Abstract
KCNQ1 channels play vital roles in cardiovascular, gastric and other systems. The conductance and dynamics of KCNQ1 could be modulated by different single transmembrane helical auxiliary proteins (such as KCNE1, KCNE2 and others). In this study, detail KCNQ1 function modulations by different regions of KCNE1 or KCNE2 were examined using combinational methods of electrophysiology, immunofluorescence, solution NMR and related backbone flexibility analysis. In the presence of KCNE2 N-terminus, decreased surface expression and consequent low activities of KCNQ1 were observed. The transmembrane domains (TMDs) of KCNE1 and KCNE2 were illustrated to associate with the KCNQ1 channel in different modes: Ile64 in KCNE2-TMD interacting with Phe340 and Phe275 in KCNQ1, while two pairs of interacting residues (Phe340-Thr58 and Ala244-Tyr65) in the KCNQ1/KCNE1 complex. The KCNE1 C-terminus could modulate gating property of KCNQ1, whereas KCNE2 C-terminus had only minimal influences on KCNQ1. All of the results demonstrated different KCNQ1 function modulations by different regions of the two auxiliary proteins.
Similar content being viewed by others
Introduction
Voltage-gated potassium KCNQ1 (Kv7.1 or KvLQT1) channels are widely expressed in various tissues1,2 such as the brain, heart, pancreas, stomach and intestine. They play crucial roles in the regulation of membrane potential and cellular excitability3,4. The KCNQ1 channel often associates with different KCNE auxiliary subunits to form KCNQ1/KCNEs complexes, resulting in significant changes in gating properties5,6. In cardiac cells, KCNE1 associates with KCNQ1 to form channels having slow delayed rectifier IKs currents. The KCNQ1/KCNE2 channel is essential for gastric acid secretion in gastric parietal cells, especially for the pH sensitivity and for the generation of constitutive K+ currents across the cell membrane7,8,9.
Although KCNE1 and KCNE2 both contain a single transmembrane helix, they may modulate KCNQ1 conductance in different ways10. To illustrate different KCNQ1 modulations by KCNE1 or KCNE2, structural comparisons of the two auxiliary proteins and electrophysiological studies of KCNQ1/KCNE1 or KCNQ1/KCNE2 complexes should be conducted. Previous electrophysiological and biochemical studies of KCNQ1/KCNE1 interactions have shown that the transmembrane domain (TMD) and C-terminus of KCNE1 might be crucial for association and modulation of KCNQ1 function11,12,13,14. Fluorescence resonance energy transfer (FRET), co-immunoprecipitation or physiology studies verified that the cytoplasmic tail of KCNE1 could interact directly with the pore region of KCNQ1 or move close to the C-terminus of KCNQ115,16,17. The availability of the KCNE1 solution NMR structure18,19 enabled the docking analysis of the KCNE1-TMD to the tetrameric KCNQ1 channel20. However, the detailed structure of KCNE2 and the structural basis of the KCNQ1 function modulations by KCNE2 remain unclear. Furthermore, the KCNE1 solution NMR structure was determined alone in detergent micelles and could not explicitly explain the association mode with KCNQ1 in the KCNQ1/KCNE1 complex. Besides docking analysis with KCNQ1 channels, flexibility analysis of KCNE1 or KCNE2 might also provide information for potential interactions with KCNQ1 in KCNQ1/KCNEs complexes.
In the present study, we examined the KCNE1 or KCNE2 mediated regional modulation of KCNQ1 channel conductance by generating different chimeras of KCNE1 and KCNE2 to clarify the roles of the N-terminal regions, TMDs or C-terminal tails in the association with KCNQ1. Immunofluoresence data showed that the N-terminal region of KCNE2 (E2(N)) affected KCNQ1 trafficking to the cell membrane. Using Q-scanning mutation with double mutant cycle analysis and molecular dynamics (MD) based on the NMR structure of the TMDs of two KCNE subunits, models of the different association modes of KCNE1-TMD or KCNE2-TMD to KCNQ1 were developed. Comparisons of structural flexibility between the C-terminal tails of KCNE1 or KCNE2 and their deletion mutants suggested that the C-terminal tail of KCNE2 plays a minor role in the function of the KCNQ1 channel.
Results
KCNE1/KCNE2 chimeras modulate the channel properties of KCNQ1
Sequence alignment of full length KCNE1 and KCNE2 showed 32% identity and 92% similarity among the primary sequences (Fig. S1). The significant variation in the currents between KCNQ1/KCNE1 and KCNQ1/KCNE2 could be related to different modes of association between KCNE1 or KCNE2 and KCNQ1. To examine whether the subdomains of KCNEs (N terminus, TMD or C terminal tail) are involved in the modulation of KCNQ1 current, chimeras of KCNE1 and KCNE2 were constructed (Fig. S2) and co-expressed with KCNQ1 in HEK293 cells.
A previous study showed that the TMD of KCNE1 interacted with the S6 domain of KCNQ121. Two chimeras, E1[E2(T)] and E2[E1(T)] (defined in Fig. S2a), were constructed in which only the TMD was exchanged to examine the possible interactions between the TMDs of KCNEs and KCNQ1. When coexpressed with KCNQ1, E2[E1(T)] showed slowly activated currents with attenuated amplitude, whereas E1[E2(T)] showed rapidly activated currents (Fig. 1left), indicating different KCNQ1 modulation by E1(T) or E2(T).
The currents of KCNQ1 co-expressed with KCNE1, KCNE2 chimeras in HEK293 cells.
Representative current traces of Q1/E1[E2(T)], Q1/E1[E2(C)], Q1/E2[E1(T)] and Q1/E2[E1(C)]. The current traces were elicited by voltage steps ranging from −120 to +60 mV with 20 mV increments for 1.5 s (and 2 s for Q1/E2[E1(C)] due to slow activation of this channel) from a holding potential of −80 mV. The voltage protocols for these chimeras are plotted in the bottom of each trace. The Q1/E2[E1(C)] currents were evoked by a voltage protocol placed at the bottom in which the repolarizing voltage was −160 mV. All the currents were expressed as current density in pA/pF.
The role of the C terminal tail of KCNE1 or KCNE2 on channel modulation was examined with the chimeras E2[E1(C)] and E1[E2(C)] (defined in Fig. S2a). When coexpressed with KCNQ1, E1[E2(C)] fully eliminated the slowly activated currents, whereas E2[E1(C)] showed slowly activated and high amplitude currents (Fig. 1right), indicating that the KCNE1 C-terminus might modulate gating properties of KCNQ1/KCNE channels, consistent with a previous report17.
Role of the KCNE2 N-terminus in KCNQ1 surface expression
More detail electrophysiological studies of the chimera of E1[E2(N)] (defined in Fig. S2a), co-expressing with KCNQ1, were conducted. The observed peak current densities were 75 ± 6.4 pA/pF (KCNQ1 alone, n = 25), 37 ± 2.8 pA/pF (Q1/E2, n = 31) and 31 ± 3.2 pA/pF (Q1/E1[E2(N)], n = 16) for the different channels at +60 mV (Fig. 2a, b). The experimental results showed that the Q1/E1[E2(N)] current decreased to the background level of Q1/E2. This suggested that the KCNE2 N-terminus is responsible for the attenuated KCNQ1 current.
Role of KCNE2 N terminus on KCNQ1 in the HEK293 cell.
(a) Representative current traces of KCNQ1, KCNQ1/KCNE2 and KCNQ1 with chimera E1[E2(N)]. (b) Bars show averaged current density of KCNQ1, KCNQ1/KCNE2 and KCNQ1 with chimera E1[E2(N)]. Error bars represent SEM. ***P < 0.001 (Student's t test). (c) Q1-146Myc (red), EGFP (a marker, green) and merged immunofluorescence images of HEK293 cells. Upper row: Q1; middle row: Q1/E2; lower row: Q1/E1[E2(N)]. The scale bar represents 10 μm. (d) Normalized fluorescence intensities of Q1-146Myc for the transfections of Q1, Q1/E2 and Q1/E1[E2(N)]. Error bars represent SEM. ***P < 0.001 (Student's t test).
To reveal the mechanisms of the attenuated currents of KCNQ1/E1[E2(N)] chimera and the suppressed currents of KCNQ1/KCNE2 complex, surface expression of KCNQ1 in the presence of KCNE2 or chimeras were analyzed using immunofluorescence. The fluorescence intensities were measured as 100 ± 4.3% (n = 194) and 13.8 ± 1.4% (n = 198) for KCNQ1 and KCNQ1/KCNE2, respectively. The chimera E1[E2(N)] also induced the fluorescence intensity reduction (15.6 ± 1.7%, n = 174, Fig. 2c, d). The relative fluorescence intensity ratios were I(Q1/E2)/I(Q1) = 0.14 and I(Q1/E1[E2(N)])/I(Q1) = 0.16 (Fig. 2d). Previously, it was reported that surface expression of KCNQ1 was not affected by KCNE122. Therefore, it is hypothesized that the presence of KCNE2 N-terminus might cause decreased surface expression of KCNQ1 in cell membrane.
Identification of interaction sites within the KCNE-TMDs and KCNQ1
To identify the interaction sites within the KCNE-TMDs and KCNQ1, glutamine-scanning and double mutant cycle analysis were conducted for all the point-mutations. Several amino acids were individually mutated to alanine because of low expression levels.
In accordance with a previous report by Strutz-Seebohm et al. (2011), residues on the KCNE1-TMD (F57-I66) and KCNQ1-S6 (S338-G345) were firstly scanned. The V50 values were 27 mV for Q1/E1 (n = 30), −4 mV for Q1/E1(T58Q) (n = 15), 129 mV for Q1(F340Q)/E1 (n = 16) and 66 mV for Q1(F340Q)/E1(T58Q) (n = 16) (Fig. 3a). The conferred ΔΔV50 = 32 mV (Fig. 3A) strongly indicated that the residue pair of Q1(F340)-E1(T58) were involved in KCNQ1/KCNE1 interactions, which was consistent with previous results20. Similarly, the V50 values of 60 mV for Q1/E1(Y65Q) (n = 13), 23 mV for Q1(A344Q)/E1 (n = 12) and 18 mV for Q1(A344Q)/E1(Y65Q) (n = 15) conferred a ΔΔV50 = 38 mV (Fig. 3b), indicating that the second pair of interaction residues was Q1(A344)-E1(Y65). A summary of the ΔΔV50 values was plotted for mutant combinations in KCNQ1-S6 and KCNE1-TMD (Fig. 3c). Only the two pairs of interaction residues [Q1(F340)-E1(T58), Q1(A344)-E1(Y65)] were distinctive.
Identification of interaction sites between KCNQ1 and KCNE1 using double-mutant cycle analysis.
(a) Left, representative current traces for Q1/E1, Q1/E1(T58Q), Q1(F340Q)/E1 and Q1(F340Q)/E1(T58Q) as indicated. The voltage protocols are plotted at the bottom. Right, the corresponding GV curves from peak currents plotted for Q1/E1 (black), Q1/E1(T58Q) (blue), Q1(F340Q)/E1 (red) and Q1(F340Q)/E1(T58Q) (green) as indicated. (b) Left, representative current traces for Q1/E1, Q1/E1(Y65Q), Q1(A344Q)/E1 and Q1(A344Q)/E1(Y65Q) as indicated. The voltage protocols are plotted at the bottom. Right, the corresponding GV curves plotted for Q1/E1 (black), Q1/E1(Y65Q) (blue), Q1(A344Q)/E1 (red) and Q1(A344Q)/E1(Y65Q) (green) as indicated. (c) Identification of interaction sites for the Q1(S6) mutants (S338 to G345) with the E1(TM) mutants (F57 to I66). The ΔΔV50 values are plotted for the mutant combinations between Q1 and E1 as indicated. Detailed information on the V50 values is provided in Table S1. Two dotted lines are located on 20 and −20 mV. ΔΔV50 values higher than 20 mV or lower than −20 mV are considered as significant changes.
Double mutant cycle analysis was also conducted for KCNQ1 and KCNE2-TMD. The fitted V50 values of −22 mV for Q1/E2 (n = 31), −27 mV for Q1/E2(I64Q) (n = 16), −12 mV for Q1(F340Q)/E2 (n = 20) and 47 mV for Q1(F340Q)/E2(I64Q) (n = 15) conferred a ΔΔV50 = −64 mV (Fig. 4a), indicating an interaction pair of Q1(F340) and E2(I64) in the KCNQ1/KCNE2 complex. At the same time, the fitted V50 values of −27 mV for Q1/E2(I64Q) (n = 16), −82 mV for Q1(F275Q)/E2 (n = 14) and −15 mV for Q1(F275Q)/E2(I64Q) (n = 12) conferred a ΔΔV50 = −72 mV (Fig. 4b), indicating another interaction pair of Q1(F275)-E2(I64). Therefore, the residue E2(I64) could interact synchronously with both Q1(F340) and Q1(F275). The ΔΔV50 values were plotted for mutant combinations of KCNQ1 and KCNE2-TMD (Fig. 4c). Only two pairs of interaction residues [Q1(F275)-E2(I64), Q1(F340)-E2(I64)] within the KCNQ1 and KCNE2-TMD were mapped out using this method.
Identification of interaction sites between KCNQ1 and KCNE2 using double-mutant cycle analysis.
(a) Left, representative current traces for Q1/E2, Q1/E2(I64Q), Q1(F340Q)/E2 and Q1(F340Q)/E2(I64Q) as indicated. The voltage protocols are plotted at the bottom. Right, the corresponding GV curves from peak currents plotted for Q1/E2 (black), Q1/E2(I64Q) (blue), Q1(F340Q)/E2 (red) and Q1(F340Q)/E2(I64Q) (green) as indicated. (b) Left, representative current traces for Q1/E2, Q1/E2(I64Q), Q1(F275Q)/E2 and Q1(F275Q)/E2(I64Q) as indicated. The voltage protocol is plotted at the bottom. Right, the corresponding GV curves from peak currents plotted for Q1/E2 (black), Q1/E2(I64Q) (blue), Q1(F275A)/E2 (red) and Q1(F275A)/E2(I64Q) (green) as indicated. (c) Identification of interaction sites for the Q1(S6) mutants (F332, F335 and S338 to G345), the Q1(S5) mutants (F270 and F275) with the E2(TM) mutants (I63 to V72). The ΔΔV50 values are plotted for the mutant combinations between Q1 and E2 as indicated. Detailed information on the V50 values is provided in Table S1. Two dotted lines are located on 20 and −20 mV. ΔΔV50 values higher than 20 mV or lower than −20 mV are considered as significant changes.
Solution NMR structure determination of KCNE2 in detergent micelles
Differences between the KCNQ1 channel modulations by KCNE1 or KCNE2 can be examined by analyzing their interaction modes, especially the spatial contacts between KCNQ1 and KCNEs. The solution NMR structure of KCNE1 is available18 and in the present study the solution NMR structure of KCNE2 was determined, providing the basis for further analysis of the different modes of interaction between KCNQ1 and the two KCNE proteins at the atomic structure level.
Solution NMR structure analysis of KCNE2 in detergent micelles resulted in the assignment of approximately 90% of the backbone resonances (Cα, CO, HN and N)23. Here, the structure of KCNE2 was determined based on 582 restraints, including 240 dihedral angle restraints, 282 NOE restraints and 60 hydrogen-bond restraints. The statistics of the quality and precision of the final 10 lowest energy structures were summarized in Table S2. The chemical shifts for KCNE2 were deposited in the Biological Magnetic Resonance Bank (BMRB accession number: 18817) and the coordinates were deposited in the Protein Data Bank (PDB accession code: 2M0Q).
The overall structure of KCNE2 was shown in Fig. 5a, containing three α helices: an N-terminal helix (α1, residue 2–42), a transmembrane helix (α2, residues 45–78) and a C-terminal helix (α3, residues 85–118). The structures of three α helices were individually convergent (Fig. 5a and Table S2), but their relative orientations were much dynamic due to the lack of long distance restraints, especially in the presence of globular shape detergent micelles24.
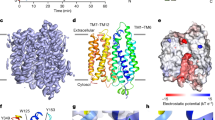
Solution structure of KCNE2 in detergent micelles and comparison of the structures and electrostatic potential surfaces of the KCNE transmembrane helical domains.
(a) Solution structure of KCNE2. Left panel, Schematic illustration of the structure of KCNE2. Middle and right panels, The backbone superposition of the final 10 conformers with the lowest energies in the transmembrane helix (middle panel, residue 45–78), the N-terminal helix (right upper panel, residues 2–42) and C-terminal helix (right lower panel, 85–118) is shown. The N-terminal helix of KCNE2 is an amphiphilic helix. The backbone is shown in blue and the hydrophobic and hydrophilic residues are respectively shown in red and green. (b) Comparison of the structures and electrostatic potential surfaces of the KCNE transmembrane helical domains KCNE2 (left panel) and KCNE1 (right panel). Red denotes negative potential, blue denotes positive potential and white denotes neutral potential. The side-chains of several important residues are indicated.
Docking and interaction models of KCNQ1/KCNE-TMDs
The primary sequences of KCNE1 and KCNE2 were listed in Supplementary Fig. S1 and comparisons of their structures were shown in Fig. 5b. The α2–α3 linker and the juxta-membrane region of α3 were more conserved than the other regions. The transmembrane helix of KCNE2(45–78) was 8 residues longer than that of KCNE1(46–71) (Fig. 5 and Supplementary Fig. S1). A kink in the transmembrane helix of KCNE1 was present in residues around Gly60, whereas it was absent in the corresponding residue Ala65 in KCNE2, resulting in a straight helix (Fig. 5 and Supplementary Fig. S1). The electrostatic potential surfaces of the transmembrane helices of KCNE1 and KCNE2 were similar (Fig. 5), both having highly hydrophobic amino acids (Supplementary Fig. S1).
The solution NMR structures of the KCNE1-TMD and KCNE2-TMD, together with the spatial proximity restraints from the double mutant cycle analysis were used to construct complex models of KCNQ1/KCNE1-TMD and KCNQ1/KCNE2-TMD using PyMol suite (Fig. 6a). The docked complex structure optimizations were processed using 2.4 ns DESMOND suite molecular dynamic simulations for KCNQ1/KCNE1-TMD or KCNQ1/KCNE2-TMD25. The optimized docking results were shown in Figure 6. In Fig. 6aleft, the distances between residue pairs as measured in the double mutant cycle analysis of KCNQ1/KCNE2-TMD were 3.8 Å and 2.6 Å for Q1(F340)-E2(I64) and Q1(F275)-E2(I64), respectively. In Figure 6aright, the distances were 3.4 Å and 2.9 Å for Q1(F340)-E1(T58) and Q1(344)-E1(Y65), respectively. Interestingly, KCNE1-TMD and KCNE2-TMD (Fig. 6b) bound to the S6 helix of KCNQ1 in an apparently different orientation. Two restrained residues between KCNQ1 and KCNE1-TMD [Q1(F340)-E1(T58) or Q1(A344)-E1(Y65)] drove interactions between KCNE1-TMD and the S4–S5 linker in a Chinese chopsticks mode (Fig. 6bright), which might constrain the movement of S6 and result in the slow activation of KCNQ1 channels. On the contrary, KCNE2-TMD only provided one residue site (I64) to interact with two sites in KCNQ1 (F340, F275), which could not restrain the movement of the S6 helix of KCNQ1 (Fig. 6bleft).
Interaction models between the transmembrane domain of KCNEs and the KCNQ1 channel.
(a) The top panel shows the side-view of the Q1/E2 (left) and Q1/E1 (right) complexes. The lower panel shows the top view with the distances between the inter-residues of Q1 to E2 (left) and Q1 to E1 (right). The sticks represent the interacting residues and the dotted lines denote distances in Å. (b) Left, schematic illustration showing that the E2 has no site interacting with the C-terminus of S6 or the S4–S5 linker. Right, schematic illustration showing that the E1 interacts with S6 or the S4–S5 linker to obstruct the channel opening via a spring-like connection.
Role of the KCNE C-terminal tails in KCNQ1 modulation
In the docking models, the KCNE1-TMD and KCNE2-TMD showed different association modes with KCNQ1 (Fig. 6b), which might lead to different orientations of the C-terminal tails of KCNE1 and KCNE2. At the same time, the physiological studies between KCNQ1 and KCNE chimeras (Fig. 1a) indicated that the C-terminus of KCNE1 might modulate gating properties of KCNQ1/KCNE channel.
To verify the possibilities, residue specific flexibilities of KCNE2 proteins were analyzed using the solution NMR backbone amide15N longitudinal relaxation (T1), transverse relaxation (T2) and backbone amide1H-15N inter-nuclear dipolar coupling (NOE) measurements (Fig. 7). Compared with previously reported backbone relaxation data of KCNE119, the C-terminus of KCNE2 was not that flexible. Therefore, high flexibility of the KCNE1 C-terminus provided possibilities for interaction with KCNQ1.
Further examinations of different roles of the flexible KCNE1 C-terminus and the relatively rigid KCNE2 C-terminus in KCNQ1 modulation were conducted. KCNE1 with truncations in the C-terminus after residues 81, 97 and 113 was co-expressed with KCNQ1 to identify the region critical for KCNQ1 current regulation. The acquired channel currents were shown as indicated: Q1/E1(1–81) (Fig. 8aleft), Q1/E1(1–97) (Fig. 8amiddle) and Q1/E1(1–113) (Fig. 8aright). All the constructs resulted in large slowly activated currents, with a significant right-shift in the GV curve (Fig. 8b). The increased GV right-shift with progressive truncation (Fig. 8b) suggested that the entire KCNE1 C-terminus may be important for channel modulation. Channel currents were also analyzed for two KCNE2 truncations (E2(1–75) and E2(1–112)) coexpressed with KCNQ1. Rapidly activated currents were observed similar as those elicited by wild-type KCNE2, with only minor right-shifts in the GV curve (Fig. 8c). Therefore, the more pronounced KCNQ1 kinetic modulations by KCNE1 C-terminal truncations than KCNE2 C-terminal truncations, together with higher flexibilities of KCNE1 C-terminus than KCNE2 C-terminus derived from solution NMR relaxation studies, implied that the rigid KCNE2 C-terminus might lead to less interaction, whereas the flexible KCNE1 C-terminus provided a mechanism for KCNQ1 association and modulation.
Currents and G-V curves for Q1/E1-truncations, Q1/E2 and Q1/E2-truncations.
(a) The currents of Q1/E1(1–81) (left) and Q1/E1(1–97) (middle) were evoked by voltage steps ranging from −80 to 140 mV with 20-mV increments from a holding potential of −80 mV, respectively and the current of Q1/E1(1–113) (right) was evoked by voltage steps ranging from −120 to 100 mV with 20-mV increments from a holding potential of −80 mV. The voltage protocols are plotted at the bottom. (b) The G-V curves were plotted for Q1/E1, Q1/E1(1–81), Q1/E1(1–97) and Q1/E1(1–113) as indicated. (c) The currents of Q1/E2, Q1/E2(1–75) and Q1/E2(1–112) were evoked by voltage steps ranging from −80 to 60 mV with 20-mV increments from a holding potential of −80 mV. The G-V curves were plotted for the three complexes at the bottom.
Discussion
In previous reports, coassembly of KCNQ1 with KCNE2 can form a background potassium channel that is functionally distinct from the slow-activated high amplitude KCNQ1/KCNE1 channel, as observed in Fig. 2a10,26. Further studies demonstrated that association of KCNE2 with KCNQ1/KCNE1 channels could suppress the current amplitude27. The observed current reduction could be attributed to two possible factors: (1) the fewer channel expression on the cell membrane due to impaired trafficking or increased internalization; (2) the reduced single channel conductance of each KCNQ1 in the presence of KCNE2.
Different from the membrane targeting incapability of KCNE1, the KCNE2 alone can be found in cell membrane28. When KCNE2 associated with KCNQ1, the impaired KCNE2 membrane location and partial targeting of KCNE2 to lipid rafts were observed29. Alternatively, the decreased amount of KCNQ1 were detected in lipid rafts in the presence of KCNE2, whereas no obvious changes in the abundance of KCNQ1 in lipid rafts were observed when KCNQ1 associated with KCNE129. Consistent with these reports, KCNQ1 was significantly down-regulated when it co-expressed with KCNE2 (Fig. 2), confirming that association with KCNE2 could bring negative influences on the membrane targeting of KCNQ1. Furthermore, the association of KCNQ1 with chimera of E1[E2(N)] also resulted in decreased channel current and reduced expression level of KCNQ1. These observations strongly indicated that the N-terminal region of KCNE2 might be responsible for the down-regulation of KCNQ1 membrane targeting. Detail mechanism of how the KCNE2 N-terminus affected the KCNQ1 trafficking or channel behaviors should be in more concerns and awaited further studies.
Physiology studies of KCNQ1 associated with chimeras E1[E2(T)] or E2[E1(T)] resulted in much different channel kinetics from the wild type KCNE1 or KCNE2 (Fig. 1), strongly indicating that both KCNE1-TMD and KCNE2-TMD can associate with and modulate KCNQ1.
Several previous reports have demonstrated that the KCNE1-TMD interacts with the S6 segment or S4/S5 linker of KCNQ1. Residues 338–340 on the S6 segment of KCNQ1 and residues 57–59 on the TMD of KCNE1 were revealed as the interaction sites for the gating effect20,21,30. The interaction between the residue pair F340-T5830 and residue T58 on KCNE1 with residues on KCNQ1 S4 domain and S5/S6 segment have been shown previously20. In the present study, the results of double mutant cycle analysis showed that the KCNQ1/KCNE1 channel had two pairs of interaction sites: the previously reported F340-T58 pair and a new A344-Y65 pair that was mapped to the S6 segment of KCNQ1 and the TMD of KCNE1. As in our proposed Chinese chopstick model, the two interaction sites between KCNE1 and KCNQ1 could effectively hinder the movement of the S6 segment, thus affecting channel activity. At the same time, only one residue (I64, located at similar position as T58 of KCNE131) was mapped on KCNE2-TMD to interact with two sites (F340 on S6 and F275 on S5) on KCNQ1, suggesting that KCNE2 was positioned between the S5/S6 segment of KCNQ1. Despite the high similarity between the primary sequences of KCNE1-TMD and KCNE2-TMD (Fig. S1), our docking models indicated that the straight rigid helix of KCNE2-TMD (Fig. 5) and the rigid KCNE2 C-terminus (Fig. 7) determined membrane crossing and interaction with KCNQ1 without physical interaction with cytoplasmic regions of KCNQ1. Consequently, KCNE2 has a lesser effect on KCNQ1 channel gating. On the contrary, the kinked helix in KCNE1-TMD (Fig. 5) and the flexible KCNE1 C-terminus19 result in a tighter interaction with KCNQ1 (two interaction pairs) and direct contact with the cytosolic region of KCNQ1, thus slowing channel activation.
Moreover, the residues 67–78 in KCNE1 C-terminus were mapped to be important for KCNQ1 gating modulation12. Deletion truncations after residue 70 were observed not to alter the activation profile of IKs, but significantly right-shifted the voltage dependence of the channel activation by 60 mV14. Physiology studies of KCNQ1 associated with chimera E2[E1(C)] induced a slow activated KCNQ1 current, while data of KCNQ1 associated with chimera E1[E2(C)] retained the fast activation conductance of KCNQ1. These observations indicating that the C-terminus of KCNE1 can interact directly with the KCNQ1 channel.
Residue specific relaxation analysis of KCNE1 and KCNE2 in detergent micells using solution NMR demonstrated obvious different dynamics of C-terminal tails of the two channel auxiliary proteins: high rigidity in KCNE2 C-terminus versus flexibility in KCNE1 C-terminus. Biophysically, the flexible KCNE1 C-terminus provided possibilities for its interaction with some segments of KCNQ1 and consequent KCNQ1 function modulations. To verify this hypothesis, physiological studies of KCNQ1 associated with KCNEs with different C-terminal truncations were conducted. The activation kinetics of the KCNQ1/KCNE1 channel remained in a slow activated current with all three truncations E1(1–81), E1(1–97) and E1(1–113) (Fig. 8a), whereas the KCNE1 truncations E1(1–81) and E1(1–97) shifted the G-V curves of the activation currents to more positive potentials by approximately 60 mV, similar as the previously reported KCNE1 deletion mutants after residue 7014. By contrast, the G-V curves of the KCNE2 C-terminus truncations E2(1–75) and E2(1–112) showed only a slight right shift.
In a summary, chimeras were generated and Q-scanning methods were used to understand the mechanism of KCNQ1 current regulation by KCNEs. The results of physiological analysis suggested that the KCNE2 N-terminus was responsible for the decreased amplitude of the KCNQ1/KCNE2 channel and the TMD and C terminus of KCNE1 were involved in the regulation of KCNQ1/KCNE1 channel kinetics. Comparisons of solution NMR structures of KCNE1 and KCNE2 showed a straight helix in the KCNE2-TMD, whereas the KCNE1-TMD had a kinked helix. Modeling and docking analysis suggested different association modes of KCNQ1/KCNE2 versus KCNQ1/KCNE1, leading to the differential modulation of KCNQ1 channel gating. Solution NMR backbone relaxation analysis of KCNE2 demonstrated a more rigid C-terminal tail, which was different from previous reported flexible KCNE1 C-terminus. Truncation and physiological studies verified the essential roles of the flexible KCNE1 C-terminus in KCNQ1 modulations. Although different regions of KCNE1 or KCNE2 were mapped for KCNQ1 modulations, the detail mechanisms on how these segments affected KCNQ1 conductance and kinetics require further studies.
Methods
Chimeras and mutations
Constructs of the wild-type human KCNQ1 subunit or KCNQ1-146Myc were prepared as described previously22. The coding sequences of human kcne1 and kcne2 (GenBank accession NOs NG_009091 and NG_008804) were sub-cloned into pcDNA3.1(+) (Invitrogen, CA) using the appropriate restriction enzymes. All chimeras were constructed using overlap-extension PCR with Pfu polymerase (Fermentas). Mutations were introduced using the Site-directed Gene Mutagenesis Kit (Beyotime). All chimeras and point mutations were verified through direct DNA sequencing.
Cell culture and transient transfections in HEK293 cells
HEK293 cells were cultured in DMEM supplemented with 10% fetal bovine serum (FBS), 100 U/ml penicillin and streptomycin in 37°C and 5% CO2 incubators. One day before transfection, cells were transferred to 24-well plates and when reaching 90% confluence, transfections were performed with Lipofectamine 2000 (Invitrogen, CA). At 4–6 h after transfection, cells were transferred to poly-D-lysine (Sigma, MO) coated slides for patch-clamp recording or imaging. For all co-transfection experiments, the ratio of KCNQ1 coding plasmids to KCNE1 (or KCNE2) coding plasmids was 1:1.5.
Immunofluorescence imaging in HEK293 cells
Approximately 24 h after transfection, cells were fixed with 2% paraformaldehyde in Phosphate Buffered Saline (PBS) for 10 min, washed with PBS (5 min × 3) and blocked with 5% FBS for 1 h. Then, cells were incubated with mouse monoclonal anti-human Myc antibody (1:300) (Abcam) overnight at 4°C, washed with 2.5% FBS in PBS (5 min × 6) and incubated with Rhodamine-conjugated goat anti-mouse IgG (H + L) (1:300) (Proteintech Group) for 2 h at room temperature (25°C). Cells were then washed (5 min × 6) and soaked in PBS, then were visualized using a confocal laser scanning microscopy (Olympus IX71) and a 100× oil immersion lens (NA1.30). Parameter selection, sample scanning and image acquisition were all controlled by the Andor IQ 2.0 software (Andor Technology plc, Springvale Business Park, United Kingdom). 16-bit digital images were obtained with an EM CCD camera (DU897K, ANDOR iXon, United Kingdom). The quantification of the average fluorescence intensity of KCNQ1-Myc was performed in ImageJ 1.45 (Wayne Rasband, National Institutes of Health). Following background subtraction, a 4-pixel-wide region along plasma membrane in the middle layer was drawn for the quantification. Adjacent membranes from two transfected cells were rejected from the analysis. Fields for imaging and analysis were selected using EGFP fluorescence to identify transfected cells. All experiments were performed at least three times. For each group, approximately 60 cells were examined and normalized to the intensity of the control group. Representative images were deconvolved with AutoQuant X2 software (Media Cybernetics Inc., MD) and arranged by Adobe Photoshop CS5.
Solutions
For the whole-cell recordings, the pipette solution contained the following (in mM): 5 Ethylene glycol-bis(2-aminoethylether)-N,N,N′,N′- tetraacetic acid (EGTA), 140 KCl, 10 NaCl, 1 MgCl2, 10 HEPES (pH 7.0) titrated with KOH. Bathing solution contained the following (in mM): 150 NaCl, 4 KCl, 2 CaCl2, 1 MgCl2, 10 HEPES (pH 7.0) titrated with NaOH. All chemicals were purchased from Sigma.
Patch clamp recording
Transfected HEK293 cells were bathed in a solution containing the following (in mM): 150 NaCl, 4 KCl, 2 CaCl2, 1 MgCl2, 10 HEPES one day after transfection. All the experiments were carried out in whole-cell patches. Patch pipettes were pulled from borosilicate glass capillaries with a resistance of 2–3 megohms and filled with the pipette solution. Experiments were performed using an Axon 200B patch-clamp amplifier with its software (Axon Instruments, CA, USA). Currents were typically digitized at 10 kHz and filtered at 5 kHz. All experiments were performed at room temperature (22–24°C).
Data analysis
Recording data were analyzed with Clampfit (Axon Instruments) and Sigmaplot (SPSS) software. Unless stated otherwise, the data are presented as mean ± SE and significance was tested by Student's T test. Differences between the mean values were considered significant at a probability ≤0.05. The G-V curves for activation were fitted by the Boltzmann equation: G/Gmax = (1 + exp((V − V50)/κ))−1, where V50 is the voltage at which the conductance (G) is half the maximum conductance (Gmax) and κ is a factor affecting the steepness of the activation.
For double mutant cycle analysis32,33,34,35, the principle is that pairwise coupling free energy changes associated with mutations in pairs of residues located in different subunits can be calculated using a thermodynamic square comprised of the wild-type (WT) complex (αβ), the two single mutants (α*β and αβ*) and the corresponding double mutant (α*β*). Here, α, β and * denote the KCNQ1, KCNE and mutants, respectively. The thermodynamic square was described as ΔΔG = ΔGαβ* + ΔGα*β − ΔGα*β*. In this cycle, if the total free energy change ΔΔG is significant (≥1 kcal/mol), the pairwise residues on α and β subunits can be considered as coupled35. Since the free energy of a channel is directly proportional to the potential at which the open probability is 50% (V50); therefore, the V50 was measured and used for double mutant cycle analysis as follows: ΔΔV50 = ΔV50-αβ* + ΔV50-α*β − ΔV50-α*β* − ΔV50-αβ. In this method, significant changes of ΔΔV50 ≥ 20 mV (or 1 kcal/mol) can be used as the criteria to examine whether the two sites in KCNQ1 or KCNEs are involved in the direct interaction.
Structure determination of KCNE2 using solution NMR
The human KCNE2 protein was over-expressed using an E. coli bacterial expression system and purified in LMPG micelles. A series of TROSY-based multi-dimensional NMR experiments were recorded on a 600 MHz Bruker spectrometer. The 2D1H,15N-HSQC, 3D HNCO, HNCOCA, HNCA, HNCACB and HN(CO)CACB spectra were collected using a2H/13C/15N labeled KCNE2 sample in LMPG micelles at pH 6.0 and 40°C. The 3D HBHA(CO)NH and 3D15N-edited NOESY (mixing time: 100 ms) spectra were collected using a13C/15N labeled KCNE2 sample. Details of the experimental parameters were described previously23. All of the NMR spectra were processed using NMRPipe36 and analyzed using NMRView37. Backbone resonance assignments were achieved23 and deposited in the BioMagResBank (www.bmrb.wisc.edu) with accession number 18817. Backbone structural restraints of KCNE2 in detergent micelles were collected in the form of the backbone resonance chemical shift and short-range Nuclear Overhauser Effects (NOE). After chemical shift index and protein backbone dihedral angle analysis using TALOS+38, XPLOR-NIH39 was applied to calculate the three-dimensional structure of KCNE2 in detergent micelles. In the 10 lowest energy structures of the 100 calculated structures, the backbone structures of the KCNE2 transmembrane domain [residues 46–78] were well converged to an RMSD value of 0.909 Å, although the backbone structures of the full length KCNE2 were not converged because of lack of long distance restraints. The statistics of the determined KCNE2 structure are provided in Table S1. Coordinates of the KCNE2 ensemble were deposited in the Protein Data Bank (PDB accession code: 2M0Q).
Molecular modeling and MD simulations
The KCNQ1 homology model was generated using MODELLER40. Transmembrane domains (TMD) of KCNE1 or KCNE2 obtained from NMR studies were optimized in a POPC lipid bilayer membrane using DESMOND25. Docking of the optimized KCNE1-TMD and KCNE2-TMD with the adjacent-KCNQ1 subunits homology model was performed using Rosetta41. The KCNQ1/KCNE1-TMD and KCNQ1/KCNE2-TMD were inserted into two different explicit POPC lipid bilayer membranes (300 K) and then the whole system was neutralized by addition of Cl− ions. Molecular dynamics simulations were performed with DESMOND based on a standard Dynamics Cascade pipeline for the simulation of macromolecular systems. The detailed structural conformations of KCNQ1/KCNE1-TMD and KCNQ1/KCNE2-TMD were prepared using the latest PyMOL Suite (Version 1.5.0.4 Schrödinger, LLC).
References
Goldman, A. M. et al. Arrhythmia in heart and brain: KCNQ1 mutations link epilepsy and sudden unexplained death. Sci. Transl. Med. 1, 2ra6 (2009).
Tsevi, I. et al. KCNQ1/KCNE1 channels during germ-cell differentiation in the rat: expression associated with testis pathologies. J. Cell. Physiol. 202, 400–410 (2005).
Viswanathan, P. C., Shaw, R. M. & Rudy, Y. Effects of IKr and IKs heterogeneity on action potential duration and its rate dependence: a simulation study. Circulation 99, 2466–2474 (1999).
Silva, J. & Rudy, Y. Subunit interaction determines IKs participation in cardiac repolarization and repolarization reserve. Circulation 112, 1384–1391 (2005).
Barhanin, J. et al. K(V)LQT1 and lsK (minK) proteins associate to form the I(Ks) cardiac potassium current. Nature 384, 78–80 (1996).
Sanguinetti, M. C. et al. Coassembly of K(V)LQT1 and minK (IsK) proteins to form cardiac I(Ks) potassium channel. Nature 384, 80–83 (1996).
Heitzmann, D. et al. Heteromeric KCNE2/KCNQ1 potassium channels in the luminal membrane of gastric parietal cells. J. Physiol. 561, 547–557 (2004).
Lambrecht, N. W., Yakubov, I., Scott, D. & Sachs, G. Identification of the K efflux channel coupled to the gastric H-K-ATPase during acid secretion. Physiol. Genomics 21, 81–91 (2005).
Roepke, T. K. et al. The KCNE2 potassium channel ancillary subunit is essential for gastric acid secretion. J. Biol. Chem. 281, 23740–23747 (2006).
Tinel, N., Diochot, S., Borsotto, M., Lazdunski, M. & Barhanin, J. KCNE2 confers background current characteristics to the cardiac KCNQ1 potassium channel. EMBO J. 19, 6326–6330 (2000).
Takumi, T. et al. Alteration of channel activities and gating by mutations of slow ISK potassium channel. J. Biol. Chem. 266, 22192–22198 (1991).
Tapper, A. R. & George, A. L., Jr MinK subdomains that mediate modulation of and association with KvLQT1. J. Gen. Physiol. 116, 379–390 (2000).
Melman, Y. F., Domenech, A., de la Luna, S. & McDonald, T. V. Structural determinants of KvLQT1 control by the KCNE family of proteins. J. Biol. Chem. 276, 6439–6444 (2001).
Chen, J., Zheng, R., Melman, Y. F. & McDonald, T. V. Functional interactions between KCNE1 C-terminus and the KCNQ1 channel. PLoS ONE 4, e5143 (2009).
Romey, G. et al. Molecular mechanism and functional significance of the MinK control of the KvLQT1 channel activity. J. Biol. Chem. 272, 16713–16716 (1997).
Haitin, Y. et al. Intracellular domains interactions and gated motions of I(KS) potassium channel subunits. EMBO J. 28, 1994–2005 (2009).
Zheng, R. et al. Analysis of the interactions between the C-terminal cytoplasmic domains of KCNQ1 and KCNE1 channel subunits. Biochem. J. 428, 75–84 (2010).
Kang, C. et al. Structure of KCNE1 and implications for how it modulates the KCNQ1 potassium channel. Biochemistry 47, 7999–8006 (2008).
Tian, C. et al. Preparation, functional characterization and NMR studies of human KCNE1, a voltage-gated potassium channel accessory subunit associated with deafness and long QT syndrome. Biochemistry 46, 11459–11472 (2007).
Strutz-Seebohm, N. et al. Structural basis of slow activation gating in the cardiac I Ks channel complex. Cell. Physiol. Biochem. 27, 443–452 (2011).
Melman, Y. F., Um, S. Y., Krumerman, A., Kagan, A. & McDonald, T. V. KCNE1 binds to the KCNQ1 pore to regulate potassium channel activity. Neuron 42, 927–937 (2004).
Pan, N., Sun, J., Lv, C., Li, H. & Ding, J. A hydrophobicity-dependent motif responsible for surface expression of cardiac potassium channel. Cell. Signal. 21, 349–355 (2009).
Chen, L., Lai, C., Lai, J. & Tian, C. Expression, purification, detergent screening and solution NMR backbone assignment of the human potassium channel accessory subunit MiRP1. Protein Expr. Purif. 76, 205–210 (2011).
Chou, J. J., Kaufman, J. D., Stahl, S. J., Wingfield, P. T. & Bax, A. Micelle-induced curvature in a water-insoluble HIV-1 Env peptide revealed by NMR dipolar coupling measurement in stretched polyacrylamide gel. J. Am. Chem. Soc. 124, 2450–2451 (2002).
Bowers, K. J. et al. in Proceedings of the 2006 ACM/IEEE conference on Supercomputing 84 (ACM, Tampa, Florida, 2006).
Yang, Y. et al. Identification of a KCNE2 gain-of-function mutation in patients with familial atrial fibrillation. Am. J. Hum. Genet. 75, 899–905 (2004).
Jiang, M. et al. Dynamic partnership between KCNQ1 and KCNE1 and influence on cardiac IKs current amplitude by KCNE2. J. Biol. Chem. 284, 16452–16462 (2009).
Kanda, V. A., Lewis, A., Xu, X. & Abbott, G. W. KCNE1 and KCNE2 inhibit forward trafficking of homomeric N-type voltage-gated potassium channels. Biophys. J. 101, 1354–1363 (2011).
Roura-Ferrer, M. et al. Impact of KCNE subunits on KCNQ1 (Kv7.1) channel membrane surface targeting. J. Cell. Physiol. 225, 692–700 (2010).
Panaghie, G., Tai, K. K. & Abbott, G. W. Interaction of KCNE subunits with the KCNQ1 K+ channel pore. J. Physiol. 570, 455–467 (2006).
Liu, X. S., Zhang, M., Jiang, M., Wu, D. M. & Tseng, G. N. Probing the interaction between KCNE2 and KCNQ1 in their transmembrane regions. J. Membr. Biol. 216, 117–127 (2007).
Carter, P. J., Winter, G., Wilkinson, A. J. & Fersht, A. R. The use of double mutants to detect structural changes in the active site of the tyrosyl-tRNA synthetase (Bacillus stearothermophilus). Cell 38, 835–840 (1984).
Sadovsky, E. & Yifrach, O. Principles underlying energetic coupling along an allosteric communication trajectory of a voltage-activated K+ channel. Proc. Natl. Acad. Sci. U. S. A. 104, 19813–19818 (2007).
Serrano, L., Horovitz, A., Avron, B., Bycroft, M. & Fersht, A. R. Estimating the contribution of engineered surface electrostatic interactions to protein stability by using double-mutant cycles. Biochemistry 29, 9343–9352 (1990).
Yifrach, O. & MacKinnon, R. Energetics of pore opening in a voltage-gated K(+) channel. Cell 111, 231–239 (2002).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B. A. Using NMRView to visualize and analyze the NMR spectra of macromolecules. Methods Mol. Biol. 278, 313–352 (2004).
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J. Biomol. NMR 44, 213–223 (2009).
Schwieters, C. D., Kuszewski, J. J., Tjandra, N. & Marius Clore, G. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Das, R. et al. Simultaneous prediction of protein folding and docking at high resolution. Proc. Natl. Acad. Sci. U. S. A. 106, 18978–18983 (2009).
Acknowledgements
This study was supported by the Chinese Key Research Plan (No. 2011CB911104), Chinese National High-Tech Research Grant (No. 2006AA02A321) and Chinese Natural Science Foundation (30870489) to C.L. Tian; and Chinese Natural Science Foundation (31100847) and Fundamental Research Funds for the Central Universities of China (WK2070000033) to L.H. Zhang; and Chinese Natural Science Foundation (30971179, 31170814) and Chinese Key Research Plan (2010CB529804) to J.P. Ding.
Author information
Authors and Affiliations
Contributions
L.Z., C.T. and J.D. wrote the manuscript. P.L., H.L. and P.S. carried out the mutation construction and electrophysiology experiments. W.Z. carried out the immonofluorescence experiment. C.L. carried out the NMR experiments and F.W. prepared figure 5. S.W. prepared figure 6. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
supplementary file
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Li, P., Liu, H., Lai, C. et al. Differential Modulations of KCNQ1 by Auxiliary Proteins KCNE1 and KCNE2. Sci Rep 4, 4973 (2014). https://doi.org/10.1038/srep04973
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04973
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.