Abstract
Selective elimination of synaptic connections is a common phenomenon which occurs during both developmental and pathological conditions. Glial cells have a central role in the pruning of synapses by specifically engulfing the degenerating neurites of inappropriate connections, but its regulatory mechanisms have been largely unknown. To identify mediators of this process, we established an in vitro cell culture assay for the synapse elimination. Neuronal differentiation and synapse formation of PC12 cells were induced by culturing the cells with nerve growth factor (NGF) in a serum-free medium. To trigger synapse elimination, the NGF-containing medium was replaced with DMEM containing 10% FBS and the neurites of PC12 cells degenerated within two days. Co-culturing with MG6 cells, a mouse microglial cell line, accelerated the removal of degenerating neurites of PC12 cells by phagocytosis. When MG6 cells were pre-incubated with exosomes secreted from the differentiated PC12 cells after depolarization, the removal was further accelerated by increasing the expression levels of complement component 3 in the MG6 cells. These results define a role for exosomes as a regulator of synapse elimination and clarify a novel mechanism whereby active synapses promote the pruning of inactive ones by stimulating microglial phagocytosis with exosomes.
Similar content being viewed by others
Introduction
The creation of complex patterns of synaptic connectivity often requires the elimination of only a select subset of the connections initially established by neurons1. The dynamic refinement of synaptic connections is essential not only for the appropriate wiring of neural circuits, but also for behavioral responses to a changing environment as well as for learning and memory2. In mammalian nervous system, synapse pruning events have been reported in various places such as retinotectal system, cerebellum, parasympathetic and sympathetic autonomic ganglia and neuromuscular junctions3. Recent studies have shown that glial cells actively participate in synaptic pruning4,5. In Drosophila, glial cells engulf the degenerating axons of mushroom body gamma-neurons during metamorphosis through the engulfment receptor Draper and its downstream signaling molecule dCED-6, which were originally identified as proteins required for the phagocytosis of apoptotic cells6,7. In another study using mouse models, the classical complement components 1q and 3 (C1q and C3) work as opsonins that tag inappropriate synapses for phagocytosis by microglia8,9,10. There is evidence to suggest that synapse pruning by glial cells is triggered by neuronal activity11,12; however, the molecular mechanisms by which neuronal activity regulates the refinement of particular synaptic connections have not yet been elucidated.
Exosomes are small membrane vesicles of endosomal origin, composed of a lipid bilayer with inserted transmembrane proteins, enclosing cytosolic components derived from their producing cells13. Over the past few years, there has been increasing evidence that exosomes play important roles in intercellular communication networks, enabling the convey of information and the exchange of proteins and lipids between their producing cells and target cells14. Exosomes were also shown to carry mRNAs and microRNAs inside them, raising the possibility that exosomes transfer genetic information between cells15. In the central nervous system, exosomes can be released from all cell types including microglia, oligodendrocytes and neurons and have been proposed to contribute to the physiology of the nervous system and to the neuron-glia communication16,17. In particular, the findings that secretion of exosomes from neurons is promoted by depolarization18 and also by synaptic glutamatergic activity19 led us to hypothesize that neuronal exosomes may activate microglia to promote activity-dependent synaptic pruning. To validate this idea, we established an in vitro culture assay for synapse elimination by using rat pheochromocytoma PC12 cells. Using this assay, we demonstrated that microglia promote the clearance of degenerating neurites of PC12 cells by phagocytosis. We then found that incubating microglia with exosomes secreted from PC12 cells after depolarization increased the expression levels of C3 in the microglia, which enhanced the microglial clearance of degenerating neurites. These results indicate that exosomes secreted by neuronal activity are one of the critical regulators of synaptic pruning by stimulating microglial phagocytosis.
Results
Synaptic development and elimination of PC12 cells
The PC12 cell line has been a valuable model for studying the neurotropic and differentiating effects of nerve growth factor (NGF)20,21. When cultured in a serum-free medium in the presence of NGF, the differentiation and neurite outgrowth of PC12 cells are induced and the extended neurites are connected to each other forming synapse-like structures with many dense core-vesicles and a clathrin-coated membrane invagination22. In the previous reports, Tau-1 and synaptotagmin, markers of axons and presynaptic vesicles, were identified in the tips of neurites, which were connected to regions containing MAP-2 and drebrin, markers of dendrites and postsynaptic membranes22,23. We confirmed that the adult isoform of drebrin, drebrin A, which is exclusively localized in the postsynaptic sites of mature neurons24, was found in the regions where neurites from other cells are connected to form synapse-like structures (Fig. 1a). Thus, NGF-treated PC12 cells exhibit the properties of mature, terminally differentiated neurons forming synapses; but, unlike primary neurons, PC12 cells can survive in serum-containing medium whether or not NGF is present25.
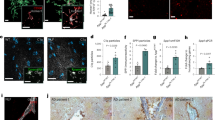
An in vitro assay for synaptic development and elimination.
(a) Differentiation and neurite outgrowth of PC12 cells were induced by culturing the cells in sf-DMEM containing 100 ng/ml NGF for 7 days. The cells were stained with antibody against drebrin A (green) and their staining profile was merged with the phase contrast image in the third panel. Scale bar, 10 μm. (b) The differentiated PC12 cells were further cultured for 2 days after replacing the culture medium with sf-DMEM in the presence (+) or absence (−) of NGF, or with NGF(−) DMEM/10F. In the third panel, Q-VD (2 μM), a potent apoptosis inhibitor, was added to NGF(−) sf-DMEM. Scale bar, 40 μm. (c) Quantification of neurite length of PC12 cells. Number of neurites longer than one cell body diameter were counted and their length were measured by using FV10i-LIV confocal microscope and FLUOVIEW software, Ver. 4.1 (Olympus, Japan). Arrows indicate representative cells that were quantified in the lower table. (d) Cells with more than one neurite were defined as neurite (+) cells and the percentage of neurite (+) cells per total number of cells was determined. (e) The average neurite length per cell was calculated as the sum of total neurite length divided by the number of neurites from a cell. 100 cells were quantified in each condition (cells were selected from 10 different fields in three independent experiments) and the average values were plotted with standard deviations (SD). *p = 0.005, **p = 0.0014; n.s., not significant, ANOVA with Bonferroni correction for multiple comparison.
When PC12 cells were differentiated in serum-free DMEM (sf-DMEM) containing 100 ng/ml NGF for 7 days, about 70–80% of PC12 cells extended neurites and formed synapse-like structures (Fig. 1b). However, when the differentiated PC12 cells were deprived of NGF, the cells underwent apoptosis and their neurites were degenerated, which were not completely inhibited by adding a potent apoptosis inhibitor Q-VD26. On the other hand, when the differentiated PC12 cells were deprived of NGF and cultured in DMEM containing 10% FBS (DMEM/10F), the cells did not undergo apoptosis and started to grow again. Nevertheless, in the absence of NGF, most of the neurites were degenerated and eliminated within 2 days. For quantitative analysis, we defined the cells that retained more than one neurite as neurite (+) cells and measured the length of each neurite (Fig. 1c). Then, the percentage of neurite (+) cells per total number of cells was determined (Fig. 1d) and the average neurite length was quantified as the sum of total neurite length divided by the number of neurites from a cell (Fig. 1e).
Clearance of degenerating neurites by microglia
Using this assay and quantification method, we examined the functions of microglia in removing the degenerating neurites by co-culturing PC12 cells with MG6, a mouse microglial cell line27,28. To distinguish between these two cell lines, the differentiated PC12 cells were labelled with CMFDA and visualized by confocal laser scanning microscopy. When PC12 cells were cultured in NGF(+) sf-DMEM, their neurites were intact and the presence of MG6 cells did not induce synapse elimination within 8 h (Fig. 2a). On the other hand, when the PC12 cells were cultured in NGF(−) DMEM/10F for 8 h, their neurites started to degenerate while most of the neurites were still retained at that point, however the presence of MG6 cells remarkably induced the removal of the degenerating neurites (Fig. 2b and c). This is not caused by the enhanced phagocytic ability of MG6 cells, because the expression of CD68, a microglia activation marker was not altered by the removal of NGF (Fig. 2d). Time-lapse imaging revealed that the MG6 cells were closely associated with the degenerating neurites of PC12 cells, promoting the removal of the neurites by phagocytosis (Fig. 2e). These data indicate that the in vitro cell culture assay established here is a good model for studying the mechanisms of synaptic pruning by microglia.
Microglia promote removal of degenerating neurites of PC12 cells.
(a) After culturing with NGF for 7 days, the differentiated PC12 cells were labelled with CMFDA (green). The PC12 cells were further cultured either in NGF(+) sf-DMEM or NGF(−) DMEM/10F in the absence or presence of MG6 cells for 8 h. PC12 cells and their neurites were visualized by confocal microscopy. Phase contrast images are shown in the lower panels, where MG6 cells are pointed by red arrows. Scale bar, 20 μm. The percentage of PC12 cells that retain neurites (b) and the average length of neurites (c) were quantified from 100 cells in each condition (cells were selected from 10 different fields in three independent experiments) and the average values were plotted with SD. *p = 0.0027, **p < 0.001; n.s., not significant. Student's t test. (d) Merged images of the phase contrast and CD68 staining (red) of MG6 cells that were co-cultured with PC12 cells either in NGF(+) sf-DMEM or NGF(−) DMEM/10F. Scale bar, 20 μm. (e) CMFDA-labelled PC12 cells (green) were co-cultured with PKH26-labelled MG6 cells (red) in NGF(−) DMEM/10F. Time-lapse images after replacing the culture medium were taken by using a confocal microscope equipped with culture chamber kept at 37°C and 5% CO2. Arrows indicate MG6 cells engulfing the degenerating neurites. Scale bar, 10 μm.
Microglia mediate synaptic pruning in PS-independent manner
In Drosophila, glial cells remove the degenerating axons via Draper, Crk, dCED-6, dCed-12, which were originally identified as proteins required for the clearance of apoptotic cells6,7,29. These findings raised the possibility that synaptic pruning and apoptotic cell clearance may share a common molecular machinery. When cells undergo apoptosis, the dying cells expose phosphatidylserine (PS) on their surface, a component of the cell plasma membrane that is kept exclusively in the inner leaflet of the lipid bilayer of healthy cells30. The exposed PS on the apoptotic cells is then recognized by the receptors on macrophages. We therefore examined whether PS works as an “eat me” signal for the synaptic pruning.
When the differentiated PC12 cells were co-cultured with MG6 cells in NGF(−) DMEM/10F for 8 h, the degenerating neurites of PC12 cells were efficiently removed by MG6 cells (Fig. 3a). The percentage of PC12 cells that retained neurites was decreased from 81.4% to 25.0% and the average length of neurites was decreased from 75.6 μm to 41.7 μm when MG6 cells were cultured together with PC12 cells (Fig. 3b and c). On the other hand, when PC12 cells were co-cultured with the NIH3T3 transformants expressing Tim4 (3T3/Tim4), which efficiently promote the clearance of apoptotic cells by recognizing PS31, the synaptic pruning of PC12 cells was not promoted. Furthermore, the addition of recombinant D89E mutant of MFG-E8 (rD89E), which inhibits PS-dependent phagocytosis32, to the culture medium did not affect the efficiency of the synaptic pruning by the MG6 cells. These results suggest that PS is dispensable for the clearance of degenerating neurites by microglia.
Synaptic pruning is not mediated by PS.
(a) After differentiation in NGF(+) sf-DMEM for 7 days, PC12 cells were labelled with CMFDA (green) and further cultured in NGF(−) DMEM/10F for 8 h in the absence or presence of 3T3/Tim4 or MG6 cells. When indicated, recombinant D89E mutant of MFG-E8 (rD89E, 3 μg/ml) that masks PS was added into the culture medium during co-culture. Phase contrast images are shown in the lower panels. Scale bar, 20 μm. The percentage of PC12 cells that retain neurites (b) and the average length of neurites (c) were quantified from 100 cells in each condition (cells were selected from 10 different fields in three independent experiments) and the average values were plotted with SD. *p < 0.001; n.s., not significant. ANOVA with Bonferroni correction for multiple comparison.
Exosomes enhance microglial ability to remove degenerating neurites
It has long been indicated that synapse elimination and engulfment by glial cells are triggered by neuronal activity11,12; however the means whereby neuronal activity regulates these processes have been elusive. Fauré et al. found that potassium-induced depolarization of neurons strongly enhances the exosome release from the neurons18. Given that microglia are the primary target of neuronal exosomes16, we examined the effects of exosomes from PC12 cells on the functions of microglia. Nanoparticle tracking analysis34 revealed that secretion of exosomes from differentiated PC12 cells was enhanced by depolarizing the cells with 25 mM KCl for 3 h (Fig. 4a). We confirmed that the average diameter of the exosomes was about 100 nm, which corresponds to the typical size of exosomes. We also confirmed that the exosomes from PC12 cells carried the typical exosomal markers33, such as MFG-E8 and flotillin-2 (Fig. 4b). When MG6 cells were pre-incubated for 16 h with the exosomes secreted from PC12 cells after depolarization, general phagocytic ability of MG6 cells, such as phagocytosis of E.coli, was not altered (Fig. 4c). Then, the ability for synaptic pruning was compared between MG6 cells pre-incubated with or without the exosomes. Interestingly, MG6 cells that have engulfed exosomes exhibited a stronger ability to remove degenerating neurites of PC12 cells than control MG6 cells without exosomes (Fig. 4d). Consequently, the addition of exosomes to MG6 cells reduced the percentage of PC12 cells that retained neurites from 26.2% to 2.1% and the average length of neurites from 40.0 μm to 1.98 μm (Fig. 4e). Unlike exosomes from PC12 cells, exosomes from NIH3T3 cells did not promote the removal of degenerating neurites of PC12 cells, suggesting cell type-specific effects of the exosomes (Fig. 4f). From these data, we considered the possibility that exosomes might up-regulate molecules in microglia that specifically accelerate the synaptic pruning.
Exosomes promote synaptic pruning by microglia.
(a) Exosomes were collected by ultra-centrifugation from the culture medium of differentiated PC12 cells that were cultured in sf-DMEM with or without 25 mM KCl for 3 h. Nanoparticle tracking analysis was performed to determine the density (left graph) and size distribution (right panel) of the exosomes in the medium. The experiments were performed in triplicate and the average values with SD were plotted. *p = 0.0015, Student's t test. (b) Exosomes secreted from PC12 cells after depolarization were collected by ultra-centrifugation and analysed by western blot using antibodies against MFG-E8 and flotillin-2, exosomal markers. (c) Phagocytosis of pHrodo Red E.coli BioParticles by MG6 cells that were exposed (solid lines) or not exposed (dot lines) to the exosomes from differentiated PC12 cells after depolarization was allowed to proceed for 2 h. The efficiency of phagocytosis was compared by FACS analysis. (d) Differentiated PC12 cells were cultured in NGF(−) DMEM/10F for 8 h in the absence or presence of MG6 cells or MG6 cells that have engulfed the exosomes from PC12 cells (red). After labelling with PKH26, the exosomes were incubated with MG6 cells for 16 h. Scale bar, 40 μm. (e) The percentage of PC12 cells that retain neurites and the average length of neurites were quantified from 100 cells in each condition (cells were selected from 10 different fields in three independent experiments). The average values were plotted with SD. *p < 0.001. ANOVA with Bonferroni correction for multiple comparison. (f) The same assay was performed with MG6 cells that have engulfed the equal amounts of exosomes (exo) from PC12 cells or NIH3T3 cells. *p = 0.0013, **p = 0.003, ***p < 0.001; n.s., not significant. ANOVA with Bonferroni correction for multiple comparison.
Complement factors induced by exosomes promote synaptic pruning
We therefore tried to identify the microglial mediators of synaptic pruning that were up-regulated by the exosomes. We examined the gene expression profiles of MG6 cells with a microarray consisting of about 40,000 mouse genes. The expression levels of 183 genes were increased more than 8-fold in MG6 cells incubated with exosomes compared to the control MG6 cells without exosomes (Fig. 5a). Functional enrichment analysis identified a number of immune-related processes significantly enriched, including “Phagosome” and “Complement and coagulation cascades” (Fig. 5b). Among them, we selected following genes with pro-phagocytic activity as candidate mediators: complement factor B (Cfb), complement component 3 (C3), macrophage receptor with collagenous structure (MARCO) and oxidized low density lipoprotein receptor 1 (Olr1). To validate the results of the microarray screening, we performed quantitative PCR (qPCR) analyses of these candidate genes and demonstrated that Cfb and C3-encoding genes were highly up-regulated (56-fold and 25-fold, respectively) in MG6 cells after incubation with exosomes secreted from PC12 cells (Fig. 5c and d). As we could not detect sufficient expression levels of C3 mRNA in the exosomes per se, it is likely that exosomes stimulated the transcriptional up-regulation of C3 in MG6 cells, rather than transferring C3 mRNA (Fig. 5e).
Exosomes up-regulate the expression of complement factors in microglia.
(a) A screening for differentially expressed genes (DEGs) in MG6 cells exposed vs. not exposed to exosomes secreted from differentiated PC12 cells was performed using microarray analysis. Two arrays per condition were used and DEGs were selected as probes detected in at least one array and with an absolute log2 fold change (M) greater than 3 (i.e. × 8 expression change). This approach resulted in 275 probes differentially expressed, which mapped to 183 known genes. (b) Functional enrichment of KEGG pathways was performed to identify the biological processes altered in microglia exposed to exosomes. Enrichment analysis resulted in a number of immune related processes significantly enriched (p < 0.05), including “Phagosome” (H2-M2, Marco, C3, Olr1, Tlr2) and “Complement and coagulation cascades” (C3, Cfb). (c) qPCR analyses were performed to validate the microarray results using gene-specific primers and RNA samples extracted from MG6 cells incubated 16 h with (black bar) or without (white bar) exosomes. Analysis was done with delta Ct method. Graphs represent gene expression levels relative to GAPDH. The experiments were performed in triplicate and the average values with SD were plotted. p < 0.001, Student's t test. (d) The gel photograph of the qPCR products is shown. (e) qPCR analysis was performed to compare C3 mRNA levels between total purified exosomes and MG6 cells that have engulfed the same amount of the exosomes. Graphs represent gene expression levels relative to GAPDH. N.D. stands for not detectable. *p < 0.001, Student's t test. (f) Differentiated PC12 cells were cultured in NGF(−) DMEM/10F for 8 h with MG6 cells or MG6 cells that have engulfed the exosomes (red). Neutralizing anti-C3a/C3 antibody (+ α-C3 Ab) or IgG isotype control antibody (+ Ctrl. Ab) was added into the culture medium at the concentration of 20 μg/ml during co-culture. Scale bar, 40 μm. (g) The percentage of PC12 cells that retain neurites and the average length of neurites were quantified from 100 cells in each condition (cells were selected from 10 different fields in three independent experiments). The average values were plotted with SD. *p < 0.001; n.s., not significant. ANOVA with Bonferroni correction for multiple comparison.
The essential roles of complement factors, in particular C3, in synaptic pruning have been well established8,9,10. In order to ascertain the effect of C3 expression on the phagocytic activity of MG6 cells, we treated MG6 cells that were activated by the exosomes with neutralizing anti-C3a/C3 antibody which blocks the cleavage and consequently the activation of C335. As shown in Fig. 5f, the blockade of C3 by the antibody abolished the exosome-induced enhancement of synaptic pruning by the microglia. As a result, anti-C3a/C3 antibody but not by IgG isotype control antibody increased the percentage of PC12 cells that retained neurites from 1.6% to 18.5% and the average length of neurites from 3.6 μm to 18.9 μm, close to the levels of PC12 cells co-cultured with control MG6 cells without exosomes (Fig. 5g). Taken together, these data suggest that the exosomes secreted by neuronal activity promote synaptic pruning by up-regulating the expression of complement factors in microglia.
Discussion
In this study, we demonstrated that PC12 cells provide a simple and reproducible in vitro model to study mechanisms of synaptic development and elimination. PC12 cells can be grown indefinitely; their availability and ease of culture make them a good alternative for in vivo studies which are time-consuming and technically demanding, precluding high-throughput experimentation. Microfluidic systems36 and compartmentalized chambers37, based on fluidic isolation of neurites from the cell body, have been widely used to study the effects of neurotropic factors for axon outgrowth and pruning. These chambers, however, are difficult to assemble and to maintain continuous control of the hydrostatic pressure difference and fluidic isolation between compartments38. The in vitro assay using PC12 cells established here is a feasible method for clarifying the molecular mechanisms of synaptic pruning. However, it is important to bear in mind the limitation of this in vitro assay, as PC12 cells are cell lines with some genetic abnormalities which may not reflect the in vivo phenomenon. In addition, this time we used MG6 cells, a mouse microglial cell line immortalized by human c-myc27,28 for the synaptic pruning of rat PC12 cells, due to its availability and ease of use, but it might be modified by using rat primary microglia in the future studies.
Several studies raised the possibility that synaptic pruning and clearance of apoptotic cells may share a common molecular mechanism6,7,29. In accordance with previous studies in dorsal root ganglion model of Wallarian degeneration39, our preliminary experiment indicated that degenerating neurites of PC12 cells express PS on their surface as determined by a positive staining with annexin V. Nevertheless, masking PS by MFG-E8/rD89E did not prevent the synaptic pruning by MG6 cells. Instead, we confirmed that complement factors actually regulate the synaptic pruning, which is consistent with the previous in vivo findings using mice deficient in C3 or its receptor8,9. Recently, Barres and colleagues have reported that not only microglia but also astrocytes mediate synapse elimination40. Interestingly, unlike microglia, astrocytes did not use complement factors for the synaptic pruning. Instead, they found that astrocytes promoted synaptic pruning through MEGF10, a mammalian ortholog of drosophila Draper and MERTK, which mediates PS-dependent phagocytosis of apoptotic cells via Gas6/Protein-S. Because expression of Draper is up-regulated by axonal injury in Drosophila7, it would be interesting to examine whether neuronal exosomes also up-regulate the expression of MEGF10 and MERTK in astrocytes to promote the synaptic pruning.
The secretion of exosomes from PC12 cells was enhanced by KCl-induced depolarization. Given that exosome secretion is often regulated by a calcium stimuli41, the KCl-induced depolarization might raise the intracellular calcium levels required for the exosome secretion. The exosomes secreted by PC12 cells were engulfed by MG6 cells, which exhibited enhanced phagocytic activity of degenerating neurites. We then examined the effect of exosomes on the gene expression profiles of MG6 cells and demonstrated that several pro-phagocytic genes, including C3-encoding gene, were up-regulated to enhance the microglial ability for synaptic pruning. However, it should be noted that exosome-mediated microglial activation is not the only prominent mechanism for synaptic pruning, but may be one important mechanism among others, because microglia without exosomes are still capable of reducing neurites and its length to great extent. The mechanisms for synaptic pruning by non-stimulated microglia remain to be elucidated.
Our results indicate that blocking C3 significantly protects neurites from engulfment by microglia, suggesting a role for this component in opsonisation of degenerating neurites. Consistently with our finding, Stevens and colleagues have shown that components of the classical complement pathway including C3 are key mediators of neurites and synapse elimination by microglia during development8,9. C3 was specifically up-regulated in microglia in the P5 lateral geniculate nucleus and down-regulated by P9, an age when pruning is largely complete. Nevertheless, the mechanisms that induce the expression of complement factors for synaptic pruning are as yet unclear. It would be interesting to examine whether exosomes are involved in this process. In addition, a model proposed over two decades ago for how activity may drive synapse elimination suggested that strong synapses, which are effective in driving postsynaptic responses, actively punish and eliminate nearby weaker synapses42. However, the entity of the “punishment” and the means whereby it promotes the synaptic pruning have not been identified. From our data, we propose a new model in which exosomes secreted from activated neurons act as the “punishment” signal for the weaker synapses (but preserving the stronger ones) by inducing the complement C3 in microglia for phagocytic clearance of the inappropriate synapses.
Synaptic pruning by microglia has been implicated not only in neural development but also in pathologies of the central nervous system2. These include neurodegenerative diseases and multiple sclerosis, in which activation of microglia is thought to play a critical role in the loss of neurons or axons43. Neuronal exosomes have also been implicated in the development of neurodegenerative diseases, as they carry several disease-related proteins such as β-amyloid, α-synuclein and TDP-43, which are involved in the pathogenesis of Alzheimer's disease, Parkinson's disease and amyotrophic lateral sclerosis, respectively44,45,46. A better understanding of the molecular mechanisms underlying synaptic pruning by microglia and its regulatory mechanisms by neuronal exosomes will provide new pathophysiological insights into these diseases. We hope the in vitro culture system established here will provide a good model system to reveal avenues towards new therapeutic approaches.
Methods
Cell Lines and Reagents
PC12 cells and MG6 cells27,28 were obtained from RIKEN BRC, Japan. PC12 cells were maintained in DMEM (Nakarai, Japan) supplemented with 10% FBS (Biowest) and 10% horse serum (Life technologies) and MG6 cells were maintained in DMEM supplemented with 10% FBS, 10 μg/ml insulin and 100 μM 2-mercaptoethanol. Recombinant rat beta-NGF and pan-caspase OPH inhibitor Q-VD26 were from R&D systems. Monoclonal antibody against C3a/C3 (clone K13/16) and IgG isotype control antibody were from BioLegend. The NIH3T3 transformants expressing Tim4 (3T3/Tim4) and rD89E were described previously31,32. When indicated, membranes of MG6 cells or exosomes were labelled with PKH26 red fluorescent cell linker kit (Sigma) according to the manufacturer's instructions.
Induction of Neurite Degeneration
PC12 cells (1 × 105 cells) were plated in Millicell EZ slide 8-well glass (Merck Millipore), which were pre-coated with 0.05% polyethyleneimine (PEI, Sigma). After overnight culture, differentiation and neurite outgrowth of PC12 cells were induced by replacing the medium with sf-DMEM containing 100 ng/ml NGF and the cells were cultured for 7 days. Fresh medium containing NGF was added every second day. For some experiments, differentiated PC12 cells were labelled with 2 μM CellTracker Green CMFDA (Life technologies) according to the manufacturer's instructions. To induce neurite degeneration, NGF was removed by washing the PC12 cells three times with DMEM/10F and cultured in NGF(−) DMEM/10F for 8 h in the absence or presence of MG6 cells (1 × 105 cells) or 3T3/Tim4 (2 × 104 cells). When indicated, rD89E (3 μg/ml), anti-C3a/C3 monoclonal antibody (20 μg/ml) or IgG isotype control antibody was added into the culture medium during co-culture. For microscopic observations, cells were washed once with PBS and then fixed in 4% paraformaldehyde solution for 10 min at room temperature (RT). The culture slides were washed twice by agitation in PBS for 5 min before mounting with VECTASHIELD mounting medium (Vector laboratories) and observed by FV10i-LIV confocal microscope (Olympus, Japan). Neurite length was determined by FLUOVIEW software, Ver. 4.1 (Olympus, Japan). The percentage of PC12 cells that retain neurites (% cells with neurites) and the average length of neurites were quantified by analysing 100 cells selected from 10 different fields in three independent experiments.
Immunocytochemistry
PC12 cells and/or MG6 cells (1 × 105 cells) cultured in Millicell EZ slide 8-well glass were washed with PBS and fixed in 4% paraformaldehyde solution for 10 min at RT. The cells were permeabilized with 0.1% Triton X-100 in PBS for 15 min and then blocked with PBS containing 5% goat serum (Sigma) and 1% BSA for 1 h. PC12 cells were stained in PBS containing 1% BSA with rabbit anti-drebrin A antibody (Takara, Japan), which was followed by staining with DyLight 488-conjugated anti-rabbit IgG antibody (BioLegend). MG6 cells were stained with rat anti-CD68 antibody (BioLegend), followed by staining with Cy3-conjugated anti-rat IgG antibody (BioLegend). After staining, cells were mounted with VECTASHIELD mounting medium and observed by FV10i-LIV confocal microscope.
Exosome Induction and Purification
PC12 cells (1 × 106 cells) were cultured in NGF(+) sf-DMEM for 7 days in 6-cm culture dishes that were pre-coated with 0.05% PEI. Preparation of exosomes was carried out following the protocol described by Fauré et al.18. In brief, exosome secretion was induced by depolarizing the differentiated PC12 cells by replacing the medium with sf-DMEM containing 25 mM KCl and culturing for 3 h. The culture medium was collected and cleared of debris by two successive centrifugations (2,000 × g for 10 min and 20,000 × g for 20 min). Exosomes were then precipitated from the medium by ultra-centrifugation at 100,000 × g for 2 h with himac CS100FNX micro ultracentrifuge (Hitachi Koki, Japan). The precipitated exosomes were labelled with PKH26, suspended in 3 ml DMEM/10F and incubated with MG6 cells (1 × 106 cells) in 6-cm culture dish. After 16 h, MG6 cells were harvested with 0.25% trypsin and 1 mM EDTA and used for the culture with PC12 cells.
Characterization of exosomes
PC12 cells (3 × 106 cells) were cultured in NGF(+) sf-DMEM for 7 days in 10-cm culture dishes and then cultured with or without KCl depolarization for 3 h. Exosomes were collected from the culture medium (9 ml) by ultra-centrifugation and suspended in 50 μl of sf-DMEM. The density and size distribution of exosomes were analysed using NanoSight LM10-HS apparatus and NTA2.3 software (Malvern). The exosomes samples were also loaded onto 10% polyacrylamide gel, separated at 200 V for 30 min and detected by western blot using monoclonal antibodies against MFG-E8 (MBL, Japan) and flotillin-2 (BD Biosciences).
Phagocytosis of E.coli BioParticles
MG6 cells (1 × 105) exposed or not exposed to the exosomes secreted from the differentiated PC12 cells were cultured in 0.5 ml of DMEM/10F in a 24-well microtitre plates. 10 μg of pHrodo Red E.coli BioParticles (Life technologies) were added to the MG6 cells and the culture was incubated at 37°C for 2 h. The cells were detached from the plate by incubating them with 0.25% trypsin and 1 mM EDTA and collected by centrifugation. In order to reduce the background signal of pHrodo under neutral conditions, the cells were suspended in 300 μl of 20 mM N-cyclohexyl-2-aminoethanesulfonic acid (CHES)–NaOH buffer (pH 9.0) containing 150 mM NaCl and 2% FCS as previously reported47 and analysed by flow cytometry on the FACSVerse apparatus (BD Biosciences).
Microarray Analysis
MG6 cells (1 × 106 cells) were incubated in 6-cm dishes for 16 h with or without exosomes secreted from PC12 cells (1 × 106 cells) after depolarization as described above. Total RNA was extracted by RNeasy Plus Mini Kit (QIAGEN) following the kit instructions. Microarray analysis was performed by Medical & Biological Laboratories (Japan) using SurePrint G3 Mouse Gene Expression 8x60K Microarrays (Agilent). Pre-processing of microarray images was performed using the software Feature Extraction version 10.10.1.1 (Agilent). Normalization was done with the Percentile 75 method implemented in the software GeneSpring GX version 12.6.1 (Agilent). Gene selection and enrichment analyses were performed in R version 3.1.2 (www.r-project.org). Differentially expressed genes were identified by plotting an MA plot (M = difference of log2 intensities between two conditions; A, average of log2 intensities of arrays) and selecting probes flagged as Detected by the FE software in any of the arrays and with an absolute log2 fold change greater than 3 (i.e. × 8 expression change). Functional enrichment analysis was performed using pathway information from the Kyoto Encyclopaedia of Genes and Genomes (KEGG; www.kegg.jp) with a hypergeometric test implemented in the GOstats Bioconductor package version 2.32.048. Pathways were considered significantly enriched at p < 0.05.
Quantitative PCR analysis
cDNA was synthesized with High Capacity RNA-to-cDNA Kit (Life technologies) and the cDNA products were amplified with a LightCycler 96 (Roche) using SYBR qPCR Mix (TOYOBO, Japan). The data were analysed with delta Ct method and normalized to the amounts of GAPDH RNA expression in each sample. The primers used for real-time PCR were as follows: Cfb-Fw (5′-GCAAGCAGCACAAGGAACAGTTG-3′), Cfb-Rv (5′-CAGGCTCTTCCCTTGCTCAGATAC-3′), MARCO-Fw (5′-AAACCTGGAGTGCAGGGTGTTC-3′), MARCO-Rv (5′-TCACCCTTGGCACCTGAAAGTC-3′), C3-Fw (5′-AGGGCAGCAACGCAAGTTCATC-3′), C3-Rv (5′-TTCTGCAGCTTCAGGGCGTTTC-3′), Olr1-Fw (5′-CCTCCTGTTGCTGCATGAAAGAC-3′), Olr1-Rv (5′- AGCTTTGCCTTTGAGCCCTCTG-3′), GAPDH-Fw (5′-TGTGTCCGTCGTGGATCTGA-3′), GAPDH-Rv (5′-TTGCTGTTGAAGTCGCAGGAG-3′).
References
Goda, Y. & Davis, G. W. Mechanisms of synapse assembly and disassembly. Neuron 40, 243–264 (2003).
Morris, G. P., Clark, I. A., Zinn, R. & Vissel, B. Microglia: a new frontier for synaptic plasticity, learning and memory and neurodegenerative disease research. Neurobiol Learn Mem 105, 40–53 (2013).
Kantor, D. B. & Kolodkin, A. L. Curbing the excesses of youth: molecular insights into axonal pruning. Neuron 38, 849–852 (2003).
Eroglu, C. & Barres, B. A. Regulation of synaptic connectivity by glia. Nature 468, 223–231 (2010).
Chung, W. S. & Barres, B. A. The role of glial cells in synapse elimination. Curr Opin Neurobiol 22, 438–445 (2012).
Awasaki, T. et al. Essential role of the apoptotic cell engulfment genes draper and ced-6 in programmed axon pruning during Drosophila metamorphosis. Neuron 50, 855–867 (2006).
MacDonald, J. M. et al. The Drosophila cell corpse engulfment receptor Draper mediates glial clearance of severed axons. Neuron 50, 869–881 (2006).
Stevens, B. et al. The classical complement cascade mediates CNS synapse elimination. Cell 131, 1164–1178 (2007).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Stephan, A. H., Barres, B. A. & Stevens, B. The complement system: an unexpected role in synaptic pruning during development and disease. Annu Rev Neurosci 35, 369–389 (2012).
Katz, L. C. & Shatz, C. J. Synaptic activity and the construction of cortical circuits. Science 274, 1133–1138 (1996).
Hua, J. Y. & Smith, S. J. Neural activity and the dynamics of central nervous system development. Nat Neurosci 7, 327–332 (2004).
Thery, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nat Rev Immunol 9, 581–593 (2009).
Turturici, G., Tinnirello, R., Sconzo, G. & Geraci, F. Extracellular membrane vesicles as a mechanism of cell-to-cell communication: advantages and disadvantages. Am J Physiol Cell Physiol 306, C621–633 (2014).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9, 654–659 (2007).
Cossetti, C. et al. Extracellular membrane vesicles and immune regulation in the brain. Front Physiol 3, 117 (2012).
Fruhbeis, C., Frohlich, D. & Kramer-Albers, E. M. Emerging roles of exosomes in neuron-glia communication. Front Physiol 3, 119 (2012).
Faure, J. et al. Exosomes are released by cultured cortical neurones. Mol Cell Neurosci 31, 642–648 (2006).
Lachenal, G. et al. Release of exosomes from differentiated neurons and its regulation by synaptic glutamatergic activity. Mol Cell Neurosci 46, 409–418 (2011).
Levi, A. & Alema, S. The mechanism of action of nerve growth factor. Annu Rev Pharmacol Toxicol 31, 205–228 (1991).
Klesse, L. J., Meyers, K. A., Marshall, C. J. & Parada, L. F. Nerve growth factor induces survival and differentiation through two distinct signaling cascades in PC12 cells. Oncogene 18, 2055–2068 (1999).
Jeon, C. Y. et al. Neurites from PC12 cells are connected to each other by synapse-like structures. Synapse 64, 765–772 (2010).
Krasnov, P. A. & Enikolopov, G. Targeting of synaptotagmin to neurite terminals in neuronally differentiated PC12 cells. J Cell Sci 113 (Pt8), 1389–1404 (2000).
Hayashi, K. et al. Modulatory role of drebrin on the cytoskeleton within dendritic spines in the rat cerebral cortex. J Neurosci 16, 7161–7170 (1996).
Greene, L. A. & Tischler, A. S. Establishment of a noradrenergic clonal line of rat adrenal pheochromocytoma cells which respond to nerve growth factor. Proc Natl Acad Sci U S A 73, 2424–2428 (1976).
Caserta, T. M., Smith, A. N., Gultice, A. D., Reedy, M. A. & Brown, T. L. Q-VD-OPh, a broad spectrum caspase inhibitor with potent antiapoptotic properties. Apoptosis 8, 345–352 (2003).
Takenouchi, T., Ogihara, K., Sato, M. & Kitani, H. Inhibitory effects of U73122 and U73343 on Ca2+ influx and pore formation induced by the activation of P2X7 nucleotide receptors in mouse microglial cell line. Biochim Biophys Acta 1726, 177–186 (2005).
Nakamichi, K. et al. Suppressive effect of simvastatin on interferon-beta-induced expression of CC chemokine ligand 5 in microglia. Neurosci Lett 407, 205–210 (2006).
Lu, T. Y., Doherty, J. & Freeman, M. R. DRK/DOS/SOS converge with Crk/Mbc/dCed-12 to activate Rac1 during glial engulfment of axonal debris. Proc Natl Acad Sci U S A 111, 12544–12549 (2014).
Nagata, S., Hanayama, R. & Kawane, K. Autoimmunity and the clearance of dead cells. Cell 140, 619–630 (2010).
Miyanishi, M. et al. Identification of Tim4 as a phosphatidylserine receptor. Nature 450, 435–439 (2007).
Hanayama, R. et al. Identification of a factor that links apoptotic cells to phagocytes. Nature 417, 182–187 (2002).
Mignot, G., Roux, S., Thery, C., Segura, E. & Zitvogel, L. Prospects for exosomes in immunotherapy of cancer. J Cell Mol Med 10, 376–388 (2006).
Dragovic, R. A. et al. Sizing and phenotyping of cellular vesicles using Nanoparticle Tracking Analysis. Nanomedicine 7, 780–788 (2011).
Elsner, J., Oppermann, M., Czech, W. & Kapp, A. C3a activates the respiratory burst in human polymorphonuclear neutrophilic leukocytes via pertussis toxin-sensitive G-proteins. Blood 83, 3324–3331 (1994).
Park, J. W., Vahidi, B., Taylor, A. M., Rhee, S. W. & Jeon, N. L. Microfluidic culture platform for neuroscience research. Nat Protoc 1, 2128–2136 (2006).
Campenot, R. B., Walji, A. H. & Draker, D. D. Effects of sphingosine, staurosporine and phorbol ester on neurites of rat sympathetic neurons growing in compartmented cultures. J Neurosci 11, 1126–1139 (1991).
Millet, L. J. & Gillette, M. U. New perspectives on neuronal development via microfluidic environments. Trends Neurosci 35, 752–761 (2012).
Sievers, C., Platt, N., Perry, V. H., Coleman, M. P. & Conforti, L. Neurites undergoing Wallerian degeneration show an apoptotic-like process with Annexin V positive staining and loss of mitochondrial membrane potential. Neurosci Res 46, 161–169 (2003).
Chung, W. S. et al. Astrocytes mediate synapse elimination through MEGF10 and MERTK pathways. Nature 504, 394–400 (2013).
Savina, A., Furlan, M., Vidal, M. & Colombo, M. I. Exosome release is regulated by a calcium-dependent mechanism in K562 cells. J Biol Chem 278, 20083–20090 (2003).
Balice-Gordon, R. J. & Lichtman, J. W. Long-term synapse loss induced by focal blockade of postsynaptic receptors. Nature 372, 519–524 (1994).
Gonzalez, H., Elgueta, D., Montoya, A. & Pacheco, R. Neuroimmune regulation of microglial activity involved in neuroinflammation and neurodegenerative diseases. J Neuroimmunol 274, 1–13 (2014).
Rajendran, L. et al. Alzheimer's disease beta-amyloid peptides are released in association with exosomes. Proc Natl Acad Sci U S A 103, 11172–11177 (2006).
Danzer, K. M. et al. Exosomal cell-to-cell transmission of alpha synuclein oligomers. Mol Neurodegener 7, 42 (2012).
Nonaka, T. et al. Prion-like properties of pathological TDP-43 aggregates from diseased brains. Cell Rep 4, 124–134 (2013).
Toda, S., Hanayama, R. & Nagata, S. Two-step engulfment of apoptotic cells. Mol Cell Biol 32, 118–125 (2012).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Acknowledgements
We thank W. Nakai and K. Yasuda for technical and secretarial assistance. I.B. and J.S. are supported by a research fellowship from IFReC Kishimoto Foundation and Japan Society for the Promotion of Science, respectively. This work was supported by grants from JST, Ministry of Education, Culture, Sports, Science and Technology (MEXT, No. 25713020 to R.H. and No. 25430182 to D.D.), Grants-in-Aid for Scientific Research on Innovative Areas (“Brain Environment”) from MEXT (No. 24111530 and No. 26111714 to R.H.), the Banyu Foundation, the Cell Science Research Foundation, the Kishimoto Foundation, the Mochida Memorial Foundation, the Naito Foundation, the Sumitomo Foundation, the Suzuken Memorial Foundation, the Takeda Science Foundation, the Tokyo Biochemical Research Foundation, the Uehara Memorial Foundation and a Human Frontier Science Program Career Development Award to R.H.
Author information
Authors and Affiliations
Contributions
I.B. and R.H. designed research; I.B. and J.S. performed research; I.B., D.D. and R.H. analysed data and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Bahrini, I., Song, Jh., Diez, D. et al. Neuronal exosomes facilitate synaptic pruning by up-regulating complement factors in microglia. Sci Rep 5, 7989 (2015). https://doi.org/10.1038/srep07989
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07989
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.