Abstract
Epigenetic modifications in eukaryotic genomes occur primarily in the form of 5-methylcytosine (5 mC). These modifications are heavily involved in transcriptional repression, gene regulation, development and the progression of diseases including cancer. We report a new single-molecule assay for the detection of DNA methylation using solid-state nanopores. Methylation is detected by selectively labeling methylation sites with MBD1 (MBD-1x) proteins, the complex inducing a 3 fold increase in ionic blockage current relative to unmethylated DNA. Furthermore, the discrimination of methylated and unmethylated DNA is demonstrated in the presence of only a single bound protein, thereby giving a resolution of a single methylated CpG dinucleotide. The extent of methylation of a target molecule could also be coarsely quantified using this novel approach. This nanopore-based methylation sensitive assay circumvents the need for bisulfite conversion, fluorescent labeling and PCR and could therefore prove very useful in studying the role of epigenetics in human disease.
Similar content being viewed by others
Introduction
DNA methylation is one of the most important and frequently occurring epigenetic modifications in mammalian cells and plays an essential role in regulating cell growth and proliferation. In humans, the most common epigenetic modification of DNA involves the addition of a methyl group at the 5-carbon position of cytosine (5-methylcytosine or 5 mC), which occurs exclusively at symmetric CG sites on the DNA double helix and are referred to as CpG dinucleotides. Hypermethylation of the promoter sequences of various genes has generally been associated with transcriptional repression through mechanisms such as the recruitment of methylated CpG binding proteins (MBDs), histone deacetylation and chromatin remodeling1,2. Furthermore, aberrant methylation in the promoter sequences of various genes can point to specific pathways disrupted in almost every tumor type including cancers of the prostate, breast, head and neck, lung and liver, whilst correlating with disease severity and metastatic potential3,4,5,6,7,8. In fact, the tumor prevalence of many methylation markers is considerably higher than that of genetic markers4; one example being the hypermethylation of CpG dinucleotides in the promoter sequence of the glutathione S-transferase pi (GSTP1) gene and is observed in over 90% of prostate cancer patients9. Methylation analysis will therefore likely play a pivotal role in the diagnosis and treatment of such diseases.
Interestingly, cancer-specific methylated DNA from most tumor types is present in biopsy specimens and also exist at very low concentrations in the form of free-floating DNA shed by apoptotic cancer cells4. Current genome-wide methylation analysis techniques rely on bisulfite genomic sequencing10 (bisulfite conversion of DNA, PCR amplification and DNA sequencing) and typically require large sample volumes due to DNA degradation during bisulfite conversion11, can exhibit low amplification efficiency and PCR bias12 and are labor intensive. Targeted methods involving analysis at specific loci or groups of genes such as methylation specific PCR (MSP)12, MethyLight13,14 and DNA microarrays15 overcome the need for sequencing but still rely on bisulfite conversion, amplification and complex probe design. Therefore, a bisulfite free, amplification free method capable of rapidly and accurately determining the methylation status of panels of genes from minute clinical sample volumes could be of tremendous clinical value.
This paper demonstrates a new single molecule assay for determining the methylation status of DNA using solid-state nanopores. Nanopores use the principle of ionic current spectroscopy to electrically interrogate individual DNA molecules with the sensitivity to discern subtle structural motifs16,17. Fabrication of these devices typically involves the physical sputtering of a single nanometer sized aperture in a dielectric membrane using a focused electron beam18,19. The electrophoretic transport of biomolecules through these nano-scale pores has enabled the study of various biophysical phenomena at the single molecule level20, with potential applications in DNA sequencing and medical diagnostics16,21,22,23. (For reviews of nanopore research, see refs 24,25,26,27,28,29,30) Recently, methylated and unmethylated DNA has been examined optically in nanofluidic channels using fluorescently labeled proteins bound to the methylation sites31,32. Nanopore-based ionic current spectroscopy, however, is ideal for single molecule epigenetic analysis eliminating the need for optical measurements. Using nanopore based ionic current spectroscopy, the differentiation of methylcytosine from cytosine has previously been demonstrated by passing these individual nucleotides through a biological nanopore33, requiring an exonuclease based cleaving of the bases from the original molecule. To date ionic current measurements obtained using a solid-state nanopore, have yet to differentiate methylated from unmethylated single molecules of DNA34,35.
Herein, we demonstrate the electrical discrimination of unmethylated and methylated DNA using solid-state nanopores. Our technique does not require bisulfite conversion, sequencing or fluorescent tags but rather relies on the detection of methylated CpG dinucleotides in DNA by labeling with a 75 amino acid region of the methyl DNA binding protein MBD1, which includes a his-tagged single DNA binding domain and will hereafter be referred to as MBD-1x. The translocation of the methylated DNA - MBD-1x complex through a solid-state nanopore induced approximately a 3-fold increase in the measured blockage current relative to unmethylated DNA. The binding of a single MBD-1x protein to a methylated DNA fragment was sufficient for differentiation with high fidelity, thereby enabling single CpG dinucleotide sensitivity. Methylation could also be coarsely quantified based on the number of bound MBD-1x proteins per molecule, characterized by distinct timescales in the event translocation time histograms. As a result, this amplification- and fluorescent label-free, single molecule assay could be significantly useful in the rapid screening of epigenetic biomarkers for the early detection of diseases such as cancer.
Results
Detection of unlabeled methylated and unmethylated DNA
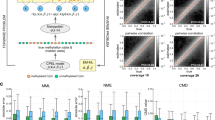
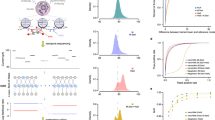
The electrophoretic transport of double stranded DNA (dsDNA) through a solid-state nanopore is illustrated in the schematic of Figure 1a, the inset showing a transmission electron microscope image of a ~4.2 nm diameter pore. The detection of unmethylated and methylated dsDNA in the absence of MBD-1x was performed using a ~4.2 nm pore fabricated in 20 nm-thick SiN membranes according to methods described previously18,19. Briefly, DNA was introduced into the cis chamber. A positive voltage was applied to the trans side resulting in the passage of dsDNA through the nanopore to the trans side. The target fragment used in these studies was an 827 bp region of DLX1 (see Supplementary Figure s1 for sequence information), a homeobox gene associated with forebrain development37. Aberrant methylation of DLX1 has been reported in several cancers, including lymphoma38 and brain tumors39. Furthermore, analysis of publicly available methylation profiling data40 identified significant hypermethylation of DLX1 promoter in lung adenocarcinomas (see Supplementary Figure s2). Therefore, methylated DLX1 promoter has potential clinical utility in cancer diagnosis. This 827 bp DLX1 region contained 36 CpG dinucleotides, which were methylated in-vitro using the M.Sss1 DNA methyltransferase. The methylation of the dsDNA was confirmed using the restriction enzyme HhaI (see Supplementary Figure s3). Methylated DLX1 will hereafter be referred to as mDLX1 and unmethylated DLX1 will be referred to as uDLX1. Ionic current traces produced by the electrophoretic transport of mDLX1 through the nanopore at various voltages are shown in Figure 1b, each downward current pulse indicative of the passage of a single mDLX1 molecule though the nanopore. A magnified view of these events is presented in Figure 1c, the key parameters of interest being the blockage current, ΔI, induced by the passage of the molecule through the pore and the event duration or translocation time, τd. Chemical structures of cytosine and methylated-cytosine, schematics of CpG dinucleotides in unmethylatd dsDNA and methylated dsDNA, data traces of uDLX1 and mDLX1 recorded at 300 mV are presented in Figure 1d.
Detection of methylated and unmethylated DNA using a solid-state nanopore.
(a) Schematic diagram of a nanopore setup. A focused electron beam of TEM is used to sculpt a nanopore in a thin (~20 nm) silicon nitride membrane; the nanopore chip is sealed between two fluidic cell chambers containing conductive electrolyte; a voltage is applied across this setup to induce the translocation of single dsDNA molecules through the pore as shown. (Inset) TEM image of typical ~4.2 nm diameter nanopore used in DNA measurements (scale bar is 10 nm). (b) Characteristic ionic current traces obtained from the translocation of mDLX1 (827 bp dsDNA with 36 potential CpG methylation sites). Traces were recorded in 600 mM KCl at pH 8.0 electrolyte at various voltage levels. (c) A typical DNA induced current blockade. Parameters of interest are the open pore current, IO, residual blocking current, IB, (occurs while a single DNA molecule translocates through the nanopore), blockage current ΔI = IB − IO and translocation time of the molecule, tduration. (d) Schematic showing (top) the chemical difference between cytosine and methylated cytosine; (middle) unmethylated versus a fully methylated CpG dinucleotide in dsDNA. Data traces of unmethylated- (bottom-left) and methylated-DLX1 (bottom-right) were recorded at 300 mV, showing similarity between both data traces. (e) Comparison of mDLX1 and uDLX1 transport. ΔI and τd plots as a function of applied voltage. τd and ΔI refers to the time constant and the blocking current respectively at each voltage. All points are the value of the fit with standard error. Second order of polynomial fit to ΔI and exponential fit to τd are also shown in short dash (black fits to uDLX1 and red to mDLX1). Each data are overlaid with over n = 1167 separate translocation events recorded per data point. Methylated and unmethylated fragments are indistinguishable. (f). τd (top) and ΔI (bottom) histograms for mDLX1 versus uDLX1 at 500 mV (n > 2153), showing similarity with τd_mDLX1 = 0.124 ± 0.006 ms, τd_uDLX1 = 0.135 ± 0.006 ms, ΔI_mDLX1 = −449.5 ± 5.4 pA and ΔI_uDLX1 = −440.8 ± 24.3 pA.
Figure 1e compares the translocation properties (ΔI and tduration) of methylated and unmethylated DLX1 through the nanopore as a function of applied voltage. Each data point on these plots consists of over 1167 separately recorded DNA translocation events. Voltage-dependent transport of both mDLX1 and uDLX1 were observed; step increases in the applied voltage resulting in higher electrophoretic forces on the molecule and therefore shorter translocation times through the pore41,42. As seen in Figure 1e, the single molecule sensitivity of a solid-state nanopore alone is not sufficient to distinguish methylated from unmethylated DNA with any statistical significance. This is reiterated by the similar τd and ΔI histograms (Figure 1f), each distribution containing over 2153 translocation events recorded at 500 mV. Notably, the time constants for mDLX1 and uDLX1 obtained by exponential fitting to the translocation time histograms of Figure 1f are within 10% of each other (τM = 0.124 ± 0.006 ms, τU = 0.135 ± 0.006 ms), confirming the inability to consistently distinguish methylated from unmethylated DNA. This result is not surprising given the subtle structural and chemical differences that exist between 5-methylcystosine and cytosine (Figure 1d). We therefore conclude that these differences along with reported differences in the nanomechanical properties of methylated versus unmethylated DNA34, are not sufficient to give rise to detectable differences in their respective ionic current signatures. Our results are consistent with previous findings35 and reiterate the need for a methylation specific label in nanopore based methylation studies.
Formation of DNA/MBD-1x complex
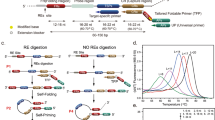
To specifically label methylated DNA, we used the 75 amino acid methylated DNA binding domain of the protein MBD1. MBD1 plays an important role in gene silencing by recruiting AFT7IP, which in turn recruits factors such as the histone methyltransferase SETDB1 and is essential in histone deacetylation and transcriptional repression in vertebrates43. Importantly, MBD1 binds symmetrically to methylated but not unmethylated CpG dinucleotides with high affinity44 and specificity43. The 75 amino acid MBD-1x was expressed in E. coli (see Materials and Methods) and protein purity was verified using Coomassie stained gels and Western blot analysis (Supplementary Figure s4). Figure 2a illustrates the crystal structures of typical B-form dsDNA and the methylated-DNA/MBD complex29,30. X-ray diffraction and NMR spectroscopy confirm that the binding domain of MBD1 occupies ~5–6 bp in the major groove of the dsDNA helix upon binding to a single methylated CpG dinucleotide45,46. It is therefore likely that only 21–25 of the 36 methyl-CpG sites in the DLX1 probe used here will serve as functional binding sites for MBD-1x, as only these regions contain sufficient spacing between sites to physically accommodate the protein. The relatively small occlusion area of MBD-1x (5–6 bps) also makes this protein ideal for nanopore based methylation analysis. Other MBD family proteins such as MBD2 and MeCP2 are known to protect ~12–14 bp around a single binding site47 and thus would provide less spatial resolution in nanopore based ionic current measurements. A top-view of MBD bound to dsDNA, derived from the crystal structure of the complex, is shown in Figure 2b. A cross-sectional diameter of ~5 nm is estimated for the complex containing a single MBD molecule, significantly larger than the ~2.2 nm cross-sectional diameter of B-DNA. With multiple bound MBD proteins, this diameter is estimated at ~7.6 nm as methylated binding sites follow the rotation of the major groove on dsDNA (Supplementary Figure s5). Gel shift assays (Figure 2c) were used to optimize binding conditions for complex formation prior to nanopore measurements. In the presence of uDLX1, no complex formation was observed (lanes 1–3). In contrast, when mDLX1 was combined with MBD-1x, robust complex formation is observed (lanes 5–9). Complex formation increases as MBD-1x concentration is increased. Importantly, this protein-DNA complex formation occurs at salt concentrations as high as 600 mM KCl (Supplementary Figure s6), which is necessary for achieving high signal to noise ratios in nanopore detection experiments. We estimate that a 30:1 excess of MBD-1x to mDLX1 is sufficient to saturate the available methylated binding sites on the target fragment.
Differentiation of unmethylated DNA from mDLX1/MBD-1x complex.
(a) Structure of B-form dsDNA (left) and methylated DNA/MBD complex (right). A single MBD protein binds to the methylated CpG site on the major groove of dsDNA, occupying about 6 bps (PDB ID: 1IG4). (b) Top-down view: the cross-sectional diameter of the complex with a single bound MBD protein is ~5 nm. Multiple bound proteins along the DNA major groove increase complex diameter to ~7.6 nm. (c) Gel-shift assay showing the high affinity and specificity of MBD-1x for methylated but not unmethylated DNA. When increasing amounts of MBD-1x protein were incubated with uDLX1, no DNA-protein complex was formed (lanes 1–3), but when mDLX1 was included a robust, dose-dependent increase in mDLX1 - MBD-1x complex formation was observed (lanes 5–9) Lane 5 and 9 show 1:5 and 1:30 (mDLX:MBD-1x), respectively. Samples were fractionated on an 8% non-denaturing polyacrylamide gel and visualized using autoradiography. (d) Nanopore ionic current traces recorded in 600 mM KCl, pH 8.0 at 600 mV; uDLX1 events (left), mDLX1/MBD-1x events (right). (e) Characteristic translocation signatures for uDLX1 (bottom) versus the complex (top) through a ~12 nm pore. Scale bar is 10 nm in the TEM image. Qualitatively, the mDLX1/MBD-1x complex induces longer, deeper current blockades relative to uDLX1. (f) tduration (left) and ΔI (right) histograms at 600 mV for uDLX1 (shown in blue − n = 857) and mDLX1 (shown in red − n = 197). Unmethylated DNA and the complex are clearly distinguishable. Exponential fits give time constants of τd_uDLX1 = 0.103 ± 0.005 ms and τd_mDLX1 = 1.43 ± 0.03 ms respectively.
Discrimination of mDLX1/MBD-1x complex from unmethylated DNA
Control experiments with nanopores of diameter of 4.5 nm and 7 nm, where these sizes are comparable with a single MBD-1x bound to DNA (5 nm) and multiple MBD-1x bound to DNA (7.6 nm), showed that mDLX1/MBD-1x complex cannot translocate through these pores (Supplementary Figure s7 and s8). Consequently, we utilized pores with larger diameters than the diameter of mDLX1/MBD-1x complex. The transport of uDLX1 at 1 nM of final concentration and the 1:30 mDLX1/MBD-1x complex at 10 pM through ~12 nm diameter pore at an applied voltage of 600 mV is shown in Figure 2d and characteristic events are shown in Figure 2e. A lower concentration of mDLX1/MBD-1x complex was used to explore a lower limit of detection. Notably, the transport of the complex induced deeper current blockades and longer translocation times relative to uDLX1. This is best represented in the τd histogram and ΔI all-points histograms of Figure 2f consisting of n = 857 unmethylated and n = 197 methylated events. Event flux (number of events per second) was expectedly less in the case of the complex versus uDLX1 as the entropic barrier associated with transport of the complex through the pore is significantly higher relative to uDLX1, in addition to more steric hindrance encountered by the complex during translocation. Fitting exponentials to the τd histogram gave time constants of τM = 1.43 ± 0.03 ms, τU = 0.103 ± 0.005 ms for mDLX1 and uDLX1 respectively, revealing the ability to statistically differentiate these populations. It should be noted that the DNA-protein interactions can be reversible as the KD can be from 106 to 870 nM44. This can indeed result in a wider distribution of the translocation duration due to varying number of bound protein on each DNA. However, we also note that the mDLX1/MBD-1x was clearly distinguishable from the uDLX1 since the translocation durations were different by over an order of magnitude. Furthermore, an all-point ΔI histogram provided a detailed view of the translocation of mDLX1/MBD-1x translocation through the nanopore. The ΔI histogram for the mDLX1/MBD-1x complex shows both a deep current blockade level and a shallower blockade level consistent with free DNA in the absence of protein. This demonstrates that the nanopore can indeed coarsely detect protein-bound regions as well as protein-free region on a single molecule, thereby enabling methylation mapping (Supplementary Figure s9). To confirm that the deeper blockade levels observed in the ΔI histogram were due to the DNA/protein complex and not due to the presence of unbound MBD-1x protein, control experiments examining the transport of the free protein were attempted. No free MBD-1x translocation events were observed (Supplementary Figure s10), because MBD-1x is positively charged in pH 8 electrolyte, thus will not migrate through the pore under the voltage polarity used in these experiments. Discrimination experiments using a mixture of uDLX1 and the mDLX1/MBD-1x complex were also carried out (Supplementary Figure s11), again with significant differences at deeper current blockage in prolonged translocation were observed in the transport of complex over shallow short duration blockages of uDLX1. This data confirms that a nanopore based technique can differentiate methylated DNA from unmethylated DNA with high confidence using a methylation specific label.
Methylation quantification
To quantify the extent of DLX1 methylation, various ratios of MBD-1x to mDLX1 were incubated and then translocated through nanopores of diameter ranging from 9 to 10nm. A pore diameter of 9–10 nm was specifically selected to allow for slower complex translocation (see supplementary video of MD simulation). Translocation data for 1:30, 1:5 and 1:1 ratios of mDLX1/MBD-1x are shown in Figures 3a, 3b and 3c respectively. Each experiment involved translocating uDLX1 as a control fragment (lower insert), followed by translocation of the DNA-protein complex through the same nanopore. Current signatures of uDLX1 and mDLX/MBD-1x complex were compared via histogram of peak blocking current (uDLX1 in black and mDLX1/MBD-1x complex in red) along with a TEM image of each of the nanopore used. Figures 3a to 3c also qualitatively show that by lowering the ratio of protein to DNA, thereby reducing the mean number of bound proteins per DNA molecule, a measurable reduction in the translocation time of the complex can be observed. This is best visualized in the normalized translocation time histogram in Figure 3d. As can be seen in Figure 3e-left, for all DNA/protein ratios examined, mDLX1/MBD-1x can be clearly distinguished from uDLX1 based on blockage amplitude, ΔI. The complex remains clearly distinguishable even at the lowest protein/DNA ratios examined. Fitting a Gaussian function to the peak value of the blocking current of ΔI gave current signatures of mDLX1/MBD-1x and uDLX1 at all ratios. Current signatures of mDLX1/MBD-1x complex were obtained at ΔI1:30 = −2.01 ± 0.5 nA, ΔI1:5 = −3.09 ± 0.44 nA and ΔI1:1 = −2.65 ± 0.37 nA, while uDLX1 through the same pores showed current signatures of ΔIuDLX1_at1:30 = −0.76 ± 0.19 nA, ΔIuDLX1_1:5 = −0.67 ± 0.07 nA and ΔIuDLX1_1:1 = −0.87 ± 0.24 nA. Overall, regardless of the ratio of MBD-1x to mDLX1, the nanopore can detect and identify mDLX1/MBD-1x complex from uDLX1 by about a 3-fold larger current signature.
Methylation quantification based on number of bound MBD-1x proteins.
MBD-1x protein was incubated with methylated DLX1 DNA at ratios of (a) 1:30, (b) 1:5 and (c) 1:1. Characteristic current signatures representing the mDLX1/MBD-1x complex (top) and unmethylated DLX1 DNA (bottom) through 9–10 nm diameter pores are shown. Current signature histogram of unmethylated DLX1 (black) and methylated DLX1-MBD-1x complex (red). Histogram was generated with peak current signature value of each event. (Inset) Scale bar is 10 nm in TEM images. (d) Translocation time histograms representing the mDLX1/MBD-1x complex and unmethylated DLX1 (inset). (e) Methylation Detection (left): Complexes formed with any ratio of MBD-1x can be discriminated from uDLX1 using blockage current alone (~3-fold increase in blockage current induced by the complex is seen). Methylation Quantification (right): Complexes formed with different ratios of protein can be differentiated based on the number of bound MBD-1x molecules. Time constants for the complexes are shown by the red circles: J1:30 = 4.51 ± 0.48 ms, J1:5 = 1.67 ± 0.17 ms and J1:1 = 1.01 ± 0.09 ms. Corresponding time constants for uDLX1 were in the range of 0.107 – 0.184 ms MBD-1x on complexes were quantified with extended translocation duration. 1:1 complex showed ~7-fold prolonged translocation duration, 1:5 at ~12-fold and 1:30 at ~31-fold respectively than unmethylated DNA.
Given a 1:1 DNA-protein ratio, the number of bound proteins per DNA molecule can be calculated using a Poisson limited random statistical distribution48. According to this model, the probability that a single DNA molecule will contain one or fewer bound proteins is ~74%. Therefore, the majority of translocation events observed in Figure 3c can be credited to the binding of one MBD-1x protein per mDLX1 molecule (free DNA translocation events not included in the histogram) and overlapping all-points histogram of blocking currents between uDLX1 and mDLX1/MBD-1x indicates one or fewer bound protein to the DNA (Supplementary Figure s12). Furthermore, as rms current noise is identical in the preceding measurements, we conclude that a methylated DNA fragment with a single bound protein can give a ~305% enhancement in ionic current relative to unmethylated DNA. This confirms that the nanopore based methylation analysis technique presented here can indeed detect the presence of a single bound protein on average on methylated DNA with the sensitivity of a single CpG dinucleotide.
Figure 3e-right, shows distinct time scales of τ1:30 = 4.51 ± 0.48 ms, τ1:5 = 1.67 ± 0.17 ms and τ 1:1 = 1.01 ± 0.09 ms calculated for the 1:30, 1:5 and 1:1 distributions respectively, based on an exponential fitting to the histogram in Figure 3d. Using this method, methylation quantification in the time domain based on the number of bound proteins is indeed possible. The uDLX1 control, fitted to in range of 0.107 ~ 0.184 ms, is shown in the inset of Figure 3d. The distinct time constants pertaining to the complex likely result from translocation involving interactions with the pore walls. To understand the nature of these protein-pore interactions, molecular dynamics (MD) simulations were conducted as shown in Figure 4. Figures 4a and 4b illustrate the transport of 63 bp dsDNA with 3 bound MBD proteins through 12 nm and 10 nm diameter nanopores respectively. Temporal snapshots from the MD trajectory reveal that the complex interacts minimally with the pore walls during translocation through a larger 12 nm pore. In contrast, interactions between the complex and the pore are observed in smaller 10 nm pores, the center of mass of the complex remaining anchored in the pore upon completion of the simulation (Figure 4d). As nanopore diameter is reduced further to 9 nm (Figure 4c), the presence of even a single protein can induce polymer-pore interactions and the capture of the complex in the pore, resulting in longer blockade times. The simulation results agree with experimental data in general. Time constants for 1:30 mDLX1/MBD-1x complexes through a ~12 nm pore (1.43 ± 0.03 ms) were more than a factor of 3 less than translocation time constants for 1:30 complexes through a ~10 nm pore (4.51 ± 0.48 ms), confirming faster translocation through larger pores. Comparable time constants were measured for 1:5 complexes through a 9 nm pore. The detailed view of an experimental data trace from an individual mDLX1/MBD-1x shows slow translocation of the complex due to polymer-pore interactions (Supplementary Figure s13). These interactions are both hydrophobic and electrostatic in nature. Once a protein or DNA contacts the pore wall, Van der Waals interactions between the biomolecule and the pore wall slow down the translocation velocity of biomolecule as reported previously with single-stranded DNA49. Electrostatic polymer-pore interactions are also likely and have been reported to slow DNA in systems where the nanopore surface charge is opposite in polarity to the charge on the translocating biomolecule42,50. As our experiments were carried out in pH 8 electrolyte and as the isoelectric points of MBD1x and the SiN pore are 8.85 and ~4 respectively51,52, we expect electrostatic interactions between the positively charged protein and the negatively charged nanopore surface. Thus, longer translocation times are expected as the number of bound proteins per DNA molecule is increased.
Molecular Dynamics (MD) simulations of methylated DNA/MBD complex through a nanopore.
Temporal MD snapshots showing translocation of 63 bp dsDNA with: (a) 3 bound MBD proteins through a 12 nm pore, (b) 3 bound MBD proteins through a 10 nm pore, (c) 1 bound MBD protein through a 9 nm pore. As pore size is reduced, hydrophobic interactions between the complex and the pore begin to dominate and can arrest the transport of the molecule through the pore. (d) Center of mass of the complex is shown distance vs. time. Smaller pore sizes can result in the trapping of the complex in the pore.
Discussion
This paper presents a new solid-state nanopore-based direct electrical analysis technique for detecting unmethylated and methylated DNA at the single molecule level. Using MBD-1x as a methylation specific label, the methylation status of nucleotide sequences corresponding to the promoter of DLX1, a potential epigenetic biomarker for cancer, could be rapidly determined without the need for bisulfite conversion, sequencing or fluorescent tags. Notably, the translocation of the mDLX1-protein complex versus uDLX1 induced a ~3 fold signal enhancement in the pore blockage current, enabling the electrical detection of a single methylated CpG dinucleotide-protein complex with high fidelity. The number of methylation sites per molecule could also be coarsely determined using this approach based on the number of bound MDB-1x proteins, characterized by distinct timescales in the corresponding translocation time histograms. While further studies are needed to determine the ultimate spatial resolution of this technique, these findings could find initial application in low-resolution gene based methylation analysis and the mapping of methylated CpG islands in the promoter sequences of various genes, essential to transcriptional repression and gene silencing3. Extending this technique to high resolution epigenetic mapping will require further improvements to the nanopore architecture. Nanopores used in these studies were 20 nm-thick in length (equivalent to ~60 bps of dsDNA) and thus multiple bound proteins contributed to the measured ionic current (Supplementary Figure s9). By reducing pore thickness to below the size of an individual protein, for example by using monolayer thick graphene nanopores53 (thickness of ~0.34 nm), it may be possible to accurately quantify and spatially map the location of individual MBD-1x proteins on a target DNA molecule. This should be feasible as the translocation of DNA-protein complexes through graphene nanopores has already been demonstrated54. Such a technology could ultimately find application in clinical settings. Cancer-specific methylated DNA from most tumor types are known to be present in biopsy specimens and in patient serum at very low concentrations. A rapid, accurate and amplification free assay to detect these biomarkers from minute sample volumes could prove invaluable in the early detection of disease, monitoring disease progression and prognosis. With continued development, solid-state nanopores could meet this unmet technological and clinical need.
Methods
Nanopore electrical measurements
Single nanopores of various diameters were sculpted using a JEOL 2010F field emission gun transmission electron microscope in 20 nm thick, low stress SiN membranes with window sizes of 50 × 50 μm2, supported on a silicon chip. Following pore formation, nanopore chips were cleaned in Piranha solution (two parts 95% H2SO4 and one part of 30% of H2O2) for 10 min and thoroughly rinsed with DI H2O. The chip was then sandwiched in a custom acrylic holder with the nanopore forming the only electrical path for ions between the two reservoirs. The recording solution for both sides was prepared with desired concentrations of KCl at pH 8.0 with 10 mM Tris·HCl and 1 mM EDTA. Ag/AgCl electrodes were immersed in the two reservoirs and an Axopatch 200B was used for applying potentials and measuring currents at a bandwidth of 10 kHz. Data was recorded at a sampling frequency of 100 kHz using a Digidata 1440A data acquisition system. Instrumental control and data analysis was performed using Clampex 10.2. All nanopore experiments were performed in a dark, double Faraday cage on an anti-vibration table at room temperature (22 ± 2°C).
DNA Preparation, Purification and Methylation
The 827 bp DNA fragment used was generated by conventional PCR of human genomic DNA (G304A, Promega, Madison, WI) and includes a region of the DLX1 gene (Homo sapiens distal-less homeobox). The region includes a nontranscribed area adjacent to a CpG island, the 5′ untranslated region (UTR), the complete first exon, part of the first intron and 36 potential CpG sites. The PCR primer sequences are; forward: gaccaatccccagtgattatgcaagac, reverse: ctcaatttgcaactatccagccaagg. The PCR product was purified using Qiaquick PCR purification kit (Qiagen, Inc., Valencia, CA). 50 μg of DNA was methylated in 10 ml using 500 U CpG Methyltransferase M.SssI, New England Biolabs (Ipswich, MA) #M0226M and 160 μM s-adenosyl-methionine (SAM) according to manufacturer's instructions. 33 μg of unmethylated control DNA was treated in the same manner except that no M.SssI was included in the reaction. Reactions were carried out at 37°C for 4 hours, then fresh SAM was added again to 160 μM (320 μM total) and incubated for another 4 hours. DNA was precipitated with ethanol and agarose gel purification was performed using Qiaquick kit with gel extraction protocol. Efficiency of methylation was shown to be high by nearly complete protection from Hha1 (a methylation sensitive enzyme) restriction digestion. There are 4 Hha1 restriction sites in this 827 bp fragment.
MBD-1x protein purification
BL21DE3pLysS E. coli that had been transformed with a bacterial expression vector encoding his-tagged MBD-1x (Kindly provided by Dr. Adrian Bird, University of Edinburgh) was exposed to 1 mM IPTG and incubated on an orbital shaker at 37°C for 3 hours. Bacteria was then chilled on ice, centrifuged at 5000 × g for 5 minutes at 4°C and subjected to 3 freeze/thaw cycles. Lysis buffer (50 mM NaH2PO4, 300 mM NaCl, 10 mM imidazole, pH 8.0) was added and the bacterial lysate was sonicated and spun at 10,000 × g for 40 minutes at 4°C. The cleared lysate was added to a column packed with nickel-NTA agarose resin (Quiagen, Valencia, CA) on an Econo Protein Purification System (BioRad, Hercules, CA) and incubated for 2 hours to allow the his-tagged protein to bind to the nickel column. Guanadinium hydrochloride (5.5 M) was added to the column to denature the protein and a linear guanadinium hydrochloride gradient (5.5–0 M) was used to refold the protein. This renaturation step was critical for MBD-1x activity. The refolded MBD-1x was eluted with increasing concentrations of imidazole (10–250 mM) in elution buffer (50 mM NaH2PO4, 300 mM NaCl, pH 8.0). Protein purity was assessed with Coomassie-stained gels and Western blot analysis using an anti-His antibody (SC-803, Santa Cruz Biotechnology, Santa Cruz, CA).
Gel shift assays
The 827 bp uDLX1 and mDLX1 DNA was end labeled with γ[31P]ATP and T4 polynucleotide kinase (New England Biolabs, Ipswich, MA) and the radiolabeled DNA was separated from free 31P using Quick Spin Columns (Roche Diagnostics Corporation, Indianapolis, IN). The indicated amounts of purified MBD-1x were added to binding buffer (15 mM Tris pH 7.5, 80 mM KCl, 0.4 mM dithiothreitol, 0.2 mM EDTA, 1 ug poly[deoxyinosine/deoxycytosine], 10% glycerol) and incubated for 15 minutes at room temperature. Radiolabeled DNA was added and incubated for 25 minutes at room temperature in a final volume of 20 μl. Samples were fractionated on a low-ionic strength polyacrylamide gel at 4°C with buffer recirculation as previously described55. Bands were visualized using autoradiography.
Molecular dynamics simulation - atomic model
The atomic model of silicon nitride membrane was constructed as described previously49. The thickness of the membrane is 20 nm. A symmetric double-conical pore was produced by removing atoms from the silicon nitride membrane with the diameter of the pore corresponds to experiment (9 nm, 10 nm and 12 nm). Atomic coordinates of mDNA-MBD complex were taken from the NMR structure of the methyl binding domain of MBD1 complexed with mDNA (Protein Data Bank entry code 1IG446). Three mDNA-MBD complex were linked together to generate a long mDNA binding with three MBD proteins, see Figure 4. The sequence of DNA is: 5′-TATCmCGGATACGTATCCGGTATCmCGGATACGTATCCGGATATATCmCGGATACGTATCCGGATA-3′. The specific binding sites (mCG) of mDNA are marked in red. The topology file of DNA and protein along with the missing hydrogen atoms was generated using the psfgen plug-in of VMD56. mDNA-MBD complex was placed in front of the pore and was solvated in a water box with 0.6 M KCl added. The final systems include ~1.1 million atoms. Simulations were performed using the program NAMD 2.8 with the CHARMM27 force field for DNA57, the CHARMM22 force field for proteins with CMAP corrections58,59 and the TIP3P water model60. Periodic boundary condition was employed. The integration time step used was 1 fs with particle-mesh Ewald (PME) full electrostatics with grid density of 1/Å3. Van der Waals energies were calculated using a 12 Å cutoff. A Langevin thermostat was assumed to maintain constant temperature at 295 K61. Each system was energy-minimized for 30,000 steps and then equilibrated for 2 ns under NPT ensemble condition to achieve a constant volume61,62. Production simulations were carried out by applying an electric field along the z-direction (perpendicular to the membrane). The applied voltage is 0.6 V as employed in experiments.
References
Robertson, K. D. DNA methylation and human disease. Nat Rev Genet 6, 597–610 (2005).
Rountree, M. R., Bachman, K. E., Herman, J. G. & Baylin, S. B. DNA methylation, chromatin inheritance and cancer. Oncogene 20, 3156–3165 (2001).
Das, P. M. & Singal, R. DNA methylation and cancer. J. Clin. Oncol. 22, 4632–4642 (2004).
Laird, P. W. The power and the promise of DNA methylation markers. Nat. Rev. Cancer 3, 253–266 (2003).
Laird, P. W. & Jaenisch, R. The role of DNA methylation in cancer genetics and epigenetics. Annu. Rev. Genet. 30, 441–464 (1996).
Strathdee, G. & Brown, R. Aberrant DNA methylation in cancer: potential clinical interventions. Expert Reviews in Molecular Medicine 4, 1–17 (2002).
Vanaja, D. K. et al. Hypermethylation of genes for diagnosis and risk stratification of prostate cancer. Cancer Invest. 27, 549–560 (2009).
Esteller, M., Corn, P. G., Baylin, S. B. & Herman, J. G. A gene hypermethylation profile of human cancer. Cancer Res. 61, 3225–3229 (2001).
Lee, W. H., Isaacs, W. B., Bova, G. S. & Nelson, W. G. CG island methylation changes near the GSTP1 gene in prostatic carcinoma cells detected using the polymerase chain reaction: A new prostate cancer biomarker. Cancer Epidemiology Biomarkers and Prevention 6, 443–450 (1997).
Murrell, A., Rakyan, V. K. & Beck, S. From genome to epigenome. Hum. Mol. Genet. 14, R3–10 (2005).
Grunau, C., Clark, S. J. & Rosenthal, A. Bisulfite genomic sequencing: systematic investigation of critical experimental parameters. Nucleic Acids Res. 29, e65 (2001).
Trinh, B. N., Long, T. I. & Laird, P. W. DNA methylation analysis by methyLight technology. Methods 25, 456–462 (2001).
Eads, C. A. et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 28, e32 (2000).
Chargari, C., Campana, F., Beuzeboc, P., Zefkili, S. & Kirova, Y. M. Preliminary experience of helical tomotherapy for locally advanced pancreatic cancer. World journal of gastroenterology : WJG 15, 4444–4445 (2009).
Sandoval, J. et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. epigenetics 6, 692–702 (2011).
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nat Nano 6, 615–624 (2011).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proceedings of the National Academy of Sciences of the United States of America 93, 13770–13773 (1996).
Kim, M. J., Wanunu, M. & Bell, D. C. A. Meller Rapid fabrication of uniformly sized nanopores and nanopore arrays for parallel DNA analysis. Advanced Materials 18, 3149–3153 (2006).
Venkatesan, B. M. et al. Highly sensitive, mechanically stable nanopore sensors for DNA analysis. Advanced Materials 21, 2771–2776 (2009).
Dekker, C. Solid-state nanopores. Nature Nanotechnology 2, 209–215 (2007).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nature Biotechnology 26, 1146 (2008).
Kasianowicz, J. J., Robertson, J. W. F., Chan, E. R., Reiner, J. E. & Stanford, V. M. Nanoscopic porous sensors. Annu Rev Anal Chem 1, 737–766 (2008).
Reiner, J. E. et al. Disease detection and management via single nanopore-based sensors. Chem Rev 112, 6431–6451 (2012).
Bayley, H. & Jayasinghe, L. Functional engineered channels and pores - (Review). Mol Membr Biol 21, 209–220 (2004).
Gu, L. Q. & Shim, J. W. Single molecule sensing by nanopores and nanopore devices. Analyst 135, 441–451 (2010).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem Soc Rev 38, 2360–2384 (2009).
Ma, L. & Cockroft, S. L. Biological nanopores for single-molecule biophysics. Chembiochem 11, 25–34 (2010).
Movileanu, L. Interrogating single proteins through nanopores: challenges and opportunities. Trends Biotechnol 27, 333–341 (2009).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nature biotechnology 26, 1146–1153 (2008).
Olasagasti, F. et al. Replication of individual DNA molecules under electronic control using a protein nanopore. Nat Nanotechnol 5, 798–806 (2010).
Cipriany, B. R. et al. Single molecule epigenetic analysis in a nanofluidic channel. Anal Chem 82, 2480–2487 (2010).
Cipriany, B. R. et al. Real-time analysis and selection of methylated DNA by fluorescence-activated single molecule sorting in a nanofluidic channel. Proceedings of the National Academy of Sciences of the United States of America 109, 8477–8482 (2012).
Clarke, J. et al. Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4, 265–270 (2009).
Mirsaidov, U. et al. Nanoelectromechanics of methylated DNA in a synthetic nanopore. Biophysical Journal 96, L32–L34 (2009).
Wanunu, M. et al. Discrimination of methylcytosine from hydroxymethylcytosine in DNA molecules. J. Am. Chem. Soc. 133, 486–492 (2010).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Petryniak, M. A., Potter, G. B., Rowitch, D. H. & Rubenstein, J. L. Dlx1 and Dlx2 control neuronal versus oligodendroglial cell fate acquisition in the developing forebrain. Neuron 55, 417–433 (2007).
Tong, W. G. et al. Genome-wide DNA methylation profiling of chronic lymphocytic leukemia allows identification of epigenetically repressed molecular pathways with clinical impact. Epigenetics : official journal of the DNA Methylation Society 5, 499–508 (2010).
Wu, X. et al. CpG island hypermethylation in human astrocytomas. Cancer Res 70, 2718–2727 (2010).
Selamat, S. A. et al. Genome-scale analysis of DNA methylation in lung adenocarcinoma and integration with mRNA expression. Genome Res 22, 1197–1211 (2012).
Wanunu, M., Sutin, J., McNally, B., Chow, A. & Meller, A. DNA translocation governed by interactions with solid-state nanopores. Biophys J 95, 4716–4725 (2008).
Venkatesan, B. M., Shah, A. B., Zuo, J. M. & Bashir, R. DNA sensing using nanocrystalline surface-enhanced Al2O3 nanopore sensors. Adv Funct Mater 20, 1266–1275 (2010).
Roloff, T., Ropers, H. H. & Nuber, U. Comparative study of methyl-CpG-binding domain proteins. BMC Genomics 4, 1 (2003).
Yu, Y. N. et al. Direct DNA methylation profiling using methyl binding domain proteins. Anal Chem 82, 5012–5019 (2010).
Drew, H. R. et al. Structure of a B-DNA dodecamer - conformation and dynamics. 1. P Natl Acad Sci-Biol 78, 2179–2183 (1981).
Ohki, I. et al. Solution structure of the methyl-CpG binding domain of human MBD1 in complex with methylated DNA. Cell 105, 487–497 (2001).
Nan, X., Meehan, R. R. & Bird, A. Dissection of the methyl-CpG binding domain from the chromosomal protein MeCP2. Nucleic Acids Res. 21, 4886–4892 (1993).
Eid J., et al. (ed. USPTO) (Pacific Biosciences of California, Inc., USA; 2010).
Aksimentiev, A., Heng, J. B., Timp, G. & Schulten, K. Microscopic kinetics of DNA translocation through synthetic nanopores. Biophysical Journal 87, 2086–2097 (2004).
Rincon-Restrepo, M., Mikhailova, E., Bayley, H. & Maglia, G. Controlled translocation of individual DNA molecules through protein nanopores with engineered molecular brakes. Nano Letters 11, 746–750 (2011).
Jorgensen, H. F., Adie, K., Chaubert, P. & Bird, A. P. Engineering a high-affinity methyl-CpG-binding protein. Nucleic Acids Res 34 (2006).
Hoogerheide, D. P., Garaj, S. & Golovchenko, J. A. Probing surface charge fluctuations with solid-State nanopores. Phys. Rev. Lett. 102, 256804 (2009).
Garaj, S. et al. Graphene as a subnanometre trans-electrode membrane. Nature 467, 190–193 (2010).
Venkatesan, B. M. et al. Stacked graphene-Al2O3 nanopore sensors for sensitive detection of DNA and DNA–protein complexes. ACS Nano 6, 441–450 (2011).
Loven, M. A., Wood, J. R. & Nardulli, A. M. Interaction of estrogen receptors alpha and beta with estrogen response elements. Mol Cell Endocrinol 181, 151–163 (2001).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J Mol Graph Model 14, 33–38 (1996).
Foloppe, N. & MacKerell, A. D. All-atom empirical force field for nucleic acids: I. Parameter optimization based on small molecule and condensed phase macromolecular target data. J Comput Chem 21, 86–104 (2000).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102, 3586–3616 (1998).
Mackerell, A. D., Feig, M. & Brooks, C. L. Extending the treatment of backbone energetics in protein force fields: Limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comput Chem 25, 1400–1415 (2004).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J Chem Phys 79, 926–935 (1983).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant-pressure molecular-dynamics algorithms. J Chem Phys 101, 4177–4189 (1994).
Feller, S. E., Zhang, Y. H., Pastor, R. W. & Brooks, B. R. Constant-pressure molecular-dynamics simulation - the langevin piston method. J Chem Phys 103, 4613–4621 (1995).
Acknowledgements
The authors would like to acknowledge support from the National Institutes of Health (R21 CA155863 to R.B. and NIDDK R01 053884 to A.M.N.). B.M.V. was a trainee supported by the Midwestern Cancer Nanotechnology Training Center (NIH-NCI R25 CA154015) at UIUC. J.S. was supported by NIH R21 CA155863. X.Z., C.S. and K.S. are grateful for the National Institutes of Health grant P41-RR005969. The authors would also like to acknowledge financial support from the Mayo Clinic/University of Illinois Strategic Alliance for Technology-Based Healthcare (http://mayoillinois.org/). The authors also gladly acknowledge supercomputer time provided by Extreme Science and Engineering Discovery Environment (XSEDE) MCA93S028 and the parallel computing resource Taub cluster provided by the Computational Science and Engineering Program at the University of Illinois. And finally, the authors also thank Drs. Adrian Bird for the MBD-1x expression vector, previously referred to as 1xMBD and Prof. Paul Soloway at (Cornell University) for technical advice on purification of the MBD-1x protein providing the coding sequence for MBD-1x.
Author information
Authors and Affiliations
Contributions
B.M.V., G.V. and R.B. developed the initial concept. J.S., B.M.V., A.M.N., F.K., G.V. and R.B. designed the experiments. J.S. performed all nanopore experiments and analyzed the data. G.I.H. and A.M.N. performed protein preparations and gel shift assays. J.M.M. prepared all DNA samples. X.Z., C.S. and K.S. performed the MD simulations. J.S, B.M.V. and R.B co-wrote the paper. R.B. supervised the study. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Interaction between complex and 9nm pore
Supplementary Information
Interaction between complex and 10nm pore
Supplementary Information
Interaction between complex and 12nm pore
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Shim, J., Humphreys, G., Venkatesan, B. et al. Detection and Quantification of Methylation in DNA using Solid-State Nanopores. Sci Rep 3, 1389 (2013). https://doi.org/10.1038/srep01389
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01389
This article is cited by
-
A novel shaped-controlled fabrication of nanopore and its applications in quantum electronics
Scientific Reports (2019)
-
Solid-state nanopore fabrication in LiCl by controlled dielectric breakdown
Biomedical Microdevices (2018)
-
Tumor and serum DNA methylation in women receiving preoperative chemotherapy with or without vorinostat in TBCRC008
Breast Cancer Research and Treatment (2018)
-
Identification of Methylation-Driven, Differentially Expressed STXBP6 as a Novel Biomarker in Lung Adenocarcinoma
Scientific Reports (2017)
-
Detection and mapping of DNA methylation with 2D material nanopores
npj 2D Materials and Applications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.