Abstract
Plant virus technology, in particular virus-induced gene silencing, is a widely used reverse- and forward-genetics tool in plant functional genomics. However the potential of virus technology to express genes to induce phenotypes or to complement mutants in order to understand the function of plant genes is not well documented. Here we exploit Potato virus X as a tool for virus-induced gene complementation (VIGC). Using VIGC in tomato, we demonstrated that ectopic viral expression of LeMADS-RIN, which encodes a MADS-box transcription factor (TF), resulted in functional complementation of the non-ripening rin mutant phenotype and caused fruits to ripen. Comparative gene expression analysis indicated that LeMADS-RIN up-regulated expression of the SBP-box (SQUAMOSA promoter binding protein-like) gene LeSPL-CNR, but down-regulated the expression of LeHB-1, an HD-Zip homeobox TF gene. Our data support the hypothesis that a transcriptional network may exist among key TFs in the modulation of fruit ripening in tomato.
Similar content being viewed by others
Introduction
In plants, annotations of the dicot Arabidopsis and the monocot rice genome sequences have predicted around 30,000–40,000 genes in the respective genomes1,2. Meanwhile, the high throughput and vast capacity of next-generation sequencing have tremendously increased the scope of whole genome sequences. High-quality, or draft, whole genome sequences of a number of higher plant species such as Arabidopsis1, rice2, maize3, potato4 and tomato5 are already publically available. These data together with bioinformatic predictions, as well as transcriptome and proteome analyses of the expression patterns of thousands of unknown genes, will certainly lead to the identification of a large number of putative genes with no designated biological functions. Forward and reverse genetics are two classical approaches to define gene function6. In practice, both approaches involve mutant analysis that provides a genetic link between genotype and phenotype. In plants, mutants can either be identified in natural populations or induced by chemical mutagenesis. Techniques including physical mutagenesis, T-DNA insertion, transposon integration and RNAi-mediated knockdown are also used to create mutants in order to assign gene function, however these genetics tools are largely restricted to model plants such as Arabidopsis, tobacco or rice6. Definite proof of gene function often involves complementation in a mutant background, which is usually achieved through stable transformation6. However, transformation is time consuming and technically challenging in many crops of economic importance. Thus to elucidate the biological function of genes predicted from whole genome sequences remains an arduous task and there is a need to develop novel functional genomics tools in order to bridge the knowledge gap from gene-to-function6.
Plant virus technology does not require transformation and it has attracted attention because it is an effective tool to delineate gene function in plants. In particular, virus-induced gene silencing (VIGS) has become a rapid reverse- and forward-genetics tool in functional genomics since the early development and refinement of VIGS vectors based on plant viruses including Tobacco mosaic virus, Potato virus X (PVX) and Tobacco rattle virus7,8,9,10. VIGS has now been applied to more than 30 plant species across the angiosperm phylogeny11,12. For instance, we have applied a PVX-based VIGS to inhibit the expression of SBP-box LeSPL-CNR and HD-Zip homeobox LeHB-1 transcription factor (TF) genes in order to reveal their essential roles in the control of fruit ripening and floral organogenesis in tomato13,14. In addition to VIGS, virus expression technology can also be used for high level production of enzymes, green fluorescent protein and foreign proteins of pharmaceutical and industrial value and for studying the biological function of heterologous genes in plants15,16,17,18,19. More recently we have reported the development of a novel virus-based RNA mobility assay for use in elucidating biological relevance of systemic florigen signalling in flower induction20,21, as well as a movement-deficient virus system to dissect intercellular RNA silencing22,23,24,25. However, the potential of virus technology to express endogenous genes to induce phenotypes, or to complement mutants, in order to investigate their functions in plants is not well recognised.
The tomato ripening inhibitor (rin) locus encodes the MADS-box TF LeMADS-RIN, which is induced at the onset of ripening26. The LeMADS-RIN gene resides on chromosome 5 adjacent to MACROCALYX (MC), the tomato APETALA1 orthologue. The rin mutation is caused by a spontaneous deletion of the 3′-end of the LeMADS-RIN gene and approximately 1 kb of the intergenic sequence between LeMADS-RIN and MC. A MADS-RIN/MC transcript is expressed in the rin mutant and the chimeric RNA cannot produce functional LeMADS-RIN and MC proteins. Elegant genetic and molecular analyses confirm that the LeMADS-RIN gene alone is responsible for ripening, providing the first insight into the dedicated fruit-specific transcriptional modulation of ripening26. The tomato LeHB-1, a class-I homeodomain leucine zipper TF protein, is involved in fruit ripening through transcriptional regulation of ACC oxidase gene expression14. Another key player in fruit ripening is the gene at the tomato Colourless non-ripening (Cnr) locus. Cnr is a pleitropic ripening mutant with colourless fruits and an altered texture resulting from loss of cell-to-cell adhesion. The gene underlying this mutation was identified as a SQUAMOSA Promoter Binding Protein (SBP)-box gene (LeSPL-CNR), belonging to the SPL gene family of TFs13. There are several other TFs, which have also been demonstrated to play a role in the control of tomato fruit ripening27, however the regulatory network of these ripening-associated TFs remains to be elucidated. Here we exploited a PVX-based viral expression vector to express LeMADS-RIN for virus-induced gene complementation (VIGC) in tomato rin mutant. We demonstrated that expression of LeMADS-RIN from VIGC vectors enabled functional complementation and caused rin fruits to ripen. Our data also favour a presumption that a regulatory network in which LeMADS-RIN up-regulates LeSPL-CNR whilst down-regulating LeHB-1 may play a role in the modulation of fruit ripening in tomato.
Results
Viral ectopic expression of LeMADS-RIN from VIGC vectors
In order to express LeMADS-RIN in plant leaf tissues and tomato fruits, the coding region for LeMADS-RIN was PCR amplified and cloned into the PVX-based gene expression vector28 to produce PVX/LeMADS-RIN (Fig. 1a). To facilitate detection of the LeMADS-RIN protein we also generated PVX/HisG::LeMADS-RIN in a modified vector PVX/HisGMF1 (Fig. S1), in which the coding sequence for a His6-Gly tag was fused in frame to the 5′ end of the LeMADS-RIN gene. As negative controls, a stop codon was introduced to replace the start codon of LeMADS-RIN and this non-sense mutated gene was cloned into the PVX and PVX/HisGMF1 vectors to produce PVX/mLeMADS-RIN and PVX/HisG::mLeMADS-RIN (Fig. 1a; Fig. S1). The integrity of the cloned genes was confirmed by sequencing.
Ectopic expression of LeMADS-RIN from VIGC vectors in Nicotiana benthamiana.
(a) Construction of VIGC vectors. The PVX vector is shown together with the wild-type LeMADS-RIN and the mutated mLeMADS-RIN (asterisked; Table S1) used to create PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN and PVX/HisG::mLeMADS-RIN vectors. PVX/LeMADS-RIN and PVX/HisG::LeMADS-RIN were designed to express either free LeMADS-RIN, or the LeMADS-RIN protein tagged with six consecutive histidine residues and one glycine residue (His6-Gly), respectively. Viral RNA dependent RNA polymerase (166K), movement proteins (25, 12 and 8K) and coat protein (CP) as well as the His6-Gly tag are indicated. The cloning sites and the position of primers (triangles) used for RT-PCR detections of the PVX::LeMADS-RIN RNA are also indicated. (b-e) Mock-inoculated N. benthamiana (b) remained healthy while plants infected with PVX/LeMADS-RIN (c), PVX/HisG::LeMADS-RIN (d), PVX/HisG::mLeMADS-RIN (e) or PVX/mLeMADS-RIN (data not shown) developed chlorosis and mosaic phenotypes, characteristic of PVX infection. Photographs were taken at 14 days post-inoculation (dpi). (f, g) Western blot detection of the LeMADS-RIN protein. Proteins were extracted from leaves of plants infected with PVX/HisG::LeMADS-RIN (RIN), PVX/HisG::mLeMADS-RIN (mRIN), or from mock-inoculated (mock) plants at 14 dpi. The blot was probed with an antibody specific for the His6-Gly polypeptide tag (g). A coomassie blue-stained SDS-PAGE gel (f) shows equal loading of samples (5 μg total soluble protein). The sizes of the molecular mass standards (marker) and position of the LeMADS-RIN protein (RIN) are indicated on the left and right, respectively.
To test whether the LeMADS-RIN protein could be efficiently expressed from the PVX-based VIGC vectors, infectious RNA transcripts were produced by in vitro transcription from PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN, PVX/HisG::mLeMADS-RIN or the empty PVX vector and mechanically inoculated onto Nicotiana benthamiana. In contrast to the mock-inoculated plant (Fig. 1b), virus-inoculated plants developed local chlorosis in inoculated leaves at 3–7 days post-inoculation (dpi) and followed later by mosaic and curling in systemic young leaves, typical phenotypes associated with PVX infection (Fig. 1c–e). Systemic young leaf tissues were then collected at 14 dpi for further molecular analysis. Recombinant viral RNAs were detected by RT-PCR and direct sequencing of the RT-PCR products confirmed that the LeMADS-RIN RNA was maintained in all VIGC vectors during the course of infection. Moreover, a HisG::LeMADS-RIN protein was readily detected by immunoblotting in plants inoculated with PVX/HisG::LeMADS-RIN, but not in PVX/HisG::mLeMADS-RIN or mock-inoculated plants using an antiserum raised against His6-Gly tag (Fig. 1f, g). These results clearly demonstrate that the tomato LeMADS-RIN protein can be expressed from VIGC vectors in N. benthamiana and over-expression of LeMADS-RIN does not have a deleterious effect on plant growth and development.
LeMADS-RIN mediated complementation in rin fruits
To investigate the feasibility of VIGC in plant gene function analysis, we tested whether ectopic viral expression of LeMADS-RIN could complement the non-ripening phenotype in rin mutant. Through the carpopodium (pedicel), immature (approx. 3–4 weeks after anthesis) or mature green fruits were needle-injected with either recombinant PVX RNA transcripts, or leaf saps from N. benthamiana plants mock-inoculated or inoculated with PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN, PVX/HisG:: mLeMADS-RIN, or the empty PVX vector. For each construct, 40–50 rin fruits were injected in repeated experiments. Not surprisingly, all mock-treated fruits and fruits injected with PVX, PVX/mLeMADS-RIN or PVX/HisG::mLeMADS-RIN remained non-ripe, showing characteristics of the rin mutant phenotype (Fig. 2a, b). However nearly 40% of fruits injected with PVX/LeMADS-RIN or PVX/HisG::LeMADS-RIN appeared to ripen at 2–3 weeks after virus-injection, evidenced by developing red sectors on the fruit (Fig. 2c, d). Thus, expression of LeMADS-RIN or His6-Gly tagged LeMADS-RIN from PVX/LeMADS-RIN or PVX/HisG::LeMADS-RIN respectively, can functionally complement the rin mutant.
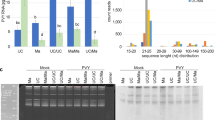
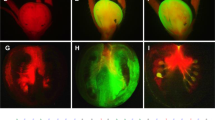
VIGC-mediated complementation of ripening in rin mutant fruits.
Rin fruits were needle-injected with TE buffer (a), or with PVX/HisG::mLeMADS-RIN (b), PVX/LeMADS-RIN (c) and PVX/HisG::LeMADS-RIN (d). Only rin fruits injected with VIGC vector PVX/LeMADS-RIN or PVX/HisG::LeMADS-RIN which are capable of expressing LeMADS-RIN or His6-Gly tagged LeMADS-RIN, respectively, resulted in ripening. Mock-injected fruits or those injected with PVX, PVX/mLeMADS-RIN or PVX/HisG::mLeMADS-RIN from which no LeMADS-RIN protein could be expressed remained green and non-ripe. Fruits were photographed at 3 weeks post-injection (wpi). (e) Presence of VIGC vectors in rin fruits. Western blot analyses of total protein extracted from rin fruits at 3 wpi of the VIGC vector indicated that PVX CP accumulated in ripe red sectors (R), but little to none in the green sectors (G) of three rin fruits (I–III) individually injected with PVX/HisG::LeMADS-RIN. No viral CP was detected in rin fruits that were mock-injected (mock) or injected with the VIGC vector but showed no complementation of ripening (Injected w/c). (f) A coomassie blue-stained SDS-PAGE gel shows equal loading of samples (10 μg total soluble proteins). The positions and sizes of the molecular mass standards (marker) are indicated. (g, h) Semi-quantitative (sq) RT-PCR assays of transient viral expression of LeMADS-RIN. Total RNA was extracted from rin fruits as described in (e) at 3 wpi and used for sqRT-PCR with 25 (g) and 30 (h) cycles of amplification. Equal amount of total RNA (100 ng) used in each of the sqRT-PCR reactions was verified by detecting a similar level of 18S rRNA in all samples. Viral ectopic expression of the LeMADS-RIN mRNA is consistent with presence of the VIGC vectors in the rin fruits. The positions and sizes of the 1-kb DNA marker (Ladder) and positions of RT-PCR products specific for PVX::LeMADS-RIN mRNA (PVX-RIN RNA) as well as 18S rRNA are indicated.
PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN, PVX/HisG::mLeMADS-RIN or PVX were initially delivered into fruits via needle-injection of the carpopodium. However all viruses were able to spread through the plants during the course of the experiment. Consequently, newly grown young systemic leaves developed chlorosis and mosaic and curling phenotypes typically associated with PVX infection. Nevertheless, virus-positive plants, similar to virus-free plants, still had the competence to flower and produce tomato fruits. Remarkably, in plants systemically infected with PVX/LeMADS-RIN or PVX/HisG::LeMADS-RIN, approximately 30% of the newly developed (but non-injected) fruits ripened, producing red ripe sectors as shown in Fig. 2c–d. These data suggest that viruses can move from the injected carpopdium through the phloem transportation system into fruits where viral ectopic expression of the LeMADS-RIN gene can cause rin fruit ripening.
To correlate VIGC of fruit ripening with viral ectopic expression of the LeMADS-RIN gene, we dissected the pericarps of the ripe (red) and non-ripe (green) sectors of rin fruits. Total protein was extracted from the separate ripe and non-ripe tissues, as well as from fruit pericarps injected with PVX/HisG::LeMADS-RIN but showing no complementation and from that of mock-injected rin fruits. Western blot assays showed that the level of the PVX coat protein (CP) was abundant in ripe sectors of rin fruits injected with PVX/HisG::LeMADS-RIN. This was evidenced by the fact that a protein of the predicted size of approximately 26 kDa was specifically recognised by an antibody raised against the PVX CP (Fig. 2e) while coomassie blue-stained SDS-PAGE gel shows equal loading of samples (Fig. 2f). There was no accumulation of PVX CP in the mock rin fruit, or in the non-complemented fruit injected with PVX/HisG::LeMADS-RIN. On the other hand, in green sectors of the complemented fruits, the level of PVX CP was mostly undetectable or significantly reduced (Fig. 2e, f). Thus, the presence of the VIGC vector PVX/HisG::LeMADS-RIN was consistent with the functional complementation of ripening in the rin fruits. However to our surprise, using an anti-His6-Gly antibody we were unable to detect the HisG::LeMADS-RIN protein in the pericarps of ripe sectors. It is possible that a low level of HisG::LeMADS-RIN sufficient to complement the rin mutant phenotype was expressed but was beyond the limit of western blot detection. Such low expression could be the result of gene silencing induced by the VIGC vector, or by loss of the inserted LeMADS-RIN RNA sequence from the PVX genomic RNA.
We thus used a more sensitive semi-quantitative (sq) RT-PCR approach to detect LeMADS-RIN mRNA expressed from the PVX-based VIGC vectors. Total RNA was extracted from ripening-inhibited fruits collected from mock-inoculated rin fruits and from the separate red ripe and green non-ripe tissues, as well as from non-complemented fruits at 3 weeks post-injection with PVX/HisG::LeMADS-RIN. Total RNA (100 ng) was used together with a set of primers PP82/PP779 (Fig. 1a; Tables S1 and S2) in one-step sqRT-PCR assays. These primers allowed specific detection of an approximately 900-bp transcript containing partial PVX RNA and the full-length LeMADS-RIN mRNA (Fig. 1a). Consistent with western blot assays, no transient viral LeMADS-RIN mRNA was detected in pericarps of the control fruits. However PVX::LeMADS-RIN RNA was detected in the red ripening sectors of complemented fruits whilst levels of these RNAs were either undetectable or obviously reduced in the green non-ripening sector of the same fruits (Fig. 2e, f). That an equal amount of total RNA was used in each of the sqRT-PCR reactions was verified by detecting a similar level of 18S rRNA in all samples using primers PP271 and PP272 (Tables S1 and S2). A similar result was also obtained in complemented rin fruits injected with PVX/LeMADS-RIN (Fig. S2). Moreover, the non-translatable mLeMADS-RIN mRNA was detectable in fruits which were injected with PVX/mLeMADS-RIN or PVX/HisG::mLeMADS-RIN and remained non-ripe (Fig. S2). Taken together, these data demonstrate that the ripening phenotypes were caused by LeMADS-RIN expression from the VIGC vectors.
LeMADS-RIN regulates LeSPL-CNR and LeHB-1 expression
Complementation of the rin mutant through VIGC offers a quick and efficient strategy to directly investigate LeMADS-RIN-dependent regulation of ripening-associated genes. To test this idea, we performed comparative sqRT-PCR assays and examined the level of RNA transcripts of LeSPL-CNR, LeHB-1 and SlTPR1 in mock-treated and VIGC fruits. These genes were chosen because they are known to be involved in fruit ripening through direct or indirect regulation of ethylene biosynthesis and signalling transduction13,14,29. The mRNA level of LeSPL-CNR was low in mock-injected fruit and in a non-complemented fruit that had been injected with the VIGC vector PVX/HisG::LeMADS-RIN. A similar expression pattern for LeHB-1 or SlTPR1 gene was also observed (Fig. 3a, b). However sqRT-PCR assays of total RNA extracted from the non-ripe green and ripening red sectors of complemented rin fruits injected with PVX/HisG::LeMADS-RIN clearly showed that viral transient expression of LeMADS-RIN had different impacts on the expression of the three ripening-associated genes. LeSPL-CNR was readily detected in the ripening sectors after 25 cycles of PCR (Fig. 3a), its mRNA level being higher than that in the non-ripening green sectors although such differential expression levels became less obvious after a saturated 30-cycle PCR (Fig. 3b). On the other hand, the mRNA level of LeHB-1 was constantly lower in ripening red than non-ripe green sectors (Fig. 3a, b). In contrast to both LeSPL-CNR and LeHB-1 TF genes, the level of SlTPR1 expression remained relatively unaffected. A similar level of 18S rRNA was detected in all samples, showing that an equal amount of total RNA (100 ng) was used in each of the sqRT-PCR reactions. Taken into account the LeMADS-RIN gene expression levels in these tissues (Fig. 2), these data indicate that LeSPL-CNR is up-regulated while LeHB-1 is down-regulated by LeMADS-RIN, but that LeMADS-RIN is not directly involved in the regulation of the SlTPR1 gene transcription.
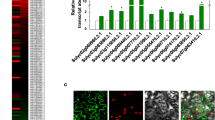
Impact of LeMADS-RIN on expression of LeSPL-CNR, LeHB1 and SlTPR1.
Total RNA was extracted from rin fruits as described in Fig. 2e at 3 wpi of the VIGC vectors and used for sqRT-PCR with 25 (a) and 30 (b) cycles of amplification. That an equal amount of total RNA (100 ng) was used in each of the sqRT-PCR reactions was verified by detecting a similar level of 18S rRNA in all samples. The positions and sizes of the 1-kb DNA marker (Ladder) and positions of RT-PCR products specific for LeSPL-CNR, LeHB1 and SlTPR1 mRNA as well as 18S rRNA are indicated.
Discussion
In this report we describe a VIGC technology for virus-induced gene complementation and provide ‘proof of concept’ that VIGC works in defining functions of fruit ripening–related TF genes in the tomato rin mutant. Several advantages of VIGC can be envisaged in its application in plant functional genomics. First, VIGC is an easy and quick technique to complement gene function in mutant plants. Ectopic over-expression of plant genes from a VIGC vector results in ‘gain-of-function’, thus providing a biological link between phenotype and genotype. Second, VIGC possesses a relatively high efficiency for mutant phenotype complementation, although this efficiency could still be improved through optimization of experimental conditions and inoculation protocols. Third, VIGC does not require time-consuming and labour-intensive stable transformation that is associated with transgenic complementation. Fourth, the feasibility of VIGC based on a single plant virus PVX was only tested in the tomato rin mutant. Further studies to develop various virus-based VIGC vectors are currently ongoing. However, the rational and potential uses of VIGC should be applicable to other plant and crop species. For gene function analysis in tomato fruit ripening, VIGC offers an additional advantage that ripening and non-ripe tissues can be collected from the same individual fruits. These tissues which are at the precisely equivalent age and stage are most suitable for comparative gene expression analysis.
Tomato is an excellent model crop to investigate fruit development and ripening. Recent advances on the understanding of genetic and molecular mechanisms that control fruit ripening have come from the study of various tomato ripening mutants, such as rin and Cnr27,30. Some of these mutant loci encode master ripening modulator TFs including LeMADS-RIN and LeSPL-CNR13,26. LeHB-1 and several other TFs have also been demonstrated to play an important role in tomato fruit ripening14,31,32,33,34,35. Recently, it has been reported that expression of TAGL1 (Tomato AGAMOUS-like 1), a member of the AGAMOUS clade of MADS-box genes, is capable of complementing the rin mutant although this contradicts the fact that the rin mutant maintains normal TAGL1 expression without ripening33,36. Nevertheless, the rationale of transcriptional control of fruit ripening is well established. However, whether a functional network exists among ripening-related TFs remains to be elucidated.
Using VIGC, we have demonstrated that ectopic expression of LeMADS-RIN resulted in functional complementation of non-ripening phenotypes and caused rin fruits to ripen (Fig. 1; Fig. 2). Such complementation was unlikely to be caused by the recombination of homologous RNA sequences. However it could be predicted that actually VIGS would take place rather than VIGC. Having an endogenous gene in the viral genome would be expected to induce VIGS and potentially produce little of the virus-expressed gene product. Thus VIGS probably contributed to our failure to detect the LeMADS-RIN protein in rin tomato, but had little effect on accumulation of the TF protein in heterologous N. benthamiana plants (Fig. 1; Fig. 2). On the other hand viral infection as well as fruit ripening are dynamic processes, VIGS usually occurs after virus establishes infection and PVX is able to evade self-silencing37, therefore PVX-based vectors are suitable to be used for VIGC. In our experiments, the observed complementation effect in the rin mutant could derive from a temporary and transient expression of the LeMADS-RIN TF. This could occur from the VIGC vectors at the beginning of the viral infection prior to occurrence of VIGS, or from VIGC viral RNA being survived the VIGS, triggering a ripening process which was only observed afterwards. VIGS and off-target gene silencing could be induced by all four VIGC vectors, however complementation was only observed in rin fruits injected with PVX/LeMADS-RIN or PVX/HisG::LeMADS-RIN, but not in any fruits injected with PVX/mLeMADS-RIN or PVX/HisG::mLeMADS-RIN. Thus this excludes the possibility that VIGS of LeMADS-RIN or siRNA-mediated silencing of off-target genes could result in the rin complementation.
Our data support the conclusion that rin complementation has taken place in a percentage (40%) of injected fruits. VIGS might be the reason that the remaining 60% of fruits do no show complementation. It is also possible that homologous recombination between the endogenous mutant rin transcript and the LeMADS-RIN RNA in the VIGC vectors occurred during virus RNA replication in the cytoplasm of cells. In this scenario the modified virus, which could incorporate part of endogenous mutant rin RNA sequences, might continue its systemic movement within the tomato plants. This could subsequently lead to VIGC vectors loosing their capacity of complementation in fruits although the probability of such type of event is unknown. In addition, instability of RNA inserts in the VIGC viral genomic RNA could have affected complementation efficiency. Partial or even complete elimination of the LeMADS-RIN RNA was observed in all PVX constructs (Fig. 3S, Supplementary Text) and that would certainly have a negative influence on the efficiency of VIGC. Moreover, experimental conditions including inoculation protocols, timing and plant growth environments could also affect VIGC.
The LeMADS-RIN-mediated VIGC process does not affect the expression of SlTPR1, which encodes an ethylene receptor-interacting protein and functions in the ethylene signalling transduction associated with fruit ripening29. However, LeMADS-RIN transiently expressed from VIGC vectors was able to up-regulate LeSPL-CNR expression (Fig. 2; Fig. 2S; Fig. 3), consistent with the observed reduction of LeSPL-CNR expression in rin mutant27. Our finding suggests that LeMADS-RIN is likely to act upstream of LeSPL-CNR in a proposed TF network (Fig. 4). On the other hand LeHB-1 is highly expressed in immature and mature green fruits, but the level of LeHB-1 mRNA declines during ripening and is maintained at a relatively low level in ripe fruits. LeHB-1 is involved in transcriptional regulation of ACC oxidase (LeACO1) gene expression through its binding to the LeACO1 promoter. Silencing of LeHB-1 in tomato fruit greatly reduces (LeACO1) mRNA levels and inhibits ripening. However, over-expression of LeHB-1 causes conversion of floral organs into fruits14. Thus expression of the homeobox TF LeHB-1 must be tightly controlled during the process of tomato fruit development and ripening. Consistent with this idea, we found that LeMADS-RIN acted as a negative modulator to down-regulate LeHB-1 transcription in the ripening sector of rin fruits (Fig. 3). Taken together, our comparative analysis of gene expression in the ripe and non-ripe sectors of rin fruits suggests that LeMADS-RIN might link with LeSPL-CNR and LeHB-1 to form a potential TF network in the modulation of fruit ripening (Fig. 4). It is possible that expression changes of LeSPL-CNR and LeHB-1 could just be a consequence of fruit ripening. In this scenario LeMADS-RIN might only have an indirect effect on these downstream TF genes. However, direct binding of LeMADS-RIN to the promoters of LeSPL-CNR and LeHB-1 has been elegantly demonstrated through various molecular and biochemical assays38,39. Thus our findings, along with experimental data reported in the literatures38,39 are in favour of a model in which LeMADS-RIN probably exerts a direct impact on LeSPL-CNR and LeHB-1 gene expression in the transcriptional regulation of tomato fruit ripening.
A regulatory network among key ripening-associated TFs.
Methionine (Met) is converted into S-adenosyl methionine (S-AdeMet) by the S-AdeMet synthetase. The key enzyme ACC synthase cleaves the amino butyrate from S-AdeMet to release 1-aminocyclopropane-1-carboxylic acid (ACC). The ACC oxidase then converts ACC to ethylene which acts as an essential phytohormone to trigger fruits to ripen in tomato. The master TF LeMADS-RIN may affect production of ACC synthase and regulate the biosynthesis of ethylene. LeMADS-RIN up-regulates (+) the transcription of LeSPL-CNR that may be directly or indirectly involved in the control of ethylene biosynthesis. Meanwhile, LeMADS-RIN serves as a negative TF to down-regulate (−) the LeHB1 expression at the transcriptional level. The latter is shown to directly modulate the ACC oxidase gene expression and affect ethylene production for fruit ripening. However, over-expression of LeHB1 also results in abnormal flower (Flower*) and fruit development14. The SlTPR1 gene product interacts with the ethylene receptor LeETR1, which exerts a positive impact on fruit ripening29.
In summary, we have exploited the potential of plant virus technology, namely VIGC, for gene expression to induce phenotypes and to complement gene function in the tomato rin mutant. Using VIGC we provide evidence to support the hypothesis that a transcriptional network may exist among three key TFs LeMADS-RIN, LeSPL-CNR and LeHB-1 in the modulation of tomato ripening. The function of the master TF LeMADS-RIN and its impact on LeSPL-CNR and LeHB-1 expression as well as on ethylene-mediated ripening are shown in a proposed model (Fig. 4). This model is supported by the findings of other research groups that the LeMADS-RIN protein can directly bind to the promoters of the LeSPL-CNR and LeHB-1 genes38,39.
Methods
Construction of VIGC vectors
A cDNA clone of the tomato (Solanum lycopersicum cultivar Ailsa Craig) LeMADS-RIN gene in pK7WGF2 (Zhong, L. & Giovannoni, J.J. unpublished data) was used as a template for construction of all VIGC vectors. It should be noted that S. lycopersicum has been recently renamed from Lycopersicon esculentum and genes isolated before and after the re-nomenclature contain either Le or Sl prefix, respectively. The wild-type and mutant LeMADS-RIN genes were PCR amplified using Pfu DNA polymerase (Promega) and a pair of primers PP797/PP799 or PP798/PP799 (Tables S1 and S2). The resulting PCR products were digested with ClaI and EagI and cloned into the ClaI/EagI sites of PVX28 to produce PVX/LeMADS-RIN and PVX/mLeMADS-RIN respectively (Fig. 1a). The wild-type and non-sense mutant LeMADS-RIN genes were also cloned in-frame to the coding sequence of a histidine-glycine (His6-Gly) tag in the ClaI/EagI sites of a modified PVX/HisGMF1 vector (Fig. S1) to generate PVX/HisG::LeMADS-RIN and PVX/HisG::mLeMADS-RIN (Fig. 1a). All constructs were confirmed by nucleotide sequencing.
Viral infection and plant maintenance
RNA transcripts were produced by in vitro transcription from each of the PVX constructs (Fig. 1) after linearization with SpeI and then were mechanically inoculated onto young leaves of N. benthamiana and S. lycopersicum cultivar Ailsa Craig plants as described28. Mock inoculation of plants with TE buffer (10 mM Tris-Cl, pH 8.0, 1 mM EDTA) was used as a negative control. All plants were maintained in an insect-free glasshouse at 25°C with supplementary lighting to give a 16-h photoperiod. Local and systemic symptom developments were assessed on a daily basis and recorded photographically using a Nikon Digital Camera Coolpix990.
VIGC (Virus-induced gene complementation)
RNA transcripts of PVX, PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN or PVX/HisG::mLeMADS-RIN were needle-injected into the carpopodium of rin tomato fruits at the mature green stage. Fruits were injected with TE buffer as negative controls. Homogenized saps prepared from systemic leaves of N. bethamiana with mock-inoculation or inoculated with PVX, PVX/LeMADS-RIN, PVX/mLeMADS-RIN, PVX/HisG::LeMADS-RIN or PVX/HisG::mLeMADS-RIN were also used to injected rin fruits on different trusses of the tomato plants in repeated experiments (more than three times). Within 2–3 weeks after injection rin fruits started to develop ripening sectors due to VIGC-mediated effect and they were routinely examined and photographed with a Nikon Coolpix 995 digital camera.
Semi-quantitative RT-PCR (sqRT-PCR)
Total RNA was extracted from leaf tissues of N. benthamiana and from tomato fruits using an RNeasy Plant Mini Kit (Qiagen). RT-PCR was performed as previously described20 with different pairs of primers PP82/PP83, PP82/PP799, PP83, PP271/PP272, PP298/PP300, PP402/PP403 and PP404/PP405 (Tables S1 and S2) to detect PVX, PVX::LeMADs-RIN RNA, plant 18S rRNA, LeSPL-CNR, LeHB1 and SlTPR1 mRNA, respectively.
Western blotting
To investigate expression of HisG::LeMADS-RIN and viral coat protein (CP) from the VIGC vectors, total protein was extracted from leaf and fruit tissues collected from N. benthamiana or tomato as previously described40. Western blot analyses of protein aliquots (10 μg) were performed with polyclonal antibodies specifically raised against PVX CP, or an anti-His-Gly tag antibody (Invitrogen) and detected using a goat anti-rabbit immunoglobulin G conjugated with alkaline phosphatase (Sigma) and 5-bromo-4-chloro-3-indolylphosphate-nitroblue tetrazolium (BCIP/NBT) substrates (Roche) as previously described37,40.
References
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Schnable, P. S. et al. The B73 Maize Genome: Complexity, Diversity and Dynamics. Science 326, 1112–1115 (2009).
The Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 475, 189–195 (2011).
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Robinson, S. J. & Parkin, I. A. P. Bridging the gene-to-function knowledge gap through functional genomics. In Plant Genomics Methods and Protocols (eds Somers, D.J., Langridge, P. & Gustafson, J.P.), Methods in Mol. Biol. 513, 153–173 (2009).
Lindbo, J. A. et al. Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5, 1749–1759 (1993).
Kumagai, M. H. et al. Cytoplasmic inhibition of carotenoid biosynthesis with virus-derived RNA. Proc. Natl. Acad. Sci. 92, 1679–1683 (1995).
Ruiz, M. T. et al. Initiation and maintenance of virus-induced gene silencing. Plant Cell 10, 937–946 (1998).
Liu, Y. et al. Virus-induced gene silencing in tomato. Plant J. 31, 777–786 (2002).
Becker, A. & Lange, M. VIGS – genomics goes functional. Trends in Plant Sci. 15, 1–4 (2009).
Senthil-Kumar, M. & Mysore, K. S. New dimensions for VIGS in plant functional genomics. Trends in Plant Sci. 16, 656–665 (2011).
Manning, K. et al. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet. 38, 948–952 (2006).
Lin, Z. F., Hong, Y. G., Yin, M. G., Li, C. Y., Zhang, K. & Grierson, D. A tomato HD-Zip homeobox protein, LeHB-1, plays an important role in floral organogenesis and ripening. Plant J. 55, 301–310 (2008).
Takamatsu, N., Ishikawa, M., Meshi, T. & Okada Y. Expression of bacterial chloramphenicol acetyltransferase gene in tobacco plants mediated by TMV-RNA. EMBO J. 6, 307–311 (1987).
Donson, J., Kearney, C. M., Hilf, M. E. & Dawson, W. O. Systemic expression of a bacterial gene by a tobacco mosaic virus-based vector. Proc. Natl. Acad. Sci. 88, 7204–7208 (1991).
Chapman, S., Kavanagh, T. & Baulcombe, D. Potato virus X as a vector for gene expression in plants. Plant J. 2, 549–557 (1992).
Zeng, X. W. et al. Expression of highly immunogenic tuberculosis proteins in plants using a versatile Potato virus X-based expression system. J. Hort. Sci. Biotechnol. 83, 4–8 (2008).
Hong, Y., Stanley, J. & van Wezel, R. Novel system for the simultaneous analysis of geminivirus DNA replication and plant interactions in Nicotiana benthamiana. J. Virol. 77, 13315–13322 (2003).
Li, C. et al. A cis element within Flowering Locus T mRNA determines its mobility and facilitates trafficking of heterologous viral RNA. J. Virol. 83, 3540–3548 (2009).
Li, C. et al. Mobile FT mRNA contributes to the systemic florigen signalling in floral induction. Sci. Rep. 1, 73; DOI:10.1038/srep00073 (2011).
Ryabov, E. V., Van Wezel, R., Walsh, J. & Hong, Y. Cell-to-Cell, but not long-distance spread of RNA silencing induced initially in individual epidermal cells. J. Virol. 78, 3149–3154 (2004).
Zhou, Y., Ryabov, E., Zhang, X. & Hong, Y. Influence of viral genes on the cell-to-cell spread of RNA silencing. J. Exp. Bot. 59, 2803–2813 (2008).
Shi, Y. et al. Suppression of local RNA silencing is not sufficient to promote cell-to-cell movement of Turnip crinkle virus in Nicotiana benthamiana. Plant Signal. Behav. 4, 15–22 (2009).
Qin, C. et al. Involvement of RDR6 in short-range intercellular RNA silencing in Nicotiana benthamiana. Sci. Rep. 2, 467; DOI:10.1038/srep00467 (2012).
Vrebalov, J. et al. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (Rin) locus. Science 296, 343–346 (2002).
Klee, H. J. & Giovannoni, J. J. Genetics and control of tomato fruit ripening and quality attributes. Annu. Rev. Genet. 45, 41–49 (2011).
Van Wezel, R., Liu, H., Tien, P., Stanley, J. & Hong, Y. Gene C2 of the monopartite geminivirus Tomato yellow leaf curl virus from China virus encodes a pathogenicity determinant that is localised in the nucleus. Mol. Plant-Microbe Interact. 14, 1125–1128 (2001).
Lin, Z. F. et al. SlTPR1, a tomato tetratricopeptide repeat protein, interacts with the ethylene receptors NR and LeETR1, modulating ethylene and auxin responses and development. J. Exp. Bot. 59, 4271–4287 (2008).
Seymour, G., Poole, M., Manning, K. & King, G. J. Genetics and epigenetics of fruit development and ripening. Curr. Opin. Plant Biol. 11, 58–63 (2008).
Itkin, M., Seybold, H., Breitel, D., Rogachev, I., Meir, S. & Aharoni, A. Tomato AGAMOUS-LIKE 1 is a component of the fruit ripening regulatory network. Plant J. 60, 1081–1095 (2009)
Pan, I., McQuinn, R., Giovannoni, J. & Irish, V. Functional diversification of AGAMOUS lineage genes in regulating tomato flower and fruit development. J. Exp. Bot. 61, 1795–1806 (2010).
Vrebalov, J. et al. Fleshy fruit expansion and ripening are regulated by the tomato SHATTERPROOF gene TAGL1. Plant Cell 21, 3041–3062 (2009).
Chung, M. et al. A tomato (Solanum lycopersicum) APETALA2/ERF gene, SlAP2a, is a negative regulator of fruit ripening. Plant J. 64, 936–947 (2010).
Karlova, R. et al. Transcriptome and metabolite profiling show that APETALA2a is a major regulator of tomato fruit ripening. Plant Cell 23, 923–941 (2011).
Gimenez, E. et al. Functional analysis of the Arlequin mutant corroborates the essential role of the ARLEQUIN/TAGL1 gene during reproductive development of tomato. PLoS ONE 5, e14427 (2010).
Van Wezel, R. & Hong, Y. Virus survival of RNA silencing without deploying protein-mediated suppression in Nicotiana benthamiana. FEBS Lett. 562, 65–70 (2004).
Martel, C., Vrebalov, J., Tafelmeyer, P. & Giovannoni, J. J. The Tomato MADS-Box transcription factor RIPENING INHIBITOR interacts with promoters involved in numerous ripening processes in a COLORLESS NONRIPENING-dependent manner. Plant Physiol. 157, 1568–1579 (2011).
Fujisawa, M. et al. Direct targets of the tomato-ripening regulator RIN identified by transcriptome and chromatin immunoprecipitation analyses. Planta 235, 1107–1122 (2012).
Hong, Y., Saunders, K., Hartley, M. R. & Stanley, J. Resistance to geminivirus infection by virus-induced expression of a plant cytotoxin. Virology 220, 119–127 (1996).
Acknowledgements
We thank David Baulcombe for providing the original PVX vector and Simon Santa Cruz for providing the PVX CP antibody. We are grateful to Jose Gutierrez-Marcos for critical reading of the manuscript. T.Z. was supported by an UK-China Scholarships for Excellence. This work was supported by the UK Biotechnology and Biological Sciences Research Council (BBSRC)–Warwick HRI core funding (BBS/E/H/00YH0271), a grant from the Royal Society (Research Grants Scheme 2007/R1/Hong), a Royal Society KC Wong Fellowship (2006R2/China/KCW/Hong) and an innovative R&D grant from Hangzhou Normal University to Y.H.
Author information
Authors and Affiliations
Contributions
T.Z., H.Z., T.L., C.Q., N.S., H.W., M.J. and S.Z. designed and performed experiments, Z.F., Y.L., Z.W., S.J., J.J.G., D.R. and P.G. contributed through discussions and revised the paper. Y.H. initiated the project, designed the experiments and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Zhou, T., Zhang, H., Lai, T. et al. Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening. Sci Rep 2, 836 (2012). https://doi.org/10.1038/srep00836
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00836
This article is cited by
-
Rapid, high efficiency virus-mediated mutant complementation and gene silencing in Antirrhinum
Plant Methods (2020)
-
An RNAi suppressor activates in planta virus–mediated gene editing
Functional & Integrative Genomics (2020)
-
A qualitative proteome-wide lysine crotonylation profiling of papaya (Carica papaya L.)
Scientific Reports (2018)
-
Comparative WGBS identifies genes that influence non-ripe phenotype in tomato epimutant Colourless non-ripening
Science China Life Sciences (2018)
-
Tomato tolerance to abiotic stress: a review of most often engineered target sequences
Plant Growth Regulation (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.