Abstract
Systemic inflammatory response syndrome (SIRS) occurs in a range of infectious and non-infectious disease processes. Toll-like receptors (TLRs) initiate such responses. We have shown that parenchymal cell TLR4 activation drives LPS-induced systemic inflammation; SIRS does not develop in mice lacking TLR4 expression on parenchymal cells. The parenchymal cell types whose TLR4 activation directs this process have not been identified. Employing a bone marrow transplant model to compartmentalize TLR4 signaling, we characterized blood neutrophil and cytokine responses, NF-κB1 activation and Tnf-α, Il6 and Ccl2 induction in several organs (spleen, aorta, liver, lung) near the time of LPS-induced symptom onset. Aorta, liver and lung gene responses corresponded with both LPS-induced symptom onset patterns and plasma cytokine/chemokine levels. Parenchymal cells in aorta, liver and lung bearing TLR4 responded to LPS with chemokine generation and were associated with increased plasma chemokine levels. We propose that parenchymal cells direct SIRS in response to LPS.
Similar content being viewed by others
Introduction
Systemic inflammatory response syndrome (SIRS) is a major cause of morbidity and mortality in the United States. The etiology of SIRS is multifactorial, with trauma, acute localized inflammation and systemic infection being of major importance1,2,3,4. Classical experimental studies have described the clinical and pathologic features associated with the onset and progression of SIRS in response to lipopolysaccharide (LPS), an endotoxin produced by Gram-negative bacteria5. The discovery of Toll-like receptors (TLR4) and of TLR4 as a key receptor for LPS has facilitated the development of studies to provide a mechanistic basis for the systemic effects of LPS6,7.
LPS binds LPS-binding protein (LBP) and engages TLR48. TLR4 associates with MD-2 (lymphocyte antigen 96) and CD14 to form an active signaling complex9,10,11,12,13,14. TLR4 ligation triggers two distinct signaling pathways – a MyD88-dependent pathway leading to activation of NF-κB and a MyD88-independent pathway leading to the production of type 1 interferons15,16. NF-κB is a well-recognized transcription factor that directs the production of pro-inflammatory chemokines8. Although many studies of SIRS have focused on TLR4 activation by circulating and infiltrating inflammatory cells, in vitro studies performed by us and others have demonstrated that parenchymal cells also respond to LPS with the generation of pro-inflammatory chemokines and may thereby contribute to the development and progression of SIRS17,18,19,20,21,22,23.
Given that both circulating inflammatory cells and resident parenchymal cells are capable of responding to LPS with chemokine generation, our initial studies sought to determine whether TLR4 signaling by bone marrow-derived inflammatory cells or by parenchymal cells contributed to the manifestations of SIRS24. We found that TLR4 expression by parenchymal cells was necessary and sufficient to produce mortality in mice administered a lethal dose of LPS24. Manifestations of SIRS appeared between 2 and 8 hours following LPS administration only in mice expressing TLR4 on parenchymal cells. Surprisingly, plasma chemokine levels – with the notable exception of CCL2 – did not predict the development of SIRS and mortality24,25.
Based on these considerations, we sought to identify potential organ/tissue source(s) of proinflammatory chemokine generation by parenchymal cells following LPS administration. As a first step, we compared chemokine generation in the spleen (an organ largely composed of bone marrow-derived inflammatory/immune cells), the lung and liver (organs composed of both parenchymal cells and resident bone marrow-derived macrophages) and aorta (an organ composed primarily of parenchymal cells). We employed mice bearing a novel NF-κB promoter-driven EGFP reporter gene, which facilitated studies to directly assay NF-κB activation in tissues harvested from mice following LPS exposure. Based on our previous experience with this model, we chose a time point – 4 hours – at which mice with parenchymal TLR4 expression were starting to develop manifestations of SIRS but before animals showed signs of multiorgan failure.
Results
Depletion of TLR4 on parenchymal cells reduces symptoms of LPS-induced systemic inflammation
We sought to define plasma and tissue chemokine levels at a time when clinical manifestations of SIRS first begin to appear. In our previous study, we employed a classical clinical definition of SIRS – ataxia, weakness, loss of activity as assessed by an experienced animal technician blinded to treatment group – to determine that the median time of LPS-induced symptom onset was ~4 hours in TLR4+/+ mice5,24. We therefore used this time point for analysis of plasma- and cell-derived chemokine expression in the studies described here.
By 4 hours post-injection, 60% of TLR4+/+ mice (TLR4+/+ mice transplanted with TLR4+/+ bone marrow) and 30% of Marrow TLR4−/− mice (TLR4+/+ mice transplanted with TLR4−/− bone marrow) given LPS had signs of LPS-induced symptom onset, as defined above, but none of the Parenchymal TLR4−/− (TLR4−/− mice transplanted with TLR4+/+ bone marrow) or TLR4−/− (TLR4−/− mice transplanted with TLR4−/− bone marrow) mice given LPS displayed any symptoms (Table 1). None of the groups became hypothermic (<32°C; data not shown) and weight loss from the time of LPS or PBS injection was similar across all groups, confirming that LPS-induced hypothermia and weight loss occur only late in the systemic inflammatory process. Moreover, all renal, hepatic and pulmonary samples showed no evidence of tissue damage or necrosis at this point in the host systemic inflammatory response (data not shown) confirming the changes we report here are not the consequence of widespread tissue damage.
Expression of TLR4 by parenchymal cells plays an important role in LPS-induced neutrophil activation
We next examined differences in neutrophil responses near the time of symptom onset. Our previous studies with this model demonstrated a marrow-derived TLR4-driven neutropenia prior to symptom onset (1 hour after LPS administration) and a nonspecific TLR4-driven neutrophilia in mice expressing TLR4 in any compartment after SIRS was established (18 hours post-injection)24. We therefore examined blood neutrophil levels both very early in the process (30 minutes after injection) and near the time of symptom onset (4 hours after injection). Only TLR4+/+ mice experienced a significant neutropenia 30 minutes after LPS injection (p = 0.0002; one-way ANOVA; Fig. 1a). At 4 hours, there was a significant neutrophilia in only Marrow TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 1b).
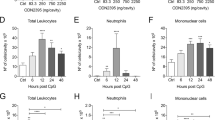
Blood neutrophil responses at 30 minutes post-injection (a) and 4 hours post-injection (b) along with several organ MPO contents at 4 hours post-injection (c-e) for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as mean ± standard error. *: p < 0.05 versus TLR4−/− given LPS; †: p < 0.05 versus Parenchymal TLR4−/− given LPS; ‡: p < 0.05 versus Marrow TLR4−/− given LPS; §: p < 0.01 versus TLR4+/+ given LPS.
Additionally, we assessed lung, kidney and liver neutrophil activity at the time of LPS-induced symptom onset (4 hours post-injection) by measuring individual organ MPO content. Lung MPO content was elevated only in the Marrow TLR4−/− mice given LPS (p = 0.0003; one-way ANOVA; Fig. 1c). In the kidneys, MPO content was elevated in both the TLR4+/+ and Marrow TLR4−/− mice given LPS compared to Parenchymal TLR4−/− and TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 1d). In the liver, MPO content was elevated in Marrow TLR4−/− mice, Parenchymal TLR4−/− mice and TLR4+/+ mice given LPS (p < 0.0001; one-way ANOVA; Fig. 1e). Based on these considerations, we conclude that parenchymal cell expression of TLR4 plays an important role in LPS-induced neutrophil activation.
Plasma cytokine levels do not predict the development of LPS-induced systemic inflammation
Since cytokine/chemokine signaling is thought to be the means by which a systemic inflammatory response is initiated from a local injury26,27, we sought to determine how these molecules differed with TLR4 expression pattern. Plasma cytokine levels were examined very early (30 minutes following LPS injection) and near the time of symptom onset (4 hours post-injection).
At 30 minutes post-injection, all plasma cytokine levels were below the detection threshold of the assay or indistinguishable from levels measured in mice given saline control; plasma FGF2, CSF2, IL1A, IL17A and VEGFA levels remained below the detection limit of the assay near the time of LPS-induced symptom onset (4 hours post-injection, data not shown). IL10 plasma levels were similar amongst all of the groups (data not shown). TNF-α, IL1-β, IL2, IL5 and CCL2 plasma levels were highest in TLR4+/+ mice given LPS, followed by Marrow TLR4−/− mice given LPS and then Parenchymal TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 2a, 2b, 2d, 2g, 2i). For IL4, IL6, CXCL11 and CCL3, TLR4+/+ and Marrow TLR4−/− mice given LPS had higher plasma levels than Parenchymal TLR4−/− and TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 2c, 2h, 2j, 2k). For IFN-γ, IL12 and CXCL10, plasma levels were highest in TLR4+/+ and Parenchymal TLR4−/− mice given LPS, followed by Marrow TLR4−/− mice given LPS and then TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 2e, 2f, 2l).
Plasma levels at 4 hours post-injection of a) TNF-α, b) IL1-β, c) IL6, d) IL2, e) IL12, f) IFN-γ, g) IL5, h) IL4, i) CCL2, j) CXCL11, k) CCL3 and l) CXCL10 for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as mean ± standard error. Dotted line indicates minimal detectable threshold for assay. *: p < 0.01 versus TLR4−/− given LPS; †: p < 0.01 versus Parenchymal TLR4−/− given LPS; ‡: p < 0.01 versus Marrow TLR4−/− given LPS.
Plasma cytokine levels correlated better with parenchymal TLR4 expression than neutrophil responses (Fig. 2a, 2b, 2c, 2g, 2h, 2j, 2k) near the time of symptom onset (4 hours). In our previous study, early (1 hour) plasma levels of TNF-α, IL6 and CCL3 were highest in mice expressing TLR4 in only bone marrow-derived cells (Parenchymal TLR4−/−). However, plasma levels of these chemokines at 4 hours – near the time of symptom onset – were highest in mice with parenchymal TLR4 expression (Fig. 2a, 2c, 2k). Since these mediators can initiate a host systemic inflammatory response, these data suggest parenchymal cells require independent recognition of injury or infection (via TLR4 in this model) before a response to these immune signals is permitted and propagated28,29. Expression of TLR4 on marrow-derived cells then serves to enhance parenchymal TLR4-driven plasma cytokine levels as this process proceeds.
Parenchymal cell expression of pro-inflammatory mediators is associated with the development of LPS-induced systemic inflammation
As an important first step in identifying the parenchymal cell source(s) responsible for the development of SIRS, a series of gene expression studies was performed in tissues harvested 4 hours after LPS administration. Tlr4 mRNA expression was considered to confirm relative amounts of marrow-derived versus parenchymal cells in each organ (using the TLR4 transplant mice given saline) and to detect any signs of LPS-driven Tlr4 induction suggestive of an inflammatory response30. To localize sources of key pro-inflammatory signals involved in the systemic inflammation process, spleen, aorta, liver and lung tissues were analyzed for NF-κB-driven EGFP reporter gene expression along with Tnf-α, Il6 and Ccl2 expression at 4 hours after LPS administration (median time of symptom onset).
Tlr4 expression
In the spleen, Tlr4 expression levels mirrored the bone marrow TLR4 genotype (p = 0.0004 amongst LPS-treated mice; p < 0.0001 amongst vehicle control mice; one-way ANOVA; Fig. 3a). Marrow TLR4−/− mice had roughly 20% of the Tlr4 expression level observed in TLR4+/+ or parenchymal TLR4−/− mice. In the aorta, Tlr4 expression levels mirrored instead the parenchymal cell TLR4 genotype (p < 0.0001 amongst LPS-treated and vehicle control mice; one-way ANOVA; Fig. 3b). Parenchymal TLR4−/− mice had roughly 3% of the aortic Tlr4 expression level observed in TLR4+/+ or Marrow TLR4−/− mice. In the liver, TLR4+/+ and Marrow TLR4−/− mice had similar levels of Tlr4 expression while Parenchymal TLR4−/− mice had roughly half of this expression level (p ≤ 0.0001 amongst LPS-treated and vehicle control mice; one-way ANOVA; Fig. 3c). In the lung, Marrow TLR4−/− and Parenchymal TLR4−/− mice had similar levels of Tlr4 expression and roughly 70% of the level of TLR4+/+ mice (p < 0.0001 amongst vehicle control treated mice; one-way ANOVA; Fig. 3d).
Tlr4 gene expression levels at 4 hours post-injection in a) spleen, b) aorta, c) liver and d) lung for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as expression level (mean ± standard error) relative to TLR4+/+ mice given saline. Dotted line depicts relative gene expression level of one. *: p < 0.001 versus TLR4−/− given LPS; †: p < 0.001 versus TLR4−/− given PBS.
Only liver and lung showed Tlr4 induction in response to LPS. TLR4+/+ mice showed a significant induction of hepatic Tlr4 expression in response to LPS compared to saline control (p = 0.026; t-test); the degree of hepatic Tlr4 induction was not significant in Parenchymal TLR4−/− mice (p = 0.11; t-test). TLR4+/+ mice and Marrow TLR4−/− mice showed an induction of lung Tlr4 expression in response to LPS (p = 0.0001 and p = 0.0037 respectively; t-test).
Basal (saline-treated) TLR4 levels reflected the cellular source of TLR4. For example, the TLR4 expression in the spleen was largely marrow-derived in origin and TLR4 expression in the aorta was largely parenchymal in origin (Fig. 3a, 3b). Liver and lung Tlr4 expression patterns amongst mice given saline suggest a mixture of marrow-derived cells and parenchymal cells. These two organs, though, showed evidence of LPS-driven Tlr4 induction (Fig. 3c, 3d). This induction was largely related to parenchymal cell TLR4 expression indicating a parenchymal cell driven inflammatory response in these two organs consistent with the parenchymal TLR4 dependence of the LPS-induced host systemic inflammatory response.
NF-κB-driven EGFP expression
NF-κB is one major pro-inflammatory transcription factor that is activated when cell surface TLR4 recognizes its ligand8. The mice used for these TLR4 transplant experiments contain an EGFP reporter gene whose promoter consists of three cis NF-κB elements allowing organ EGFP expression to be used as a specific measure of NF-κB activity in vivo31.
Splenic NF-κB activation (as measured by EGFP reporter expression) was only significantly elevated in the TLR4+/+ mice given LPS (p = 0.0009; one-way ANOVA; Fig. 4a). Hepatic EGFP NF-κB reporter expression was highest in TLR4+/+ mice amongst all TLR4 transplant mice given LPS (p = 0.04; one-way ANOVA; Fig. 4c). Aortic and pulmonary NF-κB activation was most elevated in the TLR4+/+ mice given LPS followed by Marrow TLR4−/− mice given LPS (p ≤ 0.0001; one-way ANOVA; Fig. 4b, 4d).
EGFP reporter gene expression levels at 4 hours post-injection in a) spleen, b) aorta, c) liver and d) lung for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as expression level (mean ± standard error) relative to TLR4−/− mice given LPS. Dotted line depicts relative gene expression level of one. *: p < 0.001 versus TLR4−/− given LPS; †: p < 0.05 versus TLR4−/− given LPS.
Thus, aortic and pulmonary NF-κB activity followed a purely parenchymal TLR4 expression pattern while hepatic NF-κB activity showed higher expression levels with parenchymal TLR4 expression than marrow-derived cell TLR4 expression (Fig. 4b, 4c, 4d). Spleen NF-κB activity, however, required both marrow-derived cell TLR4 expression and parenchymal cell TLR4 expression (Fig. 4a).
Tnf-α expression
Splenic Tnf-α expression was induced in both TLR4+/+ and Parenchymal TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 5a). Aortic Tnf-α expression was elevated in all TLR4 transplant mice given LPS except TLR4−/− mice (p < 0.0001; one-way ANOVA; Fig. 5b). Hepatic Tnf-α expression was only significantly elevated in TLR4+/+ mice given LPS (p = 0.015; one-way ANOVA; Fig. 5c) and pulmonary Tnf-α expression was only significantly elevated in Parenchymal TLR4−/− mice given LPS (p < 0.0001; one-way ANOVA; Fig. 5d).
Tnf-α gene expression levels at 4 hours post-injection in a) spleen, b) aorta, c) liver and d) lung for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as expression level (mean ± standard error) relative to TLR4−/− mice given LPS. Dotted line depicts relative gene expression level of one. *: p < 0.001 versus TLR4−/− given LPS; †: p < 0.05 versus TLR4−/− given LPS.
Unexpectedly, there was a wide range of Tnf-α expression levels across these four organs but these data consistently showed a clear disconnect from both parenchymal cell TLR4 expression (which drives LPS-induced systemic inflammation) and organ NF-κB activity (Fig. 4, 5). It is unclear what signals and factors drive each organ's Tnf-α expression since it is believed that NF-κB is the key inducer of murine Tnf-α expression32,33,34.
Il6 and Ccl2 expression
Splenic Il6 levels were undetectable amongst all TLR4 transplant mice given vehicle control and amongst TLR4−/− mice given LPS. Splenic Il6 expression levels were significantly elevated in only TLR4+/+ mice given LPS (p = 0.0005; one-way ANOVA; Fig. 6a). Aortic, hepatic and pulmonary IL6 levels were induced in TLR4+/+ and Marrow TLR4−/− mice given LPS (p ≤ 0.0001; one-way ANOVA; Fig. 6b, 6c, 6d).
Il6 gene expression levels at 4 hours post-injection in a) spleen, b) aorta, c) liver and d) lung for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as expression level (mean ± standard error) relative to Parenchymal TLR4−/− mice given LPS (spleen) or TLR4−/− mice given LPS (aorta, liver, lung). Dotted line depicts relative gene expression level of one. *: p < 0.001 versus TLR4−/− given LPS; †: p < 0.05 versus TLR4−/− given LPS.
Splenic Ccl2 expression levels were highest in TLR4+/+ mice given LPS with lower levels in Marrow TLR4−/− and Parenchymal TLR4−/− mice given LPS respectively (p < 0.0001; one-way ANOVA; Fig. 7a). Aortic, hepatic and pulmonary Ccl2 expression levels were only elevated in TLR4+/+ and Marrow TLR4−/− mice given LPS (p < 0.001; one-way ANOVA; Fig. 7b, 7c, 7d).
Ccl2 gene expression levels at 4 hours post-injection in a) spleen, b) aorta, c) liver and d) lung for four groups of TLR4 transplant mice given LPS (black) or saline (white).
15 TLR4+/+, 15 Marrow TLR4−/−, 15 Parenchymal TLR4−/− and 15 TLR4−/− mice were used for this experiment. Data represented as expression level (mean ± standard error) relative to TLR4−/− mice given LPS. Dotted line depicts relative gene expression level of one. *: p < 0.001 versus TLR4−/− given LPS; †: p < 0.05 versus TLR4−/− given LPS.
Both Il6 and Ccl2 contain κB motifs in their promoter sequences, so it is not surprising that Il6 and Ccl2 expression levels roughly mirrored NF-κB activity levels in each of the organs tested (Fig. 4, 6, 7)35,36,37,38. Both Il6 and Ccl2 expression in aorta, liver and lung were related to parenchymal TLR4 expression and thus also mirrored the pattern of IL6 and CCL2 plasma levels observed at 4 hours post-injection (Fig. 2c, 2i, 6, 7). In total, these data suggest that aorta, liver and lung tissues may be sources of these immune signals in the plasma and support their cellular components as key players in the host systemic inflammatory response.
Discussion
Administration of LPS is a well-established model of SIRS caused by systemic infection by gram negative bacteria5. Our current study, in which blood neutrophil levels are assessed at 30 minutes and at 4 hours, provides a more comprehensive picture of changes in blood neutrophil levels with time following LPS administration. In TLR4+/+ mice (TLR4+/+ mice transplanted with TLR4+/+ bone marrow), LPS administration produces a transient neutropenia, followed by a neutrophilia at late time points, after the onset of SIRS24. Neutrophil levels do not significantly change with LPS administration in TLR4−/− mice, which lack TLR4 expression on both parenchymal cells and bone marrow-derived cells. Marrow TLR4−/− mice do not develop neutropenia, but develop neutrophilia at 4 hours following LPS administration (Fig. 1b), which persists throughout the development of SIRS24. The development of neutropenia appears to be delayed in Parenchymal TLR4−/− mice, with no neutropenia at 30 minutes (Fig. 1a) but with neutropenia at one hour24 following LPS administration.
Lung myeloperoxidase (MPO) content was elevated in Marrow TLR4−/− mice but not TLR4+/+ mice 4 hours after LPS administration. This difference may be related to the fact that marrow TLR4−/− but not TLR4+/+ mice exhibit neutrophilia 4 hours after LPS administration (Fig. 1 b, c). We propose that mice with parenchymal cell expression of TLR4 (Marrow TLR4−/− mice) are capable of activating neutrophils, even if the neutrophils do not express TLR439. In the kidney, elevated MPO content was observed in TLR4+/+ and Marrow TLR4−/− mice, indicating that parenchymal cell expression of TLR4 is required to activate infiltrating neutrophils in the kidney. Finally, MPO content in the liver was elevated in TLR4+/+, Marrow TLR4−/− and Parenchymal TLR4−/− mice, indicating that TLR4 expression in either bone marrow-derived or parenchymal cells is sufficient to activate neutrophils in the liver. These studies demonstrate that expression of TLR4 by parenchymal cells is sufficient to drive LPS-mediated neutrophil activation.
At 4 hours after LPS administration, plasma chemokine levels, with the exception of plasma IFN-γ levels, reflected parenchymal cell expression of TLR4 (TLR4+/+ and Marrow TLR4−/−). No significant induction of any chemokine was observed following LPS administration to TLR4−/− mice. Significant induction of TNF-α, IL1β, IL2, IL12, IFNγ, IL5, CCL2, CXCL11 and CXCL10 levels were observed in Parenchymal TLR4−/− mice at 4 hours after LPS administration. Based on these considerations, we conclude that, under these experimental conditions, plasma chemokine levels may be significantly elevated in mice who do not develop manifestations of SIRS and may thereby be poor predictors of outcome. This model may provide the basis for mechanistic studies underlying the clinical observation that chemokine levels do not always correlate with SIRS and mortality in humans with sepsis40.
Analysis of TLR4 expression in various organs provided several patterns of TLR4 expression. In the spleen, TLR4 expression mirrored the bone marrow phenotype, with highest levels in TLR4+/+ and parenchymal TLR4−/− mice. Conversely, in the aorta, TLR4 expression mirrored the parenchymal cell phenotype, with highest levels observed in TLR4+/+ and Marrow TLR4−/− mice. Using an EGFP assay as a marker of NF-κB activation, we found that highest levels were observed in mice expressing TLR4 in both parenchymal cells and bone marrow-derived cells (TLR4+/+), with lower levels of NF-κB activation observed in aorta and lung of Marrow TLR4−/− mice.
Our studies provide an important first step in the identification of parenchymal cell type(s) responsible for the development and progression of SIRS. Based on our gene expression studies, the spleen does not appear to significantly contribute to the development and progression of SIRS. However, Tlr4 induction, NF-κB activation and Il6/Ccl2 expression was associated with parenchymal cell TLR4 expression in the aorta, liver and lung, suggesting that these organs may be critically involved in the host systemic inflammatory response. Remarkably, TNF-α gene expression did not correlate with parenchymal cell TLR4 expression or NF-κB activity in these organs. This finding suggests that TNF-α expression at 4 hours doe not predict the development of LPS-induced systemic inflammation.
However, our study does have some limitations that merit discussion. As with our previous study, our disease definition for LPS-induced systemic inflammation relies on clinical observation instead of continuous measurement of criteria on which the human definition of SIRS is based (i.e., temperature, heart rate, respiratory rate, white blood cell count)41,42. We did, however, measure neutrophil responses and hypothermia, which are components of the human SIRS definition. Moreover, this experimental model in which TLR4 activation in parenchymal cells and marrow-derived cells is artificially separated does not reflect the natural state of the host. Thus the applicability of these findings to the host systemic inflammatory response could be argued to be limited. These studies, however, do allow separation of necessary (and potentially causative) components of the host systemic inflammatory response from simply associative (but non-causative) components (e.g., blood neutrophil responses, organ neutrophil activity) – a feat not possible without this separation of TLR4 expression in vivo. Finally, the wider applicability of the LPS-induced systemic inflammatory response model for human disease remains in question given that this model is an intense pro-inflammatory disease model that does not recapitulate complicated, multi-faceted disease processes like human sepsis40,43. As a study focused, though, on the components and regulation of a host systemic inflammatory response, the use of a single inflammatory mediator recognized by a single host receptor allows us to parse out these components in the absence of other competing factors.
The next logical step in these studies is to identify the specific parenchymal cell type(s) responsible for the development of SIRS. One common parenchymal cell type in the aorta, liver and lung – organs likely to be important in the development and progression of SIRS – is endothelium. Endothelial cells comprise the intimal layer of the aorta and make up 3% and 30% of parenchymal cells in the liver and lung respectively41,42. Endothelial cells also actively express TLR4 and have been shown to respond to inflammatory stimuli like LPS in vitro44,45,46. Low dose LPS studies in mice whose TLR4 is solely expressed on their endothelial cells demonstrated patterns of P-selectin induction (endothelial cell activation) and neutrophilic sequestration in the lungs similar to those observed in TLR4 wild-type mice supporting a role of endothelial cell TLR4 in the host inflammatory response47. Endothelial cells are also parenchymal cells with direct access to circulating plasma factors like TNF-α, IL6 and CCL3 whose very early plasma levels were ultimately disconnected from development of SIRS and from mortality24. Therefore, endothelial cells may be one cell type playing a key role in also limiting the classical inflammatory response induced by these pro-inflammatory plasma factors early in Parenchymal TLR4−/− mice given LPS.
Based on these considerations, we conclude that neither neutrophil responses nor plasma chemokine levels predict the development of SIRS. As expected, NF-κB activity and TLR4 expression in aorta, liver and lung reflected parenchymal cell expression of TLR4. In liver, lung and aorta, expression of IL-6 and CCL2, but not TNF-α, reflected TLR4 expression by parenchymal cells and predicted the development of SIRS. These data suggest parenchymal cell types present in one or more of these organs (e.g., endothelial cells) could be responsible for the development and progression of SIRS in this model.
Methods
TLR4-positive and TLR4-negative NF-κB-enhanced green fluorescent protein (EGFP) reporter mice
All animal experiments were approved by Mayo Clinic's institutional animal care and use committee. C57BL TLR4-positive NF-κB reporter mice were obtained from the University of North Carolina31. These mice contain an NF-κB-driven EGFP reporter gene on their X chromosome to allow direct, specific measurement of this pro-inflammatory transcription factor's activity. Offspring from these EGFP reporter mice were bred with commercially obtained C57BL TLR4-knockout mice (Jackson Laboratories, Bar Harbor, ME, USA) to create a corresponding C57BL TLR4-knockout NF-κB EGFP reporter mouse colony. All mice used in these experiments were derived from these two colonies.
TLR4 transplant mouse generation
Protocols for TLR4 transplant mouse generation are described fully elsewhere24,48. In brief, recipient mice receive lethal irradiation and donor mouse bone marrow is extracted from excised femurs and tibia. Eight million donor marrow cells suspended in sterile phosphate-balanced saline (PBS) are injected IV into each recipient mouse. Engraftment is complete in 8–10 weeks. Using this procedure, two types of chimeric TLR4 mice – Marrow TLR4−/− (transplantation of TLR4−/− marrow into a TLR4+/+ mouse) and Parenchymal TLR4−/− (transplantation of TLR4+/+ marrow into a TLR4−/− mouse) - were generated along with two types of TLR4 transplant control mice – TLR4+/+ (transplantation of TLR4+/+ marrow into a TLR4+/+ mouse) and TLR4−/− (transplantation of TLR4−/− marrow into a TLR4−/− mouse). Previous experiments using this transplantation procedure between TLR4 wild-type and TLR4 knockout mice demonstrated that >99% circulating leukocytes and tissue macrophages acquire the donor phenotype by the end of the engraftment process48,49,50. All bone marrow engraftments conducted in this study were confirmed at 7 weeks post-transplantation by the presence or absence of ex vivo production of TNF-α by isolated peripheral blood mononuclear cells after an 18 hour exposure to LPS (data not shown).
LPS-induced systemic inflammation
TLR4-dependent systemic inflammation was induced using intraperitoneal injection of ultra-pure E. coli LPS (Invitrogen, San Diego, CA, USA) dissolved in PBS24. Each lot of LPS was standardized for biologic activity using a chromogenic Limulus amebocyte lysate assay (GenScript, Piscataway, NJ, USA) to adjust for any variation between lots. A dose of ~1.5 × 1015 EU/kg was used in these experiments (a dose that achieves systemic inflammation with high mortality in TLR4+/+ mice). As a vehicle control, a matching volume of PBS was injected intraperitoneal.
Onset of a host systemic inflammatory response to this dose of LPS occurs between 2–8 hours post-injection in >80% of TLR4+/+ and Marrow TLR4−/− mice24. Blood was therefore collected at 30 minutes and 4 hours post-injection and organs (spleen, aorta, lung, liver, kidney) were harvested at 4 hours. Signs of murine LPS-induced systemic inflammation (ataxia, weakness, activity and appetite loss) were assessed at 4 hours (just prior to sacrifice)5,24. Additionally, all mice were assessed for hypothermia (<32°C core temperature) via rectal thermometer prior to anesthesia induction for sacrifice and animal weights were measured before LPS/saline injection and again before sacrifice to assess weight change.
Plasma cytokines, blood neutrophil content, tissue histology and myeloperoxidase content
In brief, all blood was collected in 7.5% K2EDTA. Plasma was isolated from each blood sample by centrifugation and stored at −80°C. Plasma cytokine levels were assessed using the Luminex mouse cytokine 20-plex panel on a Luminex 100 machine (Invitrogen, Carlsbad, CA, USA). IL12 and CXCL9 levels were not considered for these experiments as per Invitrogen's assay validation data on EDTA plasma samples. The remaining packed blood cells were stained with neutrophil marker Gr1-PerCP (eBiosciences, San Diego, CA, USA) and processed using a whole blood lysing kit (R & D Systems, Minneapolis, MN, USA). Flow cytometry of these fixed cells was conducted using a BD FACS Calibur cytometer (BD Biosciences, San Jose, CA, USA). White blood cell populations were separated from cell fragments using forward and side scatter gating. Neutrophils were then separated from the lymphocyte/monocyte population based on both forward/side scatter and positive Gr1-PerCP staining. Percentage of neutrophils in blood was calculated as the number of neutrophils divided by total number of gated white blood cells.
Organs were flash frozen upon harvest and stored at −80°C for later analysis. Myeloperoxidase (MPO) content was used as a marker of neutrophil activity and was assessed in frozen lung, kidney and liver samples for each TLR4 transplant mouse using a commercially available murine MPO ELISA kit (HyCult Biotech, Plymouth Meeting, PA, USA). Renal, hepatic and pulmonary samples were also fixed in 10% neutral buffered formalin, dehydrated and embedded in paraffin using standard techniques. Representative sections were stained with hematoxylin & eosin and reviewed for signs of damage or necrosis by a pathologist blinded to mouse and intervention type.
Spleen, aorta, liver and lung gene expression studies
Total RNA was isolated from flash frozen spleens, aortas, livers and lungs using the RNeasy Mini kit with on-column DNase treatment (spleen), the RNeasy Plus Mini kit (liver, lung), or the RNeasy Fibrous Tissue Mini kit (aorta) (Qiagen, Valencia, CA, USA). All extracted RNA samples were analyzed for RNA integrity by Mayo Clinic's Advanced Genomic Technology Center Gene Expression Core to exclude any RNA samples unsuitable for analysis. cDNA was prepared from 500 ng (aorta, liver, lung) or 1000 ng (spleen) total RNA and the iScript cDNA synthesis kits per kit instructions (Bio-Rad, Hercules, CA, USA). SYBR green RT-PCR kits (Roche Applied Science, Indianapolis, IN, USA) and an iQ5 RT-PCR Detection System along with the manufacturer's recommended 96-well plates and films (Bio-Rad, Hercules, CA, USA) were used to conduct all gene expression studies.
Two RNA samples were excluded from use in RT-PCR analysis due to low RNA quality (one lung and one aorta). The average RNA Integrity Number (RIN) for all spleen, aorta, liver and lung total RNA samples used for these gene expression studies was 7.2 (SD 1.0), 8.9 (SD 0.2), 9.7 (SD 0.2) and 9.9 (SD 0.1) respectively. There was no difference in RINs between TLR4 transplant groups or between LPS and saline-treated mice for each organ (data not shown).
All RT-PCR primer pairs used were selected from previous published studies and all primer pairs were validated for RT-PCR prior to use (See Supplementary Table S1 online)51,52,53. All primer pairs underwent standard PCR on spleen and lung cDNA samples from experimental control mice (untransplanted TLR4 wild-type mice given LPS and untransplanted TLR4 knockout mice given PBS) using AmpliTaq Gold with GeneAmp 10x PCR buffer (Life Technologies, Carlsbad, CA, USA). PCR target sequences were then confirmed by Mayo Clinic's Advanced Genomic Technology Center DNA Sequencing Core. These PCR products were isolated to create standard curves for each RT-PCR primer pair (109 – 103 copies/μL) for measuring RT-PCR performance. Standard curve samples were run along with positive and negative control cDNA samples to confirm a single amplicon (melt curve analysis) and measure PCR efficiencies (See Supplementary Table S1 online).
For each organ of interest, Rn18s, Actb, Hprt and Gapdh were considered for use as reference genes. Rn18s was the only target found to be expressed at a constant level across all TLR4 transplant groups and interventions within each organ analyzed and thus was selected as the reference gene for use in these studies.
Statistical analysis
LPS-induced blood neutrophil levels, tissue MPO content and plasma cytokine levels were compared across all four TLR4 transplant groups initially using one-way ANOVA and then Tukey-Kramer Honestly Significant Differences test for all subsequent pair-wise testing to account for multiple comparisons. LPS-induced gene expression changes (Tlr4, EGFP reporter, Tnf-α, Il6, Ccl2) in spleen, aorta liver and lung were expressed relative to reference gene expression (Rn18s) and normalized to an appropriate reference group (Tlr4: TLR4+/+ mice given PBS; EGFP, Tnf-α, Il6, Ccl2: TLR4−/− given LPS if there were detectable expression levels). LPS-induced gene expression was compared across all four TLR4 transplant groups initially using one-way ANOVA and then using Dunnett's test for subsequent comparisons to the experimental negative control group (TLR4−/− mice) to account for multiple comparisons. Organ Tlr4 induction in response to LPS was assessed using t-tests assuming unequal variances by comparing LPS-treated and saline-treated mice within each TLR4 transplant group. A p-value < 0.05 was considered statistically significant.
References
Bhatia, M. Acute pancreatitis as a model of SIRS. Front Biosci 14, 2042–2050 (2009).
Bhatia, M., He, M., Zhang, H. & Moochhala, S. Sepsis as a model of SIRS. Front Biosci 14, 4703–4711 (2009).
Dahiya, P. Burns as a model of SIRS. Front Biosci 14, 4962–4967 (2009).
Robertson, C. M. & Coopersmith, C. M. The systemic inflammatory response syndrome. Microbes Infect 8, 1382–1389, 10.1016/j.micinf.2005.12.016 (2006).
Thomas, L. The physiological disturbances produced by endotoxins. Annu Rev Physiol 16, 467–490, 10.1146/annurev.ph.16.030154.002343 (1954).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
Qureshi, S. T. et al. Endotoxin-tolerant mice have mutations in Toll-like receptor 4 (Tlr4). J Exp Med 189, 615–625 (1999).
Lu, Y. C., Yeh, W. C. & Ohashi, P. S. LPS/TLR4 signal transduction pathway. Cytokine 42, 145–151 (2008).
Suntharalingam, G. et al. Cytokine storm in a phase 1 trial of the anti-CD28 monoclonal antibody TGN1412. N Engl J Med 355, 1018–1028 (2006).
Prince, J. M. et al. Toll-like receptor-4 signaling mediates hepatic injury and systemic inflammation in hemorrhagic shock. J Am Coll Surg 202, 407–417 (2006).
DeMaria, E. J., Pellicane, J. V. & Lee, R. B. Hemorrhagic shock in endotoxin-resistant mice: improved survival unrelated to deficient production of tumor necrosis factor. J Trauma 35, 720–724; discussion 724–725 (1993).
Barsness, K. A. et al. Hemorrhage-induced acute lung injury is TLR-4 dependent. Am J Physiol Regul Integr Comp Physiol 287, R592–599 (2004).
Zhai, Y. et al. Evidence for the Pivotal Role of Endogenous Toll-Like Receptor 4 Ligands in Liver Ischemia and Reperfusion Injury. Transplantation 85, 1016–1022 (2008).
Levy, R. M. et al. Systemic inflammation and remote organ damage following bilateral femur fracture requires Toll-like receptor 4. Am J Physiol Regul Integr Comp Physiol 291, R970–976 (2006).
Kobbe, P. et al. Local Exposure of Bone Components to Injured Soft Tissue Induces Toll-Like-Receptor-4 Dependent Systemic Inflammation with Acute Lung Injury. Shock (2008).
Breslin, J. W., Wu, M. H., Guo, M., Reynoso, R. & Yuan, S. Y. Toll-like receptor 4 contributes to microvascular inflammation and barrier dysfunction in thermal injury. Shock 29, 349–355 (2008).
Grande, J. P., Jones, M. L., Swenson, C. L., Killen, P. D. & Warren, J. S. Lipopolysaccharide induces monocyte chemoattractant protein production by rat mesangial cells. J Lab Clin Med 124, 112–117 (1994).
Cena, J., Lalu, M., Rosenfelt, C. & Schulz, R. Endothelial dependence of matrix metalloproteinase-mediated vascular hyporeactivity caused by lipopolysaccharide. European Journal of Pharmacology 582, 116–122 (2008).
Frank, D., Ahrens, F. & Kramer, T. Cytokine release by porcine livers perfused with lipopolysaccharide or live Salmonella choleraesuis. American Journal of Veterinary Research 57, 472–476 (1996).
Saban, R. et al. Effects of Pasteurella haemolytica leukotoxin and lipopolysaccharide on histamine, prostanoid and leukotriene release by bovine lung parenchyma in vitro. American Journal of Veterinary Research 58, 1227–1231 (1997).
Salzer, W. & McCall, C. Primed stimulation of isolated perfused rabbit lung by endotoxin and platelet activating factor induces enhanced production of thromboxane and lung injury. Journal of Clinical Investigation 85, 1135–1143 (1990).
Takahashi, Y., Negoro, M. & Wakabayashi, I. Decreased modulation by lipopolysaccharide of aortic smooth muscle contractility in streptozotocin-induced hyperglycemic rats. Journal of Cardiovascular Pharmacology 41, 162–170 (2003).
Tran-Thi, T. et al. Production of tumor necrosis factor-alpha, interleukin-1 and interleukin-6 in the perfused rat liver. Eur Cytokine Netw 4, 363–370 (1993).
Juskewitch, J. E. et al. LPS-induced murine systemic inflammation is driven by parenchymal cell activation and exclusively predicted by early MCP-1 plasma levels. The American journal of pathology 180, 32–40, 10.1016/j.ajpath.2011.10.001 (2012).
Leonard, E. J. & Yoshimura, T. Human monocyte chemoattractant protein-1 (MCP-1). Immunol Today 11, 97–101 (1990).
Bone, R. C. Immunologic dissonance: a continuing evolution in our understanding of the systemic inflammatory response syndrome (SIRS) and the multiple organ dysfunction syndrome (MODS). Annals of internal medicine 125, 680–687 (1996).
Bone, R. C. Toward a theory regarding the pathogenesis of the systemic inflammatory response syndrome: what we do and do not know about cytokine regulation. Critical care medicine 24, 163–172 (1996).
Tracey, K. J. et al. Shock and tissue injury induced by recombinant human cachectin. Science 234, 470–474 (1986).
Davatelis, G. et al. Macrophage inflammatory protein-1: a prostaglandin-independent endogenous pyrogen. Science 243, 1066–1068 (1989).
Roger, T. et al. Critical role for Ets, AP-1 and GATA-like transcription factors in regulating mouse Toll-like receptor 4 (Tlr4) gene expression. The Biochemical journal 387, 355–365, 10.1042/BJ20041243 (2005).
Magness, S. T. et al. In vivo pattern of lipopolysaccharide and anti-CD3-induced NF-kappa B activation using a novel gene-targeted enhanced GFP reporter gene mouse. J Immunol 173, 1561–1570 (2004).
Collart, M. A., Baeuerle, P. & Vassalli, P. Regulation of tumor necrosis factor alpha transcription in macrophages: involvement of four kappa B-like motifs and of constitutive and inducible forms of NF-kappa B. Molecular and cellular biology 10, 1498–1506 (1990).
Shakhov, A. N., Collart, M. A., Vassalli, P., Nedospasov, S. A. & Jongeneel, C. V. Kappa B-type enhancers are involved in lipopolysaccharide-mediated transcriptional activation of the tumor necrosis factor alpha gene in primary macrophages. J Exp Med 171, 35–47 (1990).
Drouet, C., Shakhov, A. N. & Jongeneel, C. V. Enhancers and transcription factors controlling the inducibility of the tumor necrosis factor-alpha promoter in primary macrophages. Journal of immunology 147, 1694–1700 (1991).
Dendorfer, U., Oettgen, P. & Libermann, T. A. Multiple regulatory elements in the interleukin-6 gene mediate induction by prostaglandins, cyclic AMP and lipopolysaccharide. Molecular and cellular biology 14, 4443–4454 (1994).
Yeagley, D. & Lang, C. Endotoxin-induced IL-6 promoter activation in skeletal muscle requires an NF-kappaB site. International Journal of Interferon, Cytokine and Mediator Research 2, 9–21 (2010).
Ping, D., Boekhoudt, G. H., Rogers, E. M. & Boss, J. M. Nuclear factor-kappa B p65 mediates the assembly and activation of the TNF-responsive element of the murine monocyte chemoattractant-1 gene. Journal of immunology 162, 727–734 (1999).
Diaz Encarnacion, M. M. et al. n-3 fatty acids block TNF-(alpha)-stimulated MCP-1 expression in rat mesangial cells. American Journal of Physiology - Renal Physiology F1142–F1151, 10.1152/ajprenal.00064.2011 (2011).
Abraham, E., Carmody, A., Shenkar, R. & Arcaroli, J. Neutrophils as early immunologic effectors in hemorrhage- or endotoxemia-induced acute lung injury. American journal of physiology. Lung cellular and molecular physiology 279, L1137–1145 (2000).
Hotchkiss, R. S. & Karl, I. E. The pathophysiology and treatment of sepsis. N Engl J Med 348, 138–150 (2003).
Blouin, A., Bolender, R. P. & Weibel, E. R. Distribution of organelles and membranes between hepatocytes and nonhepatocytes in the rat liver parenchyma. A stereological study. The Journal of cell biology 72, 441–455 (1977).
Crapo, J. D., Barry, B. E., Gehr, P., Bachofen, M. & Weibel, E. R. Cell number and cell characteristics of the normal human lung. Am Rev Respir Dis 126, 332–337 (1982).
Remick, D. G. & Ward, P. A. Evaluation of endotoxin models for the study of sepsis. Shock 24 Suppl 1, 7–11 (2005).
Zhang, F. X. et al. Bacterial lipopolysaccharide activates nuclear factor-kappaB through interleukin-1 signaling mediators in cultured human dermal endothelial cells and mononuclear phagocytes. The Journal of biological chemistry 274, 7611–7614 (1999).
Faure, E. et al. Bacterial lipopolysaccharide activates NF-kappaB through toll-like receptor 4 (TLR-4) in cultured human dermal endothelial cells. Differential expression of TLR-4 and TLR-2 in endothelial cells. The Journal of biological chemistry 275, 11058–11063 (2000).
Faure, E. et al. Bacterial lipopolysaccharide and IFN-gamma induce Toll-like receptor 2 and Toll-like receptor 4 expression in human endothelial cells: role of NF-kappa B activation. Journal of immunology 166, 2018–2024 (2001).
Bone, R. C., Sprung, C. L. & Sibbald, W. J. Definitions for sepsis and organ failure. Critical care medicine 20, 724–726 (1992).
Spangrude, G. J. Assessment of lymphocyte development in radiation bone marrow chimeras. Curr Protoc Immunol Chapter 4, Unit 4 6, doi:10.1002/0471142735.im0406s81 (2008).
Andonegui, G. et al. Endothelium-derived Toll-like receptor-4 is the key molecule in LPS-induced neutrophil sequestration into lungs. J Clin Invest 111, 1011–1020, 10.1172/JCI16510 (2003).
Tavener, S. A. et al. Immune cell Toll-like receptor 4 is required for cardiac myocyte impairment during endotoxemia. Circ Res 95, 700–707, 10.1161/01.RES.0000144175.70140.8c 01.RES.0000144175.70140.8c [pii] (2004).
Shi, H. et al. TLR4 links innate immunity and fatty acid-induced insulin resistance. J Clin Invest 116, 3015–3025, 10.1172/JCI28898 (2006).
Patil, C., Zhu, X., Rossa, C., Jr, Kim, Y. J. & Kirkwood, K. L. p38 MAPK regulates IL-1beta induced IL-6 expression through mRNA stability in osteoblasts. Immunol Invest 33, 213–233 (2004).
Charlton, M. et al. Fast food diet mouse: novel small animal model of NASH with ballooning, progressive fibrosis and high physiological fidelity to the human condition. American journal of physiology. Gastrointestinal and liver physiology 301, G825–834, 10.1152/ajpgi.00145.2011 (2011).
Acknowledgements
We thank Cherish Grabau for her secretarial assistance, Marshall Behrens for his assistance with the Luminex plasma cytokine assays and Kim Butters, Andrea Tilden and Anuradha Krishnan, PhD, for their assistance with developing the quantitative RT-PCR experiments. We would also like to thank Mayo Clinic's Advanced Genomic Technology Center Gene Expression and DNA Sequencing cores for their help with our studies. This work was supported in part by the National Institutes of Health (F30 DK084671, UL1 RR024150, P01 HL85307) and by Mayo Clinic's Department of Laboratory Medicine and Pathology. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
JEJ, JLP, GB and JPG designed the experiments, JEJ, BEK and KLK executed the studies and analysis, JEJ wrote the manuscript and all authors revised and approved of the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Juskewitch, J., Platt, J., Knudsen, B. et al. Disparate roles of marrow- and parenchymal cell-derived TLR4 signaling in murine LPS-induced systemic inflammation. Sci Rep 2, 918 (2012). https://doi.org/10.1038/srep00918
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00918
This article is cited by
-
P38/MAPK contributes to endothelial barrier dysfunction via MAP4 phosphorylation-dependent microtubule disassembly in inflammation-induced acute lung injury
Scientific Reports (2015)
-
Natural small molecule FMHM inhibits lipopolysaccharide-induced inflammatory response by promoting TRAF6 degradation via K48-linked polyubiquitination
Scientific Reports (2015)
-
Chloroquine relieves acute lung injury in rats with acute hemorrhagic necrotizing pancreatitis
Journal of Huazhong University of Science and Technology [Medical Sciences] (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.