. Some studies even suggest that a change in the alveolar microenvironment, particularly regarding the blood vascular system, is indispensible for the circulating OC-precursors in the peripheral blood to differentiate into OCs7. Previous investigations revealed that OC formation is rapidly induced after application of orthodontic force and that it primarily occurs in vascular canals of the alveolar bone crest on the pressure side4. However, the cellular targets and molecular mechanisms leading to OC formation and maturation upon orthodontic force application are still poorly understood.
Most scholars believe that early OTM is an acute inflammatory reaction characterized by the dilation of periodontal blood vessels and migration of white blood cells into the vasculature, followed by gradual alleviation of the initial acute reaction, which is replaced by a chronic inflammatory reaction8. Past research has proved that phosphorylation of p65 in the NF-κB pathway plays an important role in bone remodeling associated with OTM9. In addition, other studies demonstrated that NF-κB is essential for OC formation and survival when activated by reactive oxygen species10. One gene typically controlled by NF-κB factors is manganese superoxide dismutase (SOD2)11, whose promoter contains a functional κB cis element. SOD2 belongs to a group of SOD enzymes engaged in fending off cellular stress as the first line of defense against the damaging effects of reactive oxygen species. Among these, SOD2 is a highly regulated cytoprotective against oxidative damage and inflammatory responses, which rapidly converts superoxide radicals into hydrogen peroxide and molecular oxygen12. Accordingly, SOD2 presumably constitutes one of the principal cellular defense mechanisms against inflammatory and toxic agents causing oxidative stress, such as tumor necrosis factor α (TNF-α)13,14, interleukin-1 β (IL-1β)15, dinitrophenol16 and PMA17. Researchers proved that superoxide production has been linked with bone resorption activity18 and is specifically localized to the osteoclast ruffled border membrane19. Moreover, SOD2 gene expression is up-regulated in osteoporosis specimens characterized by low BMD, thus making SOD2 a potentially susceptible target for bone resorption diseases20,21. However, the detailed relationship between SOD2 and OTM-related OC formation, which is mainly caused by mechanical force, is not clear.
As OC formation and function represent a central aspect in the molecular and cellular processes involved in OTM, the aim of the present study was to investigate the effect of static loading on OC formation and function in the periodontal microenvironment. Our data in this study suggested that the 150-kpa static force loading for 1.5 h promoted human cord monocytes (HMNCs) to significantly differentiate into OCs. The protein expression profiles among HMNCs, HMNCs subjected to static force and mature OCs were established via 2-DE and MALDI-TOF-MS analyses. Five significantly changed proteins were identified including plasminogen activator inhibitor 2 (PAI-2), peroxiredoxin-6 (PRD-6), manganese superoxide dismutase (SOD2), Rho GDP-dissociation inhibitor 2 (Rho-GDI2,) and L-lactate dehydrogenase B chain (L-LDH). More importantly, we revealed that SOD2 was required to maintain monocyte differentiation into functional OCs as a response to force stimuli.
Results
HMNCs could be successfully induced into mature OCs by 1α,25-(OH)2D3
Identification of CD markers via flow cytometry on HMNCs confirmed that the cells contained many sources. As expected, the monocytes accounted for the vast majority of HMNCs as reflected by CD11b (42.3%) and CD14 (65.6%) analyses. Furthermore, hematopoietic stem cell-related CD34 (22.1%) and CD45 (20.4%) were also positive. More importantly, HMNCs also contained a small population of mesenchymal stromal cells, as assayed by CD105 (13.7%) and CD146 (11.9%) analyses (Fig. 1a). After induction by 1 × 10−8 mol/L 1α, 25-(OH)2D3 for 12 days, HMNCs differentiated into matured OCs, which displayed TRAP-positive staining. Moreover, Giemsa-staining clearly showed multinuclear morphologies of matured OCs (Supplementary Fig. 1 and Fig. 1b). However, the induced OCs could not form any bone-resorbing pits on bone slices (Fig. 1b). OC-associated markers, tartrate-resistant acid phosphatase (TRAP), cathepsin K (CTSK), receptors for calcitonin (CTR) and MMP-9 both increased as indicated by real-time PCR and western blotting after OC induction (Fig. 1c). In addition, the expression of OPG decreased while the expression of RANKL increased after OC induction. These results suggested that 1α, 25-(OH)2D3 could induce HMNCs to differentiate into OCs successfully, mainly because the HMNCs contained several different cell sources.
Identification of induced osteoclasts.
(A), Analyses of cell surface markers were performed via flow cytometry detecting PE, FITC or PerCP conjugated monoclonal antibodies for human CD11b, CD14, CD34, CD45, CD105, CD146, or isotypematched control IgGs. (B), HMNCs induced for 12 d with 1α, 25-(OH)2D3 (10−8 M) showing matured to TRAP positive (red staining) multinuclear cells. Giemsa staining (blue) clearly showed multinuclear morphologies of matured OCs. Bone resorption pits were analysed by Toluidine-blue staining. No resorption pits were formed. Magnification: ×100. (C), The expressions of CTSK, TRAP, CTR, MMP-9, OPG and RANKL were analyzed by real-time PCR and western blotting. Cont, control; MNCs, human cord monocytes; OCs, osteoclasts. BPs, Bone resorption pits. Data represent mean ± SD of the relative ratio of the gene signal to the β-actin signal. *P < 0.05, n = 6.
Static force promoted HMNCs to differentiate into OCs depending on loading time and magnitude
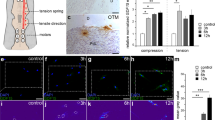
To determine the appropriate loading time and magnitude of static force application on HMNCs differentiation, six different levels of magnitude and five different duration times were tested with regard to the expression level of TRAP mRNA and protein. Real-time PCR assay and western blotting analysis showed that static force promoted TRAP expression depending on the loading time and magnitude, especially under 150 kpa loading for 1.5 h (Fig. 2a). We next tested the effect of static force on differentiation of HMNCs. HMNCs were cultured in induction medium with 1α, 25-(OH)2D3 and simultaneously loaded under 150 kpa static force for 1.5 h every 3 days after the medium was renewed. The results showed that TRAP-positive cells increased significantly with loading static force at day 3 compared with the non-loading group and many multinuclear OCs were formed in the loading group at day 5 (Fig. 2b and Supplementary Fig. 2). More importantly, bone-resorbing pits were observed in the loading group, while almost no pits could be seen in the non-loading group (Fig. 2c). These results suggested that the 150 kpa static force loading for 1.5 h significantly promotes differentiation of HMNCs into functional-OCs.
Pick up effective loading time and magnitude of static force.
(A), TRAP mRNA and protein expression for six force levels with five different force duration times evaluated via real-time PCR and western blotting. (B), (C), HMNCs were cultured in induction medium with 1α, 25-(OH)2D3 and simultaneously loaded with 150 kpa static force for 1.5 h every 3 days after the medium renew. OC formation was analysed by TRAP staining (B) after loading for 3 and 5 days and bone resorption pits were examined via Toluidine-blue staining (C) under the same loading conditions for 9 days. Cont, HMNCs were cultured in induction medium with 1α, 25-(OH)2D3; SF, HMNCs were cultured in induction medium with 1α, 25-(OH)2D3 and simultaneously loaded with 150 kpa static force for 1.5 h every 3 days after the medium was renewed. magnification × 100 and × 200. Data represent mean ± SD.*P < 0.05, n = 6.
Protein expression profiles among HMNCs, HMNCs subjected to static force and mature OCs were established via proteomic analysis
A total of nine 2-DE experiments were performed for three HMNCs control samples, three HMNC samples with 150 kpa static force application for 1.5 h and three induced-mature OC samples. For these groups, total protein spot numbers of 549 ± 13, 612 ± 19 and 634 ± 16, respectively, were detected in each of the gels over a pH range of 3 to 10 by image analysis (Fig. 3a–c). The comparative analysis of the normalized volumes of each spot revealed that a total of five spots were differentially expressed among the groups. Three spots were up-regulated after static force loading and two spots were only expressed in mature OCs (Fig. 3d). An ESI-Q-TOF MS/MS analyzer was used for accurate identification of the proteins and the mass identification information for the five key proteins is listed in Table 1. The five proteins were identified as plasminogen activator inhibitor 2 (PAI-2, Spot 1), peroxiredoxin-6 (PRD-6, Spot 3), manganese superoxide dismutase (SOD2, Spot 6), Rho GDP-dissociation inhibitor 2 (Rho-GDI2, Spot 11) and L-lactate dehydrogenase B chain (L-LDH, Spot 15) (Fig. 4).
Established protein expression profiles among HMNCs, loaded HMNCs and OCs.
(A)–(C), Representative 2-DE maps of the whole-cell lysates for HMNCs (A), HMNCs + static force (B), induced OCs (C). (D), Magnified proteins spots indicating the differential expression trends among the three groups investigated. MNCs, human cord monocytes; OCs, osteoclasts; SF, static force. Data represent mean ± SD. *P < 0.05, n = 3.
Peptide fingerprint mass spectra analyses of key proteins.
(A)–(E), The five key proteins plasminogen activator inhibitor 2 (A: PAI-2, Spot 1), peroxiredoxin-6 (B: PRD-6, Spot 3), manganese superoxide dismutase (C: SOD2, Spot 6), Rho GDP-dissociation inhibitor 2 (D: Rho-GDI2, Spot 11) and L-lactate dehydrogenase B chain (E: L-LDH, Spot 15) identified via 2-DE maps were trypsin digested and evaluated by mass spectrometry.
Then, the expression of PAI-2, PRD-6, SOD2, Rho-GDI2 and L-LDH was confirmed by RT-PCR (Fig. 5a) and western blotting (Fig. 5b) to validate the result of the 2-DE analyses. The expression levels of SerpinB2 (PAI-2), PRDX-6 and L-LDH (LDHB) were increased when HMNCs were exposed to static force and reached even higher levels in mature OCs. Additionally, Rho-GDI 2 (D4 GDI) was only expressed in mature OCs. Interestingly, the expression of SOD2 contrasted with these results, as its level was higher in the static force group than in the OCs group.
Validation of the five proteins identified by 2-DE and MS.
(A), mRNA levels of PAI-2, PRD-6, SOD2, Rho-GDI2 and L-LDH analyzed by RT-PCR. Data represent mean ± SD of the relative ratio of the gene signal to the β-actin signal. (B), Western blotting analysis for the five proteins. β-actin was used as loading control. MNCs, human cord monocytes; OCs, osteoclasts; SF, static force. *P < 0.05, n = 6.
SOD2 was required to maintain differentiation of HMNCs into mature OCs
To elucidate the function of SOD2 in OC formation under the static force, siRNA-coding oligos against mouse SOD2 were transfected into the RAW264.7 cells. Among the three si-SOD2 oligos investigated, si-832 was the most effective siRNA, leading to a greater than 4-fold reduction of SOD2 expression both on mRNA and on protein levels (Supplementary Fig. 3). After down-regulating the expression level of SOD2, the viability of RAW264.7 cells showed no significant difference. However, after adding static force, the viability of RAW264.7 cells reduced obviously in the SOD2 knockdown group, as shown by the MTT assay (Fig. 6a). RAW264.7 cells then were cultured in induction medium with 50 ng/ml RNAKL and simultaneously loaded with 150 kpa static force for 1.5 h every 3 days after the medium was renewed. The apoptosis rate of the SOD2 knockdown group showed almost no difference at day 3 but increased significantly at day 5 compared to the control group (Fig. 6b, c and Supplementary Fig. 4). TRAP staining showed that the cell numbers of TRAP-positive osteoclasts, as well as the bone resorption areas, were fewer after down-regulating the SOD2 expression than in the control groups, which is presented in Supplementary Fig. 3 and Fig. 6d–f. Furthermore, the expression of osteoclast specific genes, namely CK, TRAP and MMP-9, were reduced both in terms of mRNA and protein levels after down-regulating SOD2 expression (Fig. 6g).
SOD2 plays an important role in OCs formation and function under static force.
(A), The viability of RAW264.7 cells, which were divided as Cont siRNA with or without static force and SOD2 siRNA with or without static force, was analysed by MTT assay. (B), RAW264.7, which were transfected with Cont siRNA or SOD2 siRNA in advance, were cultured in induction medium with 50 ng/ml RNAKL and simultaneously loaded with 150 kpa static force for 1.5 h every 3 days after the medium was renewed. Apoptosis was determined by flow cytometric analysis using double of cells with Annexin V/PI. The percentage of apoptotic cells was calculated as early apoptosis (AV) and late apoptosis (AV + PI). (C), Quantification of apoptosis analysis. (D), Under similar induction medium, OC formation was analysed by TRAP staining after loading for 3 and 5 days and bone resorption pits were examined via Toluidine-blue staining under the same loading conditions for 9 days. (E), (F), The number of TRAP-positive cells, as well as occurrence of bone resorption areas, was evaluated. (G), The expression levels of osteoclast-specific genes CTSK, TRAP and MMP-9 were consistently reduced after down-regulating SOD2 expression both on mRNA and protein levels. Cont siRNA, transfected negative control siRNA; SOD2 siRNA, knockdown SOD2 expression; SF, static force. magnification × 100. Data represent the mean ± SD. *P < 0.05, n = 3.
Next, we chose mouse bone marrow monocyte cells (BMMCs) to further confirm the effect of SOD2 on osteoclastogenesis. Static force loading promoted the expression of SOD2 at day 1 and then began to decrease, but was still higher than the expression in the control group (Fig. 7a). We then induced BMMCs into mature osteoclasts by 20 ng/ml M-CSF and 100 ng/ml RANKL and simultaneously loaded 150 kpa static force for 1.5 h every 3 days. TRAP staining showed that TRAP-positive osteoclasts, as well as the bone resorption areas, were fewer in number after down-regulating the SOD2 expression as compared to the control groups (Fig. 7b–d). Furthermore, the expression of osteoclast specific genes was reduced both in terms of mRNA and protein levels after down-regulating SOD2 expression (Fig. 7e). Together, these data indicated that SOD2 may play an important part in osteoclast differentiation, formation and function.
SOD2 is also necessary for OCs formation and function in the bone marrow system.
(A), The expression level of SOD2 in BMMCs was analyzed by western blotting after loading 150 kpa static force for 1.5 h/day for 1, 3 and 5 days. (B), BMMCs, which was transfected with Cont siRNA or SOD2 siRNA in advance, were cultured in induction medium with 20 ng/mlM-CSF, 100 ng/ml RNAKL and simultaneously loaded with 150 kpa static force for 1.5 h every three days after the medium was renewed. OC formation was analysed by TRAP staining after loading for 3 and 5 days and bone resorption pits were examined via Toluidine-blue staining under the same loading conditions for 9 days. (C), (D), The number of TRAP-positive cells, as well as the occurrence of bone resorption areas, was evaluated. (E), The expression levels of osteoclast-specific genes CTSK, TRAP and MMP-9 were consistently reduced after down-regulating SOD2 expression both on mRNA and protein levels. Cont siRNA, transfected negative control siRNA; SOD2 siRNA, knockdown SOD2 expression; SF, static force. magnification × 100. Data represent the mean ± SD. *P < 0.05, n = 3.
Discussion
OTM process is closely associated with the activity of orthodontic force which is regulated by orthodontic force and bone resorption initially occurs on the pressure side of the alveolar bone22. Our studies reveal that static force promoted OC formation and bone resorption may via regulating the expression of SOD2. Baumrind23 stated that orthodontic forces are transmitted to all tissues in the region of force application as a function of the elastic band properties and that undermining bone resorption stimulated by inflammatory cytokines and by hydraulic pressures in the adjacent alveolar bone precedes the structural changes initiated in the PDL. As the blood vessels in the vascular canals of the alveolar bone are not directly compressed by orthodontic force, OCs and their precursors, both mobilized by increased blood flow in the PDL, are thought to be recruited from hemapoietic tissues and responsible for the pathological processes in the early phase of tooth movement24,25. OCs are believed to originate from a precursor cell of the monocyte/macrophage lineage1, but the stage at which the monocyte and osteoclast differentiation pathways diverge is yet unknown. However, it is generally thought that adult monocytes are too committed to their own lineage to differentiate into OCs26. Therefore, we developed long-term cultures of human cord monocytes, which have greater potential compared with adult monocytes and easily induced to express OC phenotypic features in vitro7,27,28. Surprisingly, HMNCs could be successfully induced into mature osteoclasts by 1α, 25-(OH)2D3. We inferred this mainly because the HMNCs are a mixed population of cells, containing a group of CD105 and CD146 positive mesenchymal stromal cells. This is consistent with the findings of previous studies, which have shown that the presence of 1α, 25-(OH)2D3 and a bone-derived stromal cell population were absolute requirements for OC differentiation7. More importantly, static force could promote differentiation of HMNCs into mature and functional OCs in induction medium, which could form resorption pits on bone slices. This finding suggests that the circulating OC precursors and mesenchymal stromal cells are present in the monocyte fraction and that particular stimuli such as orthodontic force can induce the differentiation of these circulating precursor cells into functional OCs. However, the precise function of static force in the induction and functional maturation of OCs still has to be clarified.
Based on our 2-DE and MALDI-TOF-MS analyses, we established the protein profiles among HMNCs, HMNCs subjected to static force and mature OCs. The proteins PAI-229, PRD-630,31, SOD220, Rho-GDI232 and L-LDH33 were identified. According to the references, these proteins are all related to bone metabolism and are mainly specific to OC formation and function. After all the results were validated by RT-PCR and western blotting, we found that the expression level of SOD2 was higher in HMNCs loaded with static force than in induced OCs, but they both expressed higher levels than HMNCs. Then, we used the osteoclast progenitor cell line RAW264.7 to further investigate the relationship between SOD2 and osteoclast formation and function. The results showed high expression of SOD2 in RAW264.7 compared to HMNCs and down-regulation of SOD2 in RAW264.7, confirming that it was required to maintain the differentiation and function of OCs. This plasma membrane SOD may help protect osteoclasts from the effects of extracellular superoxide anions released during bone resorption34. Furthermore, SOD2 catalyzes the production of hydrogen peroxide (H2O2) from superoxide and interestingly, H2O2 may stimulate osteoclast differentiation35. Key et al.22,23 proposed that superoxide generation was defective in osteoclasts from OP patients and was related to the defect in bone resorption, thus emphasizing the role played by osteoclastic superoxide in bone resorption36. OTM required OCs for the resorption of alveolar bone on the pressure side. Therefore, combined with our results, we reveal that SOD2 is required to maintain OCs formation and function, which may not only serve as a potential target to regulate the efficiency of OTM but also as a therapeutic target for other osteoclast-related disorders.
Methods
Cell culture of HMNCs and induction of OCs
The experimental protocols dealing with human umbilical cord blood were approved by the institutional review board at our institute (School of Stomatology, The Fourth Military Medical University, Ethics Committee). Signed informed consent was obtained from six donors and all protocols were in accordance with the approved guidelines. HMNCs were isolated from the umbilical cord blood (n = 6) using Ficoll-Hypaque (Pharmacia Biotech, Hertford, U.K.), density gradient centrifugation. Monocytes and lymphocyte-rich band were collected, washed twice in PBS and then resuspended in α-MEM (Gibco, Paisley, U.K.) with 15% FBS at 37°C in a humidified 5% CO2 atmosphere. 1 × 107/ml cells were cultured on a 24-well plate containing sterilized coverslips. After 12 h of culturing, nonadherent lymphocytes were washed off and new α-MEM containing 1α, 25-(OH)2D3 (1 × 10−8 M, R&D Systems, Inc., Minneapolis, MN, USA). Half of the culture medium was replaced every 3 d for a total of 12 days.
Murine monocyte/macrophage cell line RAW264.7 was seeded at a density of 1 × 105 cells/well in 12-well plates and maintained in DMEM (Gibco) supplemented with 10% FBS. OC formation was induced by addition of 50 ng/ml murine recombinant RANKL (PeproTech) from d 1 until d 12.
All of the procedures that involved animals were approved by the Animal Use and Care Committee of the Fourth Military Medical University (license number: SCXK 2007-007) and protocols were in accordance with the approved guidelines. Murine bone marrow cells (BMCs) were collected by frequent injection of PBS throughout the entire marrow cavity of the tibiae and femora from 8-week-old mice and 0.5 × 106 BMCs were suspended in α-MEM (Invitrogen) containing 15% FBS (Equitech-Bio), L-glutamine (Invitrogen), penicillin and streptomycin (Invitrogen) and 20 ng/ml M-CSF (PeproTech) in a 24-well plate for 48 hours. The adherent cells were then cultured with 20 ng/ml M-CSF (PeproTech) and 100 ng/ml RANKL (PeproTech) for another 7 days.
Flow cytometric analysis for CD markers
To investigate the subgroups of HMNCs, 5 × 105 cells were incubated with PE, FITC or PerCP conjugated monoclonal antibodies for human CD11b, CD14, CD34, CD45, CD105, CD146 (R&D Systems, Inc.), or isotype-matched control IgGs. Cells were subjected to flow cytometric analysis using a Beckman Coulter Epics XL (Beckman Coulter, Fullerton, CA, USA).
Tartrate-resistant acid phosphatase (TRAP) staining
OCs were washed in PBS, fixed in 4% paraformaldehyde and stained for TRAP using a commercially available kit (Acid Phosphatase Kit 387-A; Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer's instructions. Five random fields of view per well were captured under microscope (Leica Microsystems, Heerbrugg, Switzerland) (100 × magnification) to quantify the number of OC-positive cells showing a red cytoplasmic staining as much as the number of nuclei in each cell. Experiments were performed in triplicate.
Bone resorption assay
To evaluate bone resorption, 1 × 105 cells/well were cultured under the same culture condition as indicated above on 12-well plates containing sterilized calf cortical bone slices (100 pm). After 12 d, TRAP-positive multinucleated OCs appeared and were removed with a solution of 20% sodiumhypochlorite for 5 min. Bone discs were stained with toluidine blue for 4 min at room temperature for a better analysis of the resorption areas showing pit formation. The resorbed areas were defined manually and measured using Image J (NIH, version 1.42) to obtain the number of pixels within each resorbed area. Experiments were performed in triplicate.
Real-time quantitative reverse transcription polymerase chain reaction (RT-PCR)
Total RNA was extracted from each sample using TRIzol reagents (TRIzol; Invitrogen) and converted to cDNA (SuperScript First-Strand Synthesis Kit; Invitrogen). RT-PCR was performed using the QuantiTect SYBR Green PCR Kit (Toyobo, Osaka, Japan) and the Applied Biosystems 7500 Real-Time PCR Detection System. For quantification, target gene mRNA expressions were normalized to β-actin.
Western blot analysis
Whole cell proteins were extracted via RIPA lysis buffer (P0013B, Beyotime Co., Shanghai, China), quantified by BCA assay (Sigma-Aldrich Co), separated on NuPAGE 10%–12% polyacrylamide gels, transferred to PVDF membranes (Millipore, Billercia, MA) and blocked in 5% BSA in TBST. Incubation was performed with antibodies against Cathepsin K (Abcam, Cambridge, England, 1:1000), TRAP (Abcam:1000), MMP-9 (Cell signalling, 1:1000), Calcitonin receptor (Abcam, 1:800), SerpinB2 (Abcam, 1:800), Peroxiredoxin 6 (Abcam, 1:1000), SOD2 (Abcam, 1:2000), D4 GDI (Abcam, 1:500), Lactate Dehydrogenase (Abcam, 1:3000), OPG (Santa Cruz Biotechnology, 1:1000), RANKL (Santa Cruz Biotechnology, 1:1000), β-actin (Cell-signaling, 1:2000). After washing with TBST, blots were incubated with goat anti-rabbit, goat anti-mouse or rabbit anti-goat horseradish peroxidase-conjugated secondary antibodies previous to visualization with enhanced chemi-luminescence ECL kits (Sigma). Relative band intensities in scanned images were analyzed with Image J software.
Static force loading of HMNCs
For evaluation of static force magnitudes and effecting duration times, HMNCs were seeded at a density of 1 × 105 cells/well in 24-well plates. Static force was loaded in a series of magnitudes of 50 kpa,100 kpa,150 kpa,200 kpa and 300 kpa along with different effecting times of 0.5 h,1 h,1.5 h,2 h and 3 h in static force equipment, which is developed by temporal-mandibular joint lab in our stomatological hospital37,38. Cultures were maintained at 37°C in a humidified 5% CO2 atmosphere in the force equipment, equal to the conditions in the incubator. TRAP staining was used to quantify the number of mature osteoclasts containing at least 3 nuclei per cell. Five random fields of view per well were captured under microscope (Leica Microsystems, Heerbrugg, Switzerland).
Isoelectric focusing (IEF) and SDS-PAGE
On the basis of both the RT-PCR and TRAP-staining results, 150 kpa of static force loaded for 1.5 h was chosen to be applied to HMNCs seeded at a density of 1 × 106 cells/well in a 6-well plate. These cells were subsequently prepared for further two-dimensional electrophoresis (2-DE) and image analysis.
HMNCs, HMNCs loaded with 150 kpa static force for 1.5 h and OCs were lysed in RIPA buffer, extracted whole-cell lysates were centrifuged and supernatants were collected to remove cell debris. Hereupon, supernatants were acetone precipitated, followed by resuspension in the sample solution (7 M Urea, 2 M Thiourea, 0.5 M Tris, 4% CHAPS, 0.5 M EDTA, 65 mM DTT, 1% IPG buffer (pH 3–10), protein inhibitor cocktail). The concentrations of protein in each sample were determined via modified Bradford methods, using BSA as standard.
IEF was performed for three technical replicates per sample to determine protein amounts. Linear 13 cm pH 3–10 IPG strips were used according to the manufacturer's protocol (Bio-Rad) with minor modifications. In brief, a sample containing 60 μg of protein was diluted to 250 μl rehydration solution (7 M urea, 2%CHAPS, 100 mM DTT, 0.5% IPG buffer and bromophenol blue) and applied to the strips by overnight rehydration under passive conditions. The IPG strips were equilibrated for 15 min in equilibration solution (6 M Urea, 2% SDS, 0.05 M Tris-HCl, pH 8.8 and 20% glycerol) containing 2% DTT and then for 15 min in the same buffer containing 2.5% iodoacetamide. The equilibrated IPG strips were transferred to 12.5% uniform polyacrylamide gels at a current setting of 5 mA/gel for 1 h and at 25 mA/gel thereafter. The stained 2D gels were imaged using the VersaDoc imaging system (Bio-Rad).
Image analysis
Dried gels were scanned with an image scanner at 300 dpi. Analysis of gel images was carried out using ImageMaster TM 2D Platinum (version 6.0, Amersham Biosciences, Sweden). The protein spot volume was evaluated based on the lowest boundary mode of background selection. Normalization was carried out using the “total spot volume” method, dividing the volume of each spot by the total volume of all spots in the gel. The process of spot matching was repeated with a reference gel. For each of the spots, the relative % volume was averaged and expressed as mean ± SD.
MALDI-TOF analysis
Five protein spots chosen for excision were carried out using the solution-phase nitrocellulose method. Briefly, α-cyano-4-hydro-xycinnamic acid (40 mg/ml) and nitrocellulose (20 mg/ml) were prepared separately in acetone and mixed with 2-propanol at a ratio of 2:1:1. Then, 2 μl of the mixture was added to 2 μl of the peptide sample solution and 1 μl of each solution was spotted on a MALDI plate for 5 min. The plates were then washed with 5% formic acid, followed by water. Dried spots were analyzed using a Voyager-DE STR MALDI-TOF mass spectrometer (Applied Biosystems). Proteins were identified by peptide mass finger printing using the program Aldente (http://www.expasy.org/).
Transfection of SOD2 siRNA
SiRNAs of SOD2 (Gene-Pharma, ShaiHai, China) were chemically modified (2′-O-Methyl oligos). RAW264.7 cells were transfected at 60% confluence in 100-cm plates in serum-free medium (DMEM, Gibco, Paisley, U.K.) by using lipofectamine reagent (Invitrogen) according to the manufacturer's instructions. For silencing, cells were transfected with 50 μM of SOD2 siRNA or with 50 nM of Silencer Negative Control no. 1 siRNA. After 48 h of transfection, cells were washed, trypsinized and harvested for evaluation of mRNA and protein expression. Primer sequences are listed in table 2.
MTT assay
To examine the viability of RAW264.7 cells, which were divided as Cont siRNA with or without static force and SOD2 siRNA with or without static force, 5 × 103 cells/well were cultured in 96-well plates. The 3-(4, 5-dimethylthiazol-2yl)-2, 5-diphenyltetrazolium bromide (MTT) assay was carried out for 7 days according to the manufacturer's protocol (Sigma). Absorbance was determined at 490 nm with a microplate reader (Bio-Tek Instruments, Winooski, VT, USA). Experiments were performed in triplicate.
Flow cytometric analysis for apoptosis
After being transfected by Cont siRNA or SOD2 siRNA, RAW264.7 cells was loaded with 150 kpa static force for 3 and 5 days respectively. Apoptosis was analyzed by flow cytometry following Annexin V (AV) and propidium iodide (PI) labeling following the manufacturer's staining protocol (Neo-Bioscience, Shenzhen, China). Analysis was performed using Cell Quest Pro Software (BD Biosciences). Samples were gated on the monocyte population using forward and side scatter plots with a minimum of 10,000 gated events sampled. The percentage of apoptotic cells was calculated as early apoptosis (AV) add late apoptosis (AV + PI). Experiments were performed in triplicate.
Statistics
Results are presented as mean ± SD from at least three independent experiments and analyzed by a two-tailed unpaired Student's t test using SPSS software. P values < 0.05 were considered statistically significant. For analysis of multiple groups, the P values were adjusted using the Bonferroni method.
References
Kwak, H. B. et al. Inhibition of osteoclast differentiation and bone resorption by rotenone, through down-regulation of RANKL-induced c-Fos and NFATc1 expression. Bone 46, 724–31 (2010).
Boyle, W. J., Simonet, W. S. & Lancey, D. L. Osteoclast differentiation and activation. Nature 423, 337–42 (2003).
Teitebaum, S. L. Bone resorption by osteoclasts. Science 289, 1504–8 (2000).
Yokoya, K., Sasaki, T. & Shibasaki, Y. Distributional changes of osteoclasts and pre-osteoclastic cells in periodontal tissues during experimental tooth movement as revealed by quantitative immunohistochemistry of H(+)-ATPase. J Dent Res 76, 580–7 (1997).
Karras, J. C., Miller, J. R., Hodges, J. S., Beyer, J. P. & Larson, B. E. Effect of alendronate on orthodontic tooth movement in rats. Am J Orthod Dentofacial Orthop 136, 843–7 (2009).
Kheralla, Y., Götz, W., Kawarizadeh, A., Rath-Deschner, B. & Jäger, A. IGF-I, IGF-IR and IRS1 expression as an early reaction of PDL cells to experimental tooth movement in the rat. Arch Oral Biol 55, 215–22 (2010).
Quinn, J. M., Fujikawa, Y., McGee, J. O. & Athanasou, N. A. Rodent osteoblast-like cells support osteoclastic differentiation of human cord blood monocytes in the presence of M-CSF and 1,25 dihydroxyvitamin D3. Int J Biochem Cell Biol 29, 173–9 (1997).
Yang, J. H. et al. Effect of orthodontic force on inflammatory periodontal tissue remodeling and expression of IL-6 and IL-8 in rats. Asian Pac J Trop Med 6, 757–61 (2013).
Zuo, J. et al. Nuclear factor kappaB p65 phosphorylation in orthodontic tooth movement. J Dent Res 86, 556–59 (2007).
Xu, J. et al. NF-kappaB modulators in osteolytic bone diseases. Cytokine Growth Factor Rev 20, 7–17 (2009).
Jones, P. L., Ping, D. & Boss, J. M. Tumor necrosis factor alpha and interleukin-1beta regulate the murine manganese superoxide dismutase gene through a complex intronic enhancer involving C/EBP-beta and NF-kappaB. Mol Cell Biol 17, 6970–81 (1997).
Weisiger, R. A. & Fridovich, I. J. Superoxide dismutase: Organelle specificity. J Biol Chem 248, 4793–4796 (1973).
Wong, G. H. & Goeddel, D. V. Induction of manganous superoxide dismutase by tumor necrosis factor: possible protective mechanism. Science 242, 941–94 (1988).
Mao, X., Moerman-Herzog, A. M., Wang, W. & Barger, S. W. Differential transcriptional control of the superoxide dismutase-2 kappaB element in neurons and astrocytes. J Biol Chem. 281, 35863–72 (2006).
Masuda, A. et al. Induction of mitochondrial manganese superoxide dismutase by interleukin 1. FASEB J 15, 3087–3091 (1988).
Dryer, S. E., Dryer, R. L. & Autor, A. P. Enhancement of mitochondrial, cyanide-resistant superoxide dismutase in the livers of rats treated with 2,4-dinitrophenol. J Biol Chem 255, 1054–1057 (1980).
Fujii, J. & Taniguchi, N. Phorbol ester induces manganese superoxide dismutase in tumor necrosis factor-resistant cells. J Biol Chem. 266, 23142–23146 (1991).
Garrett, I. R. et al. Oxygen-derived free radicals stimulate osteoclastic bone resorption in rodent bone in vitro and in vivo. J Clin Invest 85, 632–639 (1990).
Key, L. L., Wolf, W. C., Gundberg, C. M. & Ries, W. L. Superoxide and bone resorption. Bone 15, 431–436 (1994).
Deng, F. Y. et al. An integrative study ascertained SOD2 as a susceptibility gene for osteoporosis in Chinese. J Bone Miner Res 26, 2695–701 (2011).
Xiao, H., Shan, L., Zhu, H. & Xue, F. Detection of significant pathways in osteoporosis based on graph clustering. Mol Med Rep 6, 1325–32 (2012).
Matsumoto, T., Iimura, T., Ogura, K., Moriyama, K. & Yamaguchi, A. The role of osteocytes in bone resorption during orthodontic tooth movement. J Dent Res 92 340–345 (2013).
Baumrind, S. A reconsideration of the propriety of the “pressure-tension” hypothesis. Am J Orthod 55, 12–22 (1969).
Udagawa, N. The mechanism of osteoclast differentiation from macrophages: possible roles of T lymphocytes in osteoclastogenesis. J Bone Miner Metab 21, 337–43 (2003).
Kitaura, H. et al. Effect of cytokines on osteoclast formation and bone resorption during mechanical force loading of the periodontal membrane. ScientificWorldJournal 617032 eCollection (2014).
Roodman, G. D. Osteoclast differentiation. Crit Rev Oral Biol Med 2, 389–409. (1991).
Orcel, P., Bielakoff, J. & de Vernejoul, M. C. Formation of multinucleated cells with osteoclast precursor features in human cord monocytes cultures. Anat Rec 226, 1–9 (1990).
Cournot, G. et al. Cultured circulating mononuclear cells from osteopetrotic infants express the osteoclast-associated vitronectin receptor and form multinucleated cells in response to 1, 25-dihydroxyvitamin D3. J Bone Miner Res 8, 61–70 (1993).
Yang, J. N., Allan, E. H., Anderson, G. I., Martin, T. J. & Minkin, C. Plasminogen activator system in osteoclasts. J Bone Miner Res 12, 761–768 (1997).
Ikeda, D., Ageta, H., Tsuchida, K. & Yamada, H. iTRAQ-based proteomics reveals novel biomarkers of osteoarthritis. Biomarkers 18, 565–572 (2013).
Erttmann, S. F. et al. PGD2 and PGE2 regulate gene expression of Prx 6 in primary macrophages via Nrf2. Free Radic Biol Med 51, 626–640 (2011).
Mullin, B. H., Mamotte, C., Prince, R. L. & Wilson, S. G. Influence of ARHGEF3 and RHOA knockdown on ACTA2 and other genes in osteoblasts and osteoclasts. PLoS One 9, e98116 (2014).
Morten, K. J., Badder, L. & Knowles, H. J. Differential regulation of HIF-mediated pathways increases mitochondrial metabolism and ATP production in hypoxic osteoclasts. J Pathol 229, 755–764 (2013).
Khalkhali-Ellis, Z., Collin-Osdoby, P., Li, L., Brandi, M. L. & Osdoby, P. A human homolog of the 150 kD avian osteoclast membrane antigen related to superoxide dismutase and essential for bone resorption is induced by developmental agents and opposed by estrogen in FLG 29.1 cells. Calcif Tissue Int 60, 187–93 (1997).
Deng, F. Y. et al. Proteomic analysis of circulating monocytes in Chinese premenopausal females with extremely discordant bone mineral density. Proteomics 8, 4259–72 (2008).
Madyastha, P. R., Yang, S., Ries, W. L. & Key, L. L., Jr IFN-gamma enhances osteoclast generation in cultures of peripheral blood from osteopetrotic patients and normalizes superoxide production. J Interferon Cytokine Res 20, 645–52 (2000).
Zhang, M., Wang, J. J. & Chen, Y. J. Effects of mechanical pressure on intracellular calcium release channel and cytoskeletal structure in rabbit mandibula r condylar chondrocytes. Life Sci 78, 2480–2487 (2006).
Zhang, M., Chen, Y. J., Ono, T. & Wang, J. J. Crosstalk between integrin and G protein pathways involved in mechanotransduction in mandibular condylar chondrocytes under pressure. Arch Biochem Biophys 474, 102–108 (2008).
Acknowledgements
This work was supported by grants from the national nature science foundation of china (31200972, 81100058 and 81171001).
Author information
Authors and Affiliations
Contributions
T.G. designed the experiments and collected the data. L.Z. prepared the data. A.K. revised the whole manuscript. H.Z. participated the experiments of proteomics. F.J. and W.L. made critical intellectual contributions that formed the central concept of this study and were involved in the writing process of the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Guo, T., Zhang, L., Konermann, A. et al. Manganese superoxide dismutase is required to maintain osteoclast differentiation and function under static force. Sci Rep 5, 8016 (2015). https://doi.org/10.1038/srep08016
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08016
This article is cited by
-
Association between blood manganese and bone mineral density in US adolescents
Environmental Science and Pollution Research (2022)
-
Mate tea (Ilex paraguariensis) improves bone formation in the alveolar socket healing after tooth extraction in rats
Clinical Oral Investigations (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.