Abstract
Computational microscopy tools, in particular lensfree on-chip imaging, provide a large field-of-view along with a long depth-of-field, which makes it feasible to rapidly analyze large volumes of specimen using a compact and light-weight on-chip imaging architecture. To bring molecular specificity to this high-throughput platform, here we demonstrate the use of plasmon-resonant metallic nanoparticles to automatically recognize different cell types based on their plasmon-enhanced lensfree holograms, detected and reconstructed over a large field-of-view of e.g., ~24 mm2.
Similar content being viewed by others
Introduction
Computational imaging without lenses has been recently emerging as a new biomedical imaging technique1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21. Such lensfree imaging modalities record transmitted, scattered, or emitted photons from objects that are placed directly on or only <2–3 millimeters away from the active area of a sensor-array such as a CMOS (Complementary Metal-Oxide-Semiconductor) or a CCD (Charge-Coupled Device) chip. Due to this “on-chip” imaging architecture, the imaging field-of-view (FOV) equals the active area of each sensor-array and therefore it can routinely reach e.g., 20–30 mm2 using a standard CMOS imager or ~10–20 cm2 using a large format CCD chip. In its holographic implementation1,6,7,8,9,19, in addition to FOV, the depth-of-field (DOF) is also significantly enhanced (e.g., ~1–5 mm), which permits rapid imaging of large specimen volumes of for example >20–2,000 μL on a chip. This throughput advantage, when combined with the compactness and cost-effectiveness of its imaging architecture, which can also be integrated with microfluidic channels7,8,9, or wireless devices such as cell phones10, makes lensfree holographic on-chip imaging a promising modality especially for biomedical imaging and diagnostic needs in resource-limited settings as well as for telemedicine applications.
Unlike its coherent alternatives, some of the recent lensfree holography work focused on using partially coherent quasi-monochromatic sources that emanate from rather large apertures (e.g., 50–100 μm). This partially coherent illumination beam scatters from the body of each specimen, for example a cell and then interferes with the unscattered background light to form a lensfree in-line hologram of the object to be sampled at the sensor array. These captured lensfree holograms can then be rapidly processed to reconstruct both phase and amplitude images of the specimen. Still at a relatively early stage of development, lensfree holographic on-chip imaging has already shown significant potential in a broad range of biomedical applications, such as three dimensional (3D) tomographic imaging of model organisms11,12, tracking of micro-swimmers13, point-of-care (POC) imaging10,14, automated cell counting15,16,17 among others18,19,20,21.
Some of the remaining challenges for computational on-chip imaging techniques include spatial resolution, image contrast and molecular specificity1. Recently, a lateral resolution of ~300 nm, corresponding to a numerical aperture (NA) of 0.9, has been achieved over a large FOV of >20 mm2 in lensfree holography by capturing multiple sub-pixel shifted holograms1 that are digitally merged using a pixel super-resolution algorithm6. On the other hand, compared to these recent improvements in resolution, molecular- or cellular-specific contrast has not yet been demonstrated in lensfree imaging. Differentiation of cell sub-types is particularly important for medical diagnostics and monitoring of various diseases. For instance, the ratio of CD4+ T lymphocytes to CD8+ T lymphocytes is a crucial diagnostic marker of HIV progression and can be used to determine the need for antiretroviral therapy. However, recognition of different cell sub-types with similar or identical physical morphologies but different biological functions, e.g., expressing different surface receptors, still remains a major challenge for computational on-chip imaging techniques in general, limiting their applications in cytometry.
To provide a solution to this important need, here we demonstrate that antibody-conjugated plasmonic nanoparticles (NPs) can serve as efficient contrast agents for lensfree holograms by modulating the optical absorption and scattering properties of targeted cells. In this approach, NPs are used to encode the holographic images of the labeled cells with unique plasmon-resonance based spectral signatures that can be resolved by multispectral lensfree holographic imaging and automatically recognized by machine-learning algorithms to enable classification of cell types labeled with different NPs over a large FOV of e.g., ~24 mm2. We should note that plasmonic NPs are also widely used as contrast agents in various conventional microscopic imaging modalities22,23,24,25,26,27,28,29,30, including lens-based holographic techniques31,32,33,34. However, these previous approaches, which are all based on lenses or microscope objectives, have rather limited FOVs (e.g., <0.5 mm2) and DOFs (e.g., <0.1 mm), which significantly limit their throughput for imaging cytometry applications.
Our plasmonic lensfree holographic cytometer is also significantly different from previous on-chip cytometry approaches, which are typically miniaturized versions of conventional flow cytometers35,36,37,38,39. First, our plasmonic lensfree cytometer does not rely on a microfluidic flow system and thus it does not demand sophisticated chip fabrication or fluid handling. Second, it serves as a dual-function platform that simultaneously allows cytometric counting and single-cell imaging with sub-micron spatial resolution over a very large field-of-view, which is typically not available using other on-chip cytometry devices. Third, it allows point-to-point verification of the differentiation or counting results and permits re-interrogation of any particular cell of interest over the sample field-of-view, whereas in flow-based on-chip cytometry, once a cell has been measured, a single cell level sorting platform would be needed for separation or isolation of individual cells. These unique features make our plasmonic lensfree cytometry approach a promising platform for multi-parameter single-cell analysis on a chip.
Results
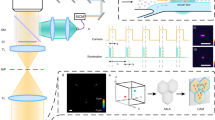
To demonstrate the proof of concept of our high-throughput plasmonic NP based imaging cytometry approach over a large FOV of e.g., ~24 mm2, we first show the discrimination of gold nanoparticle (Au NP) labeled CD4+ T cells from unlabeled ones, which will be denoted as Au-CD4 and CD4, respectively. Our lensfree in-line holographic imaging set-up is illustrated in Figure 1a, where a monochromator is used to provide partially coherent quasi-monochromatic illumination through a multi-mode optical fiber with a core diameter of 100 μm. The end of the multi-mode fiber is mounted ~10 cm (z1) away from the sample plane and is shifted stepwise in a 4 × 4 array with a step size of ~0.85 mm (blue dashed line) to enable deeply sub-micron resolution using pixel super-resolution6. Samples to be imaged are held by two glass cover slips (each ~0.1 mm thick, separated by 30-μm-diameter polystyrene beads and sealed) and are directly placed on a CMOS sensor-array that has a pixel size of 2.2 μm and an active area of ~24 mm2. The vertical distance between the sample plane and the active area of the CMOS sensor chip is ~650–700 μm (z2). This on-chip imaging geometry has a FOV of ~24 mm2 (see e.g., Figure 1b and Supplementary Figure S1), which is around two-orders of magnitude wider than the typical FOV of e.g., a 40× objective lens that has a similar resolution. This large FOV conveniently allows screening of several thousand cells within a single frame. Each specimen is imaged sequentially at seven different illumination wavelengths (λ) ranging from 480 nm to 950 nm, each with a bandwidth of ~10 nm. 100-nm diameter Au NPs (plasmon-resonance peak wavelength: ~554 nm, see Figure S2a) are conjugated with antiCD4 antibodies (denoted as Au-antiCD4) and are used to specifically label CD4 cells (see Methods)40. In our initial experiments, CD4 cells with and without Au NP labeling exhibited strong contrast differences under a conventional darkfield scattering microscope (see Figures 1c,f). For the same CD4 cells, pixel super-resolved (SR) lensfree holograms and their reconstructed amplitude images (λ = 560 nm) are also shown in Figures 1d,g and Figures 1e,h, respectively. Based on these single-wavelength lensfree reconstruction results, it is difficult to visually resolve NP-labeled cells from unlabeled ones due to their similar 2D texture and contrast.
Au NPs as contrast agents for lensfree on-chip imaging.
(a) Schematic illustration of a multispectral lensfree in-line holographic system. (b) Full FOV of a lensfree hologram of Au-CD4 cells excited at 560 nm; the center region of the white box is shown in (g). (c,f) Darkfield scattering microscopy of CD4 and Au-CD4 cells, respectively, obtained by a 60× objective lens. (d,g) Lensfree super-resolved (SR) holograms (λ ~ 560 nm) and (e,h) reconstructed amplitude images of the same regions of interest (ROIs) as the darkfield scattering images (c,f). Scale bar = 25 μm for c, d, e; Scale bar = 10 μm for f, g, h.
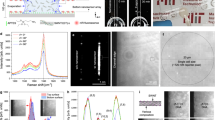
To circumvent this limitation, we implemented a multispectral lensfree imaging approach, which is also used in lens-based microscopy techniques for enhancing spectral recognition and characterization of nanostructures41,42,43. Figure 2a contains a series of SR lensfree holograms and their reconstructed amplitude images corresponding to a single Au-CD4 cell imaged by seven different wavelengths in the visible and near infrared parts of the optical spectrum. In our reconstructed holographic images, the cellular absorbance of each cell (Acell) can be defined as:

where Ibk and Icell denote the mean intensity of the surrounding background and the cell, respectively (Figure 2a). Representative absorption spectra of labeled and unlabeled CD4 cells are also shown in Figure 2b, where the unlabeled CD4 cells exhibit a decayed absorption pattern with increasing wavelength, whereas Au-labeled CD4 cells exhibit a major absorbance peak within the spectral range 560–750 nm, which appears to be broadened and red-shifted compared to the plasmon-resonant wavelength of isolated 100-nm Au NPs (λmax ~ 554 nm). A similar behavior is also observed in our Finite-Difference-Time-Domain (FDTD) simulations, which will be discussed later on. The spectral properties of these two cell types measured by lensfree on-chip holography also agree well with conventional UV-vis spectroscopy measurements, where a similar wavelength-dependent contrast behavior can be observed (see Supplementary Figure S2b and c). Conventional darkfield scattering microscopy images were also employed on the same cells to obtain the nanoparticle distribution per cell and to compare the results with the optical features measured by multispectral lensfree holographic microscopy (see Figures 1c–h and Supplementary Figures S1d–e and S3).
Differentiation of CD4 and Au-CD4 cells by multispectral lensfree on-chip holography.
(a) Lensfree SR holograms and the corresponding reconstructed amplitude images of an individual Au-CD4 cell imaged with seven different wavelengths (480–950 nm). Scale bars are 10 μm. Cellular absorbance is calculated by the logarithm of the ratio of the background intensity (Ibk) to the averaged cell intensity (Icell). (b) Representative absorption spectra of CD4 (left, n = 15) and Au-CD4 cells (right, n = 15) measured by lensfree on-chip holography. (c) PC1–PC2 plot of 624 CD4 and 508 Au-CD4 cells. (d) Classification of CD4 and Au-CD4 cells by SVM based on a seven-dimensional vector (PC1-PC7). The boundary (defined by the black curve) shows the maximum separation of unlabeled and labeled CD4 cells. Green: CD4 cells used as training; Cyan: CD4 cells classified; Red: Au-CD4 cells used as training; Magenta: Au-CD4 cells classified. 50% of the total cell population was used for training and the rest for testing. An average accuracy of 93.7 ± 0.5% was obtained over 10 independent SVM runs based on PC1–PC7.
The above outlined multispectral lensfree on-chip imaging approach was performed on >600 unlabeled and >500 Au NP labeled CD4 cells. The reconstructed images were first characterized through principal component analysis (PCA)44 and then trained by support vector machine (SVM)45. PCA is a mathematical procedure that transforms the original set of data into new coordinates called principal components (PCs) such that the largest variance of the original data lies on the first coordinate (first principal component, or PC1), the second largest variance on the second coordinate (second principal component, or PC2, which is orthogonal to PC1) and so on. PCA can significantly reduce the dimensionality of the raw data, while also extracting and presenting the most important features of the original data in the new PC space. Complementing PCA, SVM serves as a machine-learning tool that analyzes the characteristics of variables in PC space to generate a decision boundary (or support vectors) to automatically classify unknown data sets.
Figure 2c shows a scatter plot of more than 1,100 cells (624 unlabeled and 508 NP labeled) in the PC1–PC2 space, as a result of analyzing the spectral features of the lensfree reconstructed images of these cells with PCA. SVM analysis using a seven-dimensional vector (PC1–PC7) and 50% of the total cell population as the training set generated a decision boundary that maximized the separation of NP labeled and unlabeled cells (Figure 2d). For testing its classification accuracy, the SVM analysis (based on PC1–PC7) was repeated 10 times on randomly selected training data but with a fixed training/testing ratio (50%–50%). Overall, an average accuracy of 93.7% was achieved (with a standard deviation of ~0.5%) for separating NP-labeled CD4 cells from unlabeled ones. The narrow distribution of our accuracy values over multiple independent SVM runs also suggests that the decision boundary stays quite accurate regardless of the composition of the training and test sets.
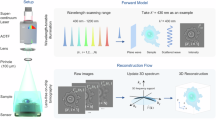
After these initial results involving a single NP type (Figure 2), next we tested the concept of nanoparticle-enhanced lensfree cytometry by using two different types of plasmon-resonant NPs (Au and Ag) for multiplexed recognition of CD4 and CD8 lymphocytes, which might especially be useful for monitoring of HIV+ patients and initiation of antiretroviral therapy46,47. For this purpose, 100-nm silver nanoparticles (Ag NPs, λmax ~ 480 nm) were conjugated with antiCD8 antibodies using a similar strategy used in preparation of Au-antiCD4 conjugates (see Methods). The success of labeling CD4 and CD8 cells with Au and Ag NPs, respectively, was verified by darkfield scattering microscopy images as before, where the cells scattered strongly at the plasmon-resonant wavelengths of labeled NPs (Figures 3c,g). In addition, labeled CD4 and CD8 cell pellets collected by slow-speed centrifugation (270 g, 5 min) showed purple (Au labeled) or yellow (Ag labeled) color in all the pellets, indicative of NP labeling (see e.g., Figures 3d,h).
Multiplexed lensfree holographic detection of CD4 and CD8 cells using Au and Ag NPs.
Lensfree holography of (a,b) CD4 and (e,f) CD8 cells. (a,e) SR holograms and (b,f) reconstructed images of CD4 and CD8 cells. (c,g) Darkfield scattering images and (d, h) photographs of Au-CD4 and Ag-CD8 cells, respectively. Cell pellets were formed in PBS buffer by centrifugation (270 g, 5 min). (i) PC1–PC2 plot of CD4, CD8, Au-CD4 and Ag-CD8 cells by PCA. (j) Multi-class SVM classification results of four types of cells generated by LIBSVM. Scale bar = 50 μm for (a), (b), (e) and (f); Scale bar = 10 μm for (c) and (g).
Next we applied our multispectral lensfree on-chip imaging cytometry approach to Ag NP-labeled CD8 cells (denoted as Ag-CD8). Compared to Au-CD4 cells, we observed a blue-shifted absorption peak at around 450–550 nm (see Supplementary Figure S4). For classification, PCA analysis was performed on multi-class data, including Au-CD4, Ag-CD8, unlabeled CD4 and unlabeled CD8 cells. Figure 3i contains a PCA scatter plot of ~2,200 cells in the PC1–PC2 domain. It shows an overlapped distribution of native CD4 (green) and CD8 cells (yellow), which is expected since the reconstructed images of unlabeled CD4 and CD8 cells look very similar to each other as illustrated in Figures 3b,f. On the contrary, Au-CD4 (red) and Ag-CD8 cells (blue) can be easily separated in the PC1–PC2 space. We then performed SVM analysis on these multi-class species using LIBSVM (which stands for a library for support vector machines, an open-source program for support vector classification)48. The results are summarized in Figure 3j. In agreement with PCA (Figure 3i), native CD4 and CD8 cells without any NP labeling were classified as the same group by multi-class SVM. Stated differently, native CD4 and CD8 cells before NP labeling only showed a classification accuracy of ~59.4 ± 0.9% (Supplementary Figure S5a), indicating a slightly better performance than flipping a coin. On the other hand, after labeling with Au and Ag NPs, our CD4–CD8 classification accuracy increased to 95.4 ± 0.8% as illustrated in Supplementary Figure S5b. These results demonstrate that multi-spectral lensfree holographic on-chip cytometry is capable of classifying multiple species using plasmonic NPs as cell-specific contrast agents.
Discussion
The mechanism of enhanced recognition of morphology-similar cells in NP-assisted lensfree holography is attributed to the modulation of optical absorption and scattering profiles of the labeled cells in the presence of plasmon-resonant NPs. Metal NPs are very well-suited contrast agents for lensfree on-chip microscopy due to their extremely high extinction coefficients (108–1010 M−1 cm−1) which are typically several orders of magnitude larger than those of organic dyes49,50,51. Moreover, noble metal NPs are considered to be chemically inert and biocompatible and have been effectively used as diagnostic imaging or therapeutic agents both in vitro and in vivo52,53.
In our NP-assisted on-chip cytometry experiments, although we achieved highly accurate CD4-CD8 characterization, reaching an accuracy level of 95.4 ± 0.8% (e.g., Figure 3 and Supplementary Fig. S5b), a significant variation among spectral features of each cell type was observed, which is also evident in the scatter plot shown in e.g., Figure 3i. To shed more light on this variation and its possible sources, we characterized the distribution of the spectral responses of each cell type by using a parameter, which we term as the Center of Mass (CoM) of the spectra (Figure 4a, inset). Based on this, we calculated the weighted center of each cellular absorption spectrum measured by our lensfree on-chip microscope as a function of either absorbance or wavelength (denoted as absorbance CoM and wavelength CoM, respectively; see Methods). With this definition, the distribution of CoM for Au NP-labeled CD4 cells is depicted in Figure 4a where each data point represents a single cell's lensfree spectral measurement. It shows that more than 98% of the cells have their wavelength CoM located within 640–740 nm range when the cells were incubated with 3.6 × 1010 Au NPs per milliliter (NP/mL). We also repeated the same lensfree Au-CD4 imaging experiments with four different NP concentrations ([Au NP]) ranging from 9.6 × 109 to 1.44 × 1011 NP/mL, while other conditions of cell incubation were kept the same. The results of these experiments on the dependence of wavelength CoM on Au NP dosage are summarized in Figure 4b, which reveals that the wavelength CoM of the labeled cells shifts to longer wavelengths at higher Au NP concentrations. It is also worth noting that this dependence of CoM wavelength on [Au NP] is not linear, i.e., the red shift of CoM wavelength becomes much smaller when [Au NP] exceeds 3.6 × 1010 NP/mL, which can be attributed to saturation of Au NP labeling on the surface of CD4 cells as well as formation of 3D clusters on the cell surface or within the cell body due to partial endocytosis of NPs.
The effect of Au NP concentration ([Au NP]) on the absorption of CD4 cells.
(a) CoM absorbance and CoM wavelength of >350 Au-CD4 cells measured by multispectral lensfree holography ([Au NP] = 3.6 × 1010 NP/mL). A schematic illustration of the CoM of the spectrum is shown in the inset, where x = CoM Wavelength and y = CoM Absorbance. (b) Distribution of CoM wavelength when incubated with different concentrations of Au NPs (from 9.60 × 109 to 1.44 × 1011 NP/mL). The width of the distribution represents the relative population of cells at that CoM wavelength. Number n on the top of each distribution represents the number of cells that each group contains. The box chart on the right side shows the statistical percentiles of each distribution (bars: the maximum and minimum; whiskers: top 5% and 95%, respectively; the box: range of 25–75%; middle line: 50%; middle dot: mean of the distribution).
To shed more light on these NP concentration experiments summarized in Figure 4, we modeled the spectral properties of random 2D distributions of 100-nm Au NPs using FDTD simulations (see Methods). Due to the small size of Au NPs compared to CD4 or CD8 cells (~1/100 ratio in diameter), we initially studied a model system where Au NPs were laterally distributed on a ‘planar’ surface in an attempt to mimic cell membrane labeling (see Figures 5a–d). With the increase of Au NP surface density, mean particle-to-particle distance reduces significantly and thus leads to a greater chance of plasmon coupling54, as also illustrated by our simulations (Figures 5a–c). Two NPs in close proximity formed an inter-particle region called a “hot spot” where the local electric field intensity was enhanced by several orders of magnitude as illustrated in Figures 5a–c (shown in red color). The number of such “hot spots” was strongly dependent on the NP surface density and the corresponding extinction spectra of the Au NP assemblies broadened significantly as the particle density increased (see Figure 5d). This change also elucidated why the CoM of wavelengths gradually shifted to the red as the incubation density of Au NPs increased (i.e., matching our experimental results reported in Figure 4b). However, we should also note that the simulated maximum possible CoM wavelength for the ‘2D’ NP assembly (~680 nm) was smaller than the largest measured mean value (~700 nm) in Figure 4b, which can be attributed to the plausible formation of not only 2D but also 3D assemblies of nanoparticles on or within the cell. The formation of 3D NP clusters can be due to partial endocytosis of NPs55 or dynamic remodeling of cell membrane curvature56. To explore it further, we also performed FDTD simulations on NPs assembled on spherical surfaces, in 3D volume-filling spaces, as well as planar multilayer structures (see e.g., Figure 5e, inset). The 3D volume-filling and multilayer models gave the largest coupling-induced red shifts up to ~90 nm, which closely match our experimental data (Figure 4). Previous simulations have also shown that plasmonic nanostructures assembled in 3D tend to exhibit much stronger plasmon-coupling shifts than 2D monolayer models57. Figure 5e presents an overall calibration function that includes not only 2D planar models (first 7 data points) but also 3D multilayer cases (last 2 data points, bilayer and trilayer, respectively). For the 3D morphology, the particle density is calculated by dividing the surface area of x-y plane by the total number of particles in 3D volume. This relationship depicted in Figure 5e predicts the saturation of the plasmon-coupling effect at a density of ~150 NP/μm2.
FDTD simulations of Au NP plasmon coupling at different nanoparticle densities.
(a–c) Simulated electric field density distribution of 30, 60 and 90 Au NPs within a 1.25-μm square. The simulation is performed using an excitation pulse with a power spectrum covering the range 490 nm to 960 nm and the results represent a weighted average of all frequencies and two orthogonal polarizations, simulating a uniform intensity across this bandwidth. (d) Normalized extinction spectra of 2D Au NP assemblies at different surface densities. Insets show randomly distributed Au NPs on planar surfaces at a particle density of 1.28 and 57.6 NP/μm2, respectively. (e) Calculated CoM wavelengths as a function of particle density. The first 7 data points are obtained by 2D assembly simulations, whereas the last 2 data points are obtained by 3D multilayer simulations (bilayer and trilayer, respectively), as illustrated in the insets.
This nonlinear relationship of the nanoparticle density and its corresponding wavelength CoM obtained through our FDTD simulations (Figure 5e) may also be used to roughly estimate the average number of bound NPs on CD4 cells. Using Figure 5e and assuming NP spatial coverage follows a uniform random variable across the cell surface, we can calculate the distribution of Au NP surface density on individual CD4 cells at different Au NP concentration values ([Au NP]) used during incubation (see Figure 6). Each histogram was also fitted by a normal distribution curve (grey), which infers the mean labeling density for each [Au NP] value. These estimations illustrate that the mean NP surface density on CD4 cells increased as a function of Au NP concentration during the incubation period and saturated at around 150 NP/μm2 when [Au NP] was above 3.6 × 1010 NP/mL (Figure 6). This saturation density of Au NPs estimated by our lensfree on-chip microscopy platform matches the values observed by lens-based imaging methods. For instance, using plasmon-coupling microscopy Reinhard et al. reported a saturation concentration of 200 NP/μm2 for 40-nm Au NPs on epidermal growth factor receptor (EGFR) expressing A431 cells58,59. Previous simulation studies on particle packing also show that the maximum coverage occurs at a particle density of around 114 NP/μm260. Based on Figure 6, the average surface coverage of Au NPs on CD4 cells can be estimated as ~50% at low [Au NP] (e.g., 9.6 × 109 NP/mL), while it reached close to 100% at higher [Au NP] (e.g., 1.92 × 1010 NP/mL). A surface coverage value that is higher than 100% implies over-packing of Au NPs on the CD4 cells and the transition of the aggregation morphology from 2D structures to 3D clusters. However, one should also emphasize that these estimated particle binding density values are based on our FDTD simulations reported in Figure 5, which treat the individual locations of nanoparticles as independent random variables. Stated differently, our simulations assume that plasmonic hot-spots appear randomly on the surface of the cell and statistically become more frequent as the density of the NPs increases. In cases where 2D and 3D clustering of nanoparticles occurs more frequently (making the individual locations of nanoparticles correlated), our simulation results (e.g., Fig. 5e) would no longer be accurate and cannot be used to estimate the NP density on the cell surface.
In summary, we demonstrated that plasmon-resonant metal NPs are effective contrast agents for lensfree on-chip holographic cytometry. Cellular-specific contrast on our lensfree imaging platform over a FOV of ~24 mm2 was achieved by first measuring the resonant optical properties of labeled cell lines with multispectral lensfree holographic imaging and second, automatically recognizing the spectral patterns by statistical analysis and machine learning. In its proof of concept, we successfully differentiated native CD4 cells from Au-CD4 cells with >93% accuracy. Furthermore, multiplexed lensfree recognition of CD4 and CD8 cells with an accuracy of >95% was also demonstrated by using antibody-conjugated Au and Ag NPs simultaneously. This lensfree holographic imaging and cytometry platform can serve as a powerful screening tool for studying cell properties or behavior with approximately two orders of magnitude larger FOV than conventional lens-based microscopy tools. Therefore, it has the unique capability of being able to simultaneously identify large numbers of different, but closely related, cell sub-types, while also allowing for high resolution inspection of individual cells. To this end, we also demonstrated, through the combination of FDTD simulations and lensfree multi-spectral imaging experiments, that holographic on-chip imaging enables the characterization of cell-receptor-mediated plasmon coupling and nanoparticle labeling density. Plausible labeling models at different Au NP concentrations were also provided. Given its simplicity, compactness and ultra-wide FOV, nanoparticle-enhanced lensfree imaging tools may find broad applications in cell biology and cytometry studies, such as quantitative screening of cells or cell surface receptors.
Methods
Chemicals and materials
100-nm diameter citrate-coated Au and Ag NPs were purchased from nanoComposix. CD4, CD8 mouse anti-human monoclonal antibody (denoted as antiCD4 and antiCD8) and recombinant human Interleukin 2 (IL2) were purchased from Invitrogen. 4,7,10,13,16,19,22,25,32,35,38,41,44,47,50,53-Hexadecaoxa-28,29-dithiahexapentacontanedioic acid di-N-succinimidyl ester (PEG7-NHS disulfide), hexadimethrine bromide (polybrene), bovine serum albumin (BSA) and phosphate buffered saline (PBS) were obtained from Sigma-Aldrich.
Preparation of antiCD4-conjugated Au NPs and antiCD8-conjugated Ag NPs
To prepare Au-antiCD4 conjugates, 24 μL of freshly prepared PEG7-NHS disulfide solution (5 mM, in PBS) was reacted with 5 μL of 1 mg/mL antiCD4 solution and the mixture was incubated for 2 h at 4°C to form PEGylated antibody disulfide complex. Next, PEGylated antibody was added to 0.3 mL of citrate-stabilized Au NPs (OD ~ 1) and mixed continuously on a shaker (100 rpm) for 3 h. AntiCD4-conjugated Au NPs (Au-antiCD4) were further stabilized by adding 5 μL of 5 mg/mL BSA. The solution was left overnight at 4°C to complete the conjugation reaction. Finally, Au-antiCD4 conjugates were collected by centrifugation (3500 g, 10 min) and resuspended in 30 μL of PBS. AntiCD8-conjugated Ag NPs (Ag-CD8) were prepared in a similar procedure as Au NPs, except that 10 μL of 5 mg/mL BSA was used as stabilizing reagent since Ag NPs were slightly less stable than Au NPs. All antibody-functionalized metal NPs were stored at 4°C for no more than two days before the labeling experiments.
Cell culture and nanoparticle labeling
Labeling CD4 cells with Au-antiCD4 conjugates followed a similar procedure described by Hansen et al.40. CD4 cells were maintained in RPMI 1640 medium under normal cell culture conditions (37°C, 5% CO2). For labeling, 0.5 mL of CD4 cells (105 cells in total) were placed in culture tubes. Polybrene was added to CD4 cells with a final concentration of 10 μg/mL. Polybrene served as a charge neutralization agent to increase the affinity of negatively charged antibody to the cell membrane during the incubation40. CD4 cells were then mixed with 4, 8, 15 and 60 μL of Au-antiCD4 conjugates (OD ~ 10) and incubated at 37°C with continuous mixing (100 rpm) overnight. After incubation, CD4 cells were fixed with 1.5% formaldehyde for 10 min at room temperature (RT). Unbound nanoparticles were removed from labeled cells by slow centrifugation (270 g, 5 min) and discarding the supernatant. The washing steps were repeated twice. Finally, labeled Au-CD4 cells were resuspended in 1× PBS buffer and store at 4°C for future imaging use. Similar incubation steps were also applied to label CD8 cells with Ag-antiCD8 conjugates. Different from CD4 cell culture, CD8 cells were maintained in RPMI 1640 medium with 20 ng/mL IL2 to promote cell growth.
Darkfield scattering microscopy
Darkfield scattering microscopy was performed on an inverted microscope (Nikon Eclipse TE2000-E) equipped with a high NA oil-immersion darkfield condenser (NA 1.2–1.43). A broadband halogen lamp was used for excitation. Images were collected by either a 60 × or 100 × objective and a digital color camera (Leica DFC 295).
Center of mass (CoM) of the absorption spectra
For the discrete cellular absorption spectra, the center of mass (CoM) was defined as:


where ai is the absorbance and λi is the wavelength for the ith data in the spectra. wi is the weighting factor calculated by (λi+1 − λi−1)/2. The wavelengths used to compute the center of mass ranged from 480 nm to 950 nm.
FDTD simulations
RSoft FullWAVE version 6.1 was used to perform the FDTD simulations. For monolayer simulations, the nanoparticle clusters occupied a region with size 1.25 μm × 1.25 μm × 0.1 μm. For bilayer simulations, the region was 0.75 μm × 0.75 μm × 0.20 μm. For trilayer simulations, the region was 0.75 μm × 0.75 μm × 0.30 μm. For monolayer simulations, the domain size was 1.322 μm × 1.322 μm × 0.172 μm. For bilayer and trilayer simulations, the domain size was 0.822 μm × 0.822 μm × 0.372 μm. The voxel size in all simulations was 5.5 nm × 5.5 nm × 5.5 nm. A perfectly matched layer with thickness 0.15 μm was included outside of the simulation domain. The time step was 0.003 μm and the total simulation time was 12 μm (where 1 μm of time is the time it takes light to travel 1 μm in vacuum.) The excitation beam was directed perpendicularly to the layered structures and was restricted to a region of 1.3 μm × 1.3 μm × 0.15 μm (0.8 μm × 0.8 μm × 0.35 μm for bilayer and trilayer structures); outside this region only the scattered light was present. The power scattered outward from this region was measured as a function of frequency, as was the power absorbed within the region. Extinction was computed as scattering plus absorption. The excitation field (either Ex or Ey depending on polarization) was a short pulse given by,

This provided a band of wavelengths in the excitation pulse. For each particle packing density, five random packings were simulated, each with both transverse-electric (TE) and transverse-magnetic (TM) polarizations and the results averaged. Random packings were generated by randomly selecting particle coordinates within the domain and placing a particle if it did not overlap with any other particles.
References
Greenbaum, A. et al. Imaging without lenses: achievements and remaining challenges of wide-field on-chip microscopy. Nat. Methods 9, 889–895 (2012).
Cui, X. et al. Lensless high-resolution on-chip optofluidic microscopes for Caenorhabditis elegans and cell imaging. Proc. Natl. Acad. Sci. USA 105, 10670–10675 (2008).
Ozcan, A. & Demirci, U. Ultra wide-field lens-free monitoring of cells on-chip. Lab Chip 8, 98–106 (2008).
Coskun, A. F., Su, T.-W. & Ozcan, A. Wide field-of-view lens-free fluorescent imaging on a chip. Lab Chip 10, 824–827 (2010).
Roda, A. et al. Portable device based on chemiluminescence lensless imaging for personalized diagnostics through multiplex bioanalysis. Anal. Chem. 83, 3178–3185 (2011).
Bishara, W., Su, T.-W., Coskun, A. F. & Ozcan, A. Lensfree on-chip microscopy over a wide field-of-view using pixel super-resolution. Opt. Express 18, 11181–11191 (2010).
Bishara, W., Zhu, H. & Ozcan, A. Holographic opto-fluidic microscopy. Opt. Express 18, 27499–27510 (2010).
Isikman, S. O., Bishara, W., Zhu, H. & Ozcan, A. Optofluidic tomography on a chip. Appl. Phys. Lett. 98, 161109 (2011).
Bishara, W., Isikman, S. & Ozcan, A. Lensfree optofluidic microscopy and tomography. Ann. Biomed. Eng. 40, 251–262 (2011).
Tseng, D. et al. Lensfree microscopy on a cellphone. Lab Chip 10, 1787–1792 (2010).
Isikman, S. O. et al. Lens-free optical tomographic microscope with a large imaging volume on a chip. Proc. Natl. Acad. Sci. USA 108, 7296–7301 (2011).
Isikman, S. O., Bishara, W. & Ozcan, A. Partially coherent lensfree tomographic microscopy. Appl. Opt. 50, H253–H264 (2011).
Su, T.-W., Xue, L. & Ozcan, A. High-throughput lensfree 3D tracking of human sperms reveals rare statistics of helical trajectories. Proc. Natl. Acad. Sci. USA 109, 16018–16022 (2012).
Mudanyali, O. et al. Compact, light-weight and cost-effective microscope based on lensless incoherent holography for telemedicine applications. Lab Chip 10, 1417–1428 (2010).
Seo, S., Su, T.-W., Tseng, D. K., Erlinger, A. & Ozcan, A. Lensfree holographic imaging for on-chip cytometry and diagnostics. Lab Chip 9, 777–787 (2009).
Seo, S. et al. High-throughput lens-free blood analysis on a chip. Anal. Chem. 82, 4621–4627 (2010).
Su, T.-W., Erlinger, A., Tseng, D. & Ozcan, A. Compact and light-weight automated semen analysis platform using lensfree on-chip microscopy. Anal. Chem. 82, 8307–8312 (2010).
Mudanyali, O., Oztoprak, C., Tseng, D., Erlinger, A. & Ozcan, A. Detection of waterborne parasites using field-portable and cost-effective lensfree microscopy. Lab Chip 10, 2419–2423 (2010).
Stybayeva, G. et al. Lensfree holographic imaging of antibody microarrays for high-throughput detection of leukocyte numbers and function. Anal. Chem. 82, 3736–3744 (2010).
Greenbaum, A., Sikora, U. & Ozcan, A. Field-portable wide-field microscopy of dense samples using multi-height pixel super-resolution based lensfree imaging. Lab Chip 12, 1242–1245 (2012).
Mudanyali, O. et al. Wide-field optical detection of nanoparticles using on-chip microscopy and self-assembled nanolenses. Nat. Photonics 7, 247–254 (2013).
Schultz, S., Smith, D. R., Mock, J. J. & Schultz, D. A. Single-target molecule detection with nonbleaching multicolor optical immunolabels. Proc. Natl. Acad. Sci. USA 97, 996–1001 (2000).
El-Sayed, I. H., Huang, X. & El-Sayed, M. A. Surface plasmon resonance scattering and absorption of anti-EGFR antibody conjugated gold nanoparticles in cancer diagnostics: applications in oral cancer. Nano Lett. 5, 829–834 (2005).
Kang, B., Austin, L. A. & El-Sayed, M. A. Real-time molecular imaging throughout the entire cell cycle by targeted plasmonic-enhanced Rayleigh/Raman spectroscopy. Nano Lett. 12, 5369–5375 (2012).
Kneipp, J., Kneipp, H., Wittig, B. & Kneipp, K. One- and two-photon excited optical pH probing for cells using surface-enhanced Raman and hyper-Raman nanosensors. Nano Lett. 7, 2819–2823 (2007).
Cognet, L. et al. Single metallic nanoparticle imaging for protein detection in cells. Proc. Natl. Acad. Sci. USA 100, 11350–11355 (2003).
Leduc, C. C., Jung, J.-M., Carney, R. R., Stellacci, F. & Lounis, B. Direct investigation of intracellular presence of gold nanoparticles via photothermal heterodyne imaging. ACS Nano 5, 2587–2592 (2011).
Wang, H. et al. In vitro and in vivo two-photon luminescence imaging of single gold nanorods. Proc. Natl. Acad. Sci. USA 102, 15752–15756 (2005).
Tai, S.-P. et al. Molecular imaging of cancer cells using plasmon-resonant-enhanced third-harmonic-generation in silver nanoparticles. Adv. Mater. 19, 4520–4523 (2007).
Tong, L., Wei, Q., Wei, A. & Cheng, J.-X. Gold nanorods as contrast agents for biological imaging: optical properties, surface conjugation and photothermal effects. Photochem. Photobio. 85, 21–32 (2009).
Atlan, M. et al. Heterodyne holographic microscopy of gold particles. Opt. Lett. 33, 500–502 (2008).
Absil, E. et al. Photothermal heterodyne holography of gold nanoparticles. Opt. Express 18, 780–786 (2010).
Warnasooriya, N. et al. Imaging gold nanoparticles in living cell environments using heterodyne digital holographic microscopy. Opt. Express 18, 3264–3273 (2010).
Verpillat, F., Joud, F., Desbiolles, P. & Gross, M. Dark-field digital holographic microscopy for 3D-tracking of gold nanoparticles. Opt. Express 19, 26044–26055 (2011).
Wu, H., Wheeler, A. & Zare, R. N. Chemical cytometry on a picoliter-scale integrated microfluidic chip. Proc. Natl. Acad. Sci. USA 101, 12809–12813 (2004).
Mao, X., Lin, S.-C. S., Dong, C. & Huang, T. J. Single-layer planar on-chip flow cytometer using microfluidic drifting based three-dimensional (3D) hydrodynamic focusing. Lab Chip 9, 1583–1589 (2009).
Cho, S. H., Qiao, W., Tsai, F. S., Yamashita, K. & Lo, Y.-H. Lab-on-a-chip flow cytometer employing color-space-time coding. Appl. Phys. Lett. 97, 093704 (2010).
Lapsley, M. I., Wang, L. & Huang, T. J. On-chip flow cytometry: where is it now and where is it going? Biomarkers in Medicine 7, 75–78 (2013).
Skommer, J. et al. Multiparameter Lab-on-a-Chip flow cytometry of the cellcycle. Biosens. Bioelectron. 42, 586–591 (2013).
Hansen, P. et al. Physics of a rapid CD4 lymphocyte count with colloidal gold. Cytom. Part A 81A, 222–231 (2011).
Luo, Y., Sun, W., Gu, Y., Wang, G. & Fang, N. Wavelength-dependent differential interference contrast microscopy: multiplexing detection using nonfluorescent nanoparticles. Anal. Chem. 82, 6675–6679 (2010).
Wang, H., Rong, G., Yan, B., Yang, L. & Reinhard, B. M. Optical sizing of immunolabel clusters through multispectral plasmon coupling microscopy. Nano Lett. 11, 498–504 (2011).
Seekell, K. et al. Hyperspectral molecular imaging of multiple receptors using immunolabeled plasmonic nanoparticles. J. Biomed. Opt. 16, 116003 (2011).
Jolliffe, I. T. Principal component analysis. (2nd ed.). (Springer, Berlin; 2002).
Press, W. H., Teukolsky, S. A., Vetterling, W. T. & Flannery, B. P. in Numberical recipes: the art of scientific computing. (3rd ed.) (Cambridge University Press, New Your; 2007).
Taylor, J. M. G., Fahey, J. L., Detels, R. & Giorgi, J. V. CD4 Percentage, CD4 number and CD4: CD8 ratio in HIV infection: which to choose and how to use. J. Acquir. Immune. Defic. Syndr. 2, 114–124 (1989).
Rodriguez, W. R. et al. A microchip CD4 counting method for HIV monitoring in resource-poor settings. PLoS Med 2, e182 (2005).
Chang, C.-C. & Lin, C.-J. A Library for support vector machines (LIBSVM). http://www.csie.ntu.edu.tw/~cjlin/libsvm/ (Accessed 4/2/2013).
Liu, X., Atwater, M., Wang, J. & Huo, Q. Extinction coefficient of gold nanoparticles with different sizes and different capping ligands. Colloids Surf. B 58, 3–7 (2007).
Umezawa, K., Nakamura, Y., Makino, H., Citterio, D. & Suzuki, K. Bright, Color-tunable fluorescent dyes in the visible-near-infrared region. J. Am. Chem. Soc. 130, 1550–1551 (2008).
Mitronova, G. Y. et al. New fluorinated rhodamines for optical microscopy and nanoscopy. Chem. Eur. J. 16, 4477–4488 (2010).
Thakor, A. S., Jokerst, J., Zavaleta, C., Massoud, T. F. & Gambhir, S. S. Gold nanoparticles: a revival in precious metal administration to patients. Nano Lett. 11, 4029–4036 (2011).
Dreaden, E. C., Alkilany, A. M., Huang, X., Murphy, C. J. & El-Sayed, M. A. The golden age: gold nanoparticles for biomedicine. Chem. Soc. Rev. 41, 2740–2779 (2012).
Halas, N. J., Lal, S., Chang, W.-S., Link, S. & Nordlander, P. Plasmons in strongly coupled metallic nanostructures. Chem. Rev. 111, 3913–3961 (2011).
Bartczak, D., Sanchez-Elsner, T., Louafi, F., Millar, T. M. & Kanaras, A. G. Receptor-mediated interactions between colloidal gold nanoparticles and human umbilical vein endothelial cells. Small 7, 388–394 (2011).
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Aaron, J., Travis, K., Harrison, N. & Sokolov, K. Dynamic imaging of molecular assemblies in live cells based on nanoparticle plasmon resonance coupling. Nano Lett. 9, 3612–3618 (2009).
Wang, J., Boriskina, S. V., Wang, H. & Reinhard, B. M. Illuminating epidermal growth factor receptor densities on filopodia through plasmon coupling. ACS Nano 5, 6619–6628 (2011).
Wang, J., Yu, X., Boriskina, S. V. & Reinhard, B. M. Quantification of differential ErbB1 and ErbB2 cell surface expression and spatial nanoclustering through plasmon coupling. Nano Lett. 12, 3231–3237 (2012).
Specht, E. The best known packings of equal circles in the unit square. http://hydra.nat.uni-magdeburg.de/packing/csq/csq.html#Overview (Accessed 4/2/2013).
Acknowledgements
Ozcan Group gratefully acknowledges the support of the Presidential Early Career Award for Scientists and Engineers, Army Research Office Young Investigator Award, National Science Foundation CAREER Award, Office of Naval Research Young Investigator Award, National Institutes of Health Director's New Innovator Award DP2OD006427 from the Office of the Director, National Institutes of Health and National Center for Advancing Translational Sciences UCLA CTSI Grant UL1TR000124. The authors also acknowledge D. Tseng of Ozcan Research Group for his assistance with the figures.
Author information
Authors and Affiliations
Contributions
Q.W. conducted the experiments and processed the resulting data. E.M. performed numerical analysis and electromagnetic simulations of the plasmonic NP effects. H.Q. and Z.W. contributed to methods and experiments. Q.W. and A.O. planned and executed the research and wrote the manuscript. R.S. and A.O. supervised the project.
Ethics declarations
Competing interests
A.O. is the co-founder of a start-up company that aims to commercialize lensfree microscopy tools.
Electronic supplementary material
Supplementary Information
Supplementary Information and Figures
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wei, Q., McLeod, E., Qi, H. et al. On-Chip Cytometry using Plasmonic Nanoparticle Enhanced Lensfree Holography. Sci Rep 3, 1699 (2013). https://doi.org/10.1038/srep01699
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01699
This article is cited by
-
Digital plasmonic holography
Light: Science & Applications (2018)
-
Demosaiced pixel super-resolution for multiplexed holographic color imaging
Scientific Reports (2016)
-
Automated single-cell motility analysis on a chip using lensfree microscopy
Scientific Reports (2014)
-
Spectral Demultiplexing in Holographic and Fluorescent On-chip Microscopy
Scientific Reports (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.