Abstract
Missense mutations in TP53 gene promote metastasis in human tumours. However, little is known about the complete loss of function of p53 in tumour metastasis. Here we show that squamous cell carcinomas generated by the specific ablation of Trp53 gene in mouse epidermis are highly metastatic. Biochemical and genome-wide mRNA and miRNA analyses demonstrated that metastases are associated with the early induction of epithelial-mesenchymal transition (EMT) and deregulated miRNA expression in primary tumours. Increased expression of miR-21 was observed in undifferentiated, prometastatic mouse tumours and in human tumours characterized by p53 mutations and distant metastasis. The augmented expression of miR-21, mediated by active mTOR and Stat3 signalling, conferred increased invasive properties to mouse keratinocytes in vitro and in vivo, whereas blockade of miR-21 in a metastatic spindle cell line inhibits metastasis development. Collectively these data identify novel molecular mechanisms leading to metastasis in vivo originated by p53 loss in epithelia.
Similar content being viewed by others
Introduction
Tumour metastasis is the major cause of mortality of human cancers. In epithelial tumours, metastasis is frequently associated with the loss of epithelial characteristics and the acquisition of mesenchymal properties by a genetically controlled process named epithelial mesenchymal transition (EMT), which is also essential during embryonic development1,2. These facts have posed tremendous efforts to understand the molecular mechanism governing EMT in order to find possible therapeutic agents that impair metastatic growth of the tumours. A number of master genes controlling EMT in tumors have been identified2. Importantly, EMT is also related to cancer cell stemness3 and the presence of these putative cancer stem cells is associated with metastatic spreading4,5.
The p53 tumour suppressor coordinates the cellular response to stress, including DNA damage, hypoxia and oncogenic stress through transcriptional mechanisms, resulting in cell cycle arrest, senescence, or apoptosis. Accordingly, p53 mutations are widely involved in human tumourigenesis6 and are also associated with poor prognosis and high metastatic potential in human tumours6,7. Recently, several groups have demonstrated that missense, gain of function-associated mutations in TP53 gene, promote tumour metastasis by interfering with integrin and TGFβ signalling8,9. However, little is known about the molecular mechanisms of metastasis due to complete loss of function of p53. Indeed, observations based on mouse knock out models have led to the assumption that complete p53 loss is not prone to metastasis10, unless other members of the p53 family are also ablated11. However, gene mutation analyses of human cancer clinical samples revealed that loss of p53 is also frequent in metastatic tumours, although the molecular mechanisms involved are not defined.
MicroRNAs (miRNAs) are a recently discovered class of small RNA molecules that negatively regulate gene expression at the post-transcriptional level. As some of these genes are involved in the control of development, proliferation, apoptosis and stress response, it is not surprising that deregulated expression of miRNAs is implicated in tumor development. Importantly, miRNAs have been associated with metastatic spreading, EMT and cancer stem cells thus opening new therapeutic avenues12,13,14. Recent evidences have also linked the p53 tumor suppressor with the altered expression or maturation of miRNAs15,16,17,18.
We and others have previously described that the specific ablation of Trp53 gene in stratified epithelia (hereafter p53ΔEC) leads to spontaneous squamous cell carcinomas (SCC) development19,20. Tumor development is associated to premature chromosome instability21 and is accelerated by the epidermal ablation of Rb1 gene (hereafter RbΔEC). This last aspect is due to the increased proliferation, promoted by pRb loss, in conjunction with activation of specific signal transduction mechanisms20,21. Such cooperative functions can also explain the findings obtained in RbΔEC mice. These RbΔEC mice showed altered differentiation and increased proliferation in epidermis, but did not develop spontaneous tumours22. Moreover, upon chemical carcinogenesis protocols RbΔEC mice exhibited reduced tumour susceptibility, although the tumors displayed more malignant characteristics due to premature p53 loss23,24. The relationship between pRb and p53 in epithelial cells is also supported by RbΔEC; p107-/- mouse models, which are prone to oncogenic transformation due, in part, to the impairment of p53 proapoptotic functions25.
Gene expression analysis of overt tumours arising in RbΔEC; p53ΔEC and p53ΔEC mice revealed no significant differences between the two genotypes26, indicating that cooperation is only evidenced at early stages of tumor development20. The majority of genes overexpressed in tumours are involved in cell cycle and, in particular, in mitosis26, in agreement with the described altered chromosome instability mediated by the epidermal loss of p5321. These genomic profiling studies also revealed a highly significant overlap with stem cell signatures and evidences of EMT processes and demonstrated that these mouse tumours share relevant characteristics with multiple human malignancies distinguished by poor prognosis, altered p53 status and, remarkably, high metastasis incidence26. These data prompted us to analyze the possible metastatic capacities of spontaneous tumours arising in RbΔEC; p53ΔEC and p53ΔEC mice. Here we show that these epidermal tumours are highly prone to form lung metastasis in association with an early development of EMT genetic program. In addition, we also observed that primary tumours leading to lung metastasis displayed deregulated expression of specific miRNAs, including miR-21. Overexpression and blockade experiments demonstrated not only the involvement of miR-21 in EMT, but also its specific implication in tumour aggressiveness and metastatic spreading. Collectively, our findings indicate that targeted therapies aimed to restore the affected pathways and/or modulate miRNA expression would be of great benefit in the treatment of tumours bearing altered p53 functions, in particular those bearing truncated p53 mutations.
Results
Trp53-deficient epidermal tumors are metastatic
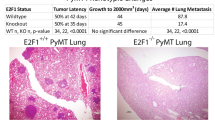
To analyze the metastatic potential of spontaneous tumours from RbΔEC; p53ΔEC and p53ΔEC mice, a full necropsy analysis was performed in mice bearing primary tumors. We found that a high proportion of these mice displayed lung metastases, in some cases macroscopically detectable (Fig 1a). Moreover, upon histology studies we found that a large number of mice displaying primary epidermal tumours also exhibited the presence of small cell masses in the lungs (Fig 1b). The expression of epidermal keratin K5 in these lesions (Fig 1b’), together with the absence of similar events in mice without primary epidermal tumours and the absence of recombination of Rb1 or Trp53 genes in lungs (not shown), strongly suggested that these tumours were metastasis derived from epidermal primary tumours. The metastases were also detected by PET studies by 30-50 days after the detection of primary tumours (Fig 1c). This delay was also evidenced by analyzing cohorts of RbΔEC; p53ΔEC and p53ΔEC mice (Fig 1d, d’). Overall, 50-65% of the mice that displayed primary epidermal tumours also showed metastatic outgrowths in the lungs. These studies also indicated that, similarly to the primary tumours (Fig 1d), in RbΔEC; p53ΔEC mice metastases occurred earlier than in p53ΔEC mice (Fig 1d’). However, the metastasis rate was similar in both groups (Fig 1e). On the other hand, when the metastasis rate was related to the histology of the primary tumour, we observed increased metastasis incidence in mice bearing spindle cell carcinomas (SpCCs) compared to squamous cell carcinomas (SCCs) (Fig 1e’). These results demonstrated that p53-deficient epidermal tumours (both RbΔEC; p53ΔEC and p53ΔEC) are highly metastatic and the metastasis are more frequent when primary tumours display spindle undifferentiated phenotype.
Spontaneous tumours in RbΔEC; p53ΔEC and p53ΔEC mice are metastasic.
(a) Example of a lung showing macroscopic metastases (red arrows). (b) H&E and (b’) anti K5 stained section from a lung showing micrometastatic outgrowths positive for the epidermal marker keratin K5. (c) Examples of in vivo imaging using PET at days 0, 20 and 40 after detection of two primary tumours in the neck area (white arrows). Note the presence of small positive areas in the lungs (small orange arrows) 40 days after the detection of the primary tumours. Kaplan Meier distribution of primary tumour (d) and metastasis (d’) in RbΔEC; p53ΔEC ( ) and p53ΔEC (
) and p53ΔEC ( ) mouse cohorts. p values correspond to Log-Rank test. Distribution of metastasis incidence (mice bearing metastasis with respect to total number of mice with primary tumors) according to the mouse genotype (e) or the differentiation status of the primary tumor type (e’).
) mouse cohorts. p values correspond to Log-Rank test. Distribution of metastasis incidence (mice bearing metastasis with respect to total number of mice with primary tumors) according to the mouse genotype (e) or the differentiation status of the primary tumor type (e’).
Trp53-deficient epidermal tumors undergo EMT
The gene expression profile of primary tumours arising in RbΔEC; p53ΔEC and p53ΔEC mice suggested that SpCCs underwent EMT26. Indeed in supervised hierarchical clustering aimed to discriminate between RbΔEC; p53ΔEC and p53ΔEC tumours, the metastatic primary tumors (denoted by arrows in Fig 2a) did not cluster together. On the contrary, when a recently developed EMT signature27 was applied to the microarray data in unsupervised clustering metastatic tumors are grouped together (Fig 2b). We thus studied the possible EMT events in SCCs and SpCCs arising in RbΔEC; p53ΔEC and p53ΔEC mice. In agreement with the spindle morphology, we observed decreased E-cadherin (Fig 2c, c’) and increased vimentin (Fig 2d, d’) expression in SpCCs. Since gene expression data revealed that the SpCC showed increased expression of essential transcriptional modulators of EMT process26, we also studied their expression by immunohistochemistry. We observed expression of Twist, Snail and FoxC2 (Fig 2e-g’) in SCCs and in SpCCs. However, in SCCs their expression occurred prior to overt EMT process in migrating cells and more undifferentiated areas of squamous tumours (Fig 2e, f, g), whilst in SpCCs (Fig 2e’, f’, g’) we observed generalized expression of these transcription factors. In agreement, western blot analyses showed increased expression of Snail, FoxC2, Twist and Zeb1 in SpCC compared to SCCs, in parallel with decreased E-Cadherin and increased Vimentin expression (Fig 2h). Finally, qRT-PCR studies showed that increased expression was found in primary tumours originating metastasis and in SpCC compared to SCCs (Fig 2i), although no major differences in Snai1 gene expression levels between RbΔEC; p53ΔEC and p53ΔEC tumours.
Primary tumors in RbΔEC; p53ΔEC and p53ΔEC mice undergo EMT.
(a-b) Hierarchical clustering of mouse tumors and normal skin showing the discrimination between genotypes (a) and according to an EMT gene signature (b). Note that metastatic tumours (denoted by arrows) cluster together only in the second case. c-g’) Examples of spontaneous squamous (c, d, e, f, g) and spindle cell carcinoma (c’, d’, e’, f’, g’) showing the expression of E-Cadherin (c, c’), Vimentin (d, d’), Twist (e, e’), Snail (f, f’) and FoxC2 (g, g’). Dashed lines in c-g denote the boundaries between differentiated and undifferentiated areas in squamous tumours. Bars = 150 μm. (h) Western blot showing the expression of the quoted EMT markers and modulators in skin, squamous and spindle cell carcinomas. i) qRT-PCR analysis showing the expression of Snai1 gene in tumours according the mouse genotype (upper panel), metastatic behaviour (middle Panel) and tumour differentiation characteristics (lower panel) n = number of tumours analysed.
Deregulated miRNA expression in metastasic mouse epidermal tumors
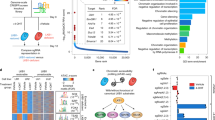
Recent findings have demonstrated that deregulated expression of specific miRNAs controls EMT and metastasis28,29,30 and is involved in TGFβ signalling pathway31. We thus investigated whether deregulated miRNA expression could also contribute to spontaneous tumor development and metastasis in RbΔEC; p53ΔEC and p53ΔEC mice. Microarray analysis showed that 210 different miRNA species (448 probes) could discriminate between epidermis and tumors (Fig 3a and Supp Table 1). This set of selected miRNAs showed both increased and decreased expression compared to normal skin and included several species previously found to be expressed in epidermis32,33 and different miRNAs previously involved in metastasis34,35. In agreement with the increased expression of multiple miRNA in tumours, we found only few tumor samples showing decreased expression of miRNA processing proteins Dicer, Dgcr8 or Drosha compared to normal samples (Supp Fig 1a), suggesting that the observed changes are not attributable to impaired miRNA processing, as reported in other tumour types36,37.
Altered miRNA expression in p53-deficient spontaneous tumors.
(a, b) Hierarchical clustering of miRNA showing the differential expression discriminating between normal skin and mouse tumours (a) and discriminating normal skin, squamous and spindle tumours (b). (c) Box plot showing the overall expression of miR-200 family of miRNAs between squamous and spindle tumours obtained by qRT-PCR. (d) Box plots showing the expression of miR-21 in metastatic and non metastatic mouse tumours (upper panel), in tumours from RbΔEC; p53ΔEC and p53ΔEC mice (middle panel) and spindle and squamous cell carcinomas (lower panel) n = number of tumours analysed. e-h′) Immunohistochemistry examples showing the expression of Sprouty2 (e, e′), phosphorylated Erk (f, f′), PTEN (g, g′) and phosphorylated Akt (h, h′) in squamous cell carcinomas (e, f, g, h) and in spindle cell tumours (e′, f′, g′, h′). i) Western blot analysis of squamous, spindle and mixed tumours (tumours showing areas of squamous and spindle cell morphology) showing reduced expression of PTEN and Sprouty 2 in undifferentiated samples in parallel with increased expression of Zeb1 and phosphorylation of Akt and Erk. (j) Quantitative RTPCR analysis of the expression of miR-21 in the same tumor samples used in i.
In this microarray analysis the metastatic tumours are not clustered (denoted by arrows in Fig 3a). To identify possible miRNAs involved in metastasis, the whole selected set of 448 probes was used to discriminate non tumoural skin, squamous and spindle cell carcinomas by supervised clustering. This analysis rendered 14 probes corresponding to miR-23, miR-29*, miR-19a, miR-21, miR-105 and miR-135b discriminating metastatic and non-metastatic primary tumors (Fig 3b). Of note, although qRT-PCR analyses showed no significant differences between SCCs and SpCCs respect to miRNAs regulated by p53 family or those belonging to the miR-200 family (Supp Fig 1b), the overall expression of miR-200 family members was decreased in SpCCs compared to SCCs (Fig 3c). Additionally, to further support the miRNA microarray analyses, we used the gene expression microarray data to monitor the expression of genes predicted to be targets of miR-21, miR-29 and miR-105 (using miRgen38) in unsupervised manner. In all the cases (Supp Fig 1c, d, e), we found that metastatic tumours were grouped together, reinforcing the relevance of these miRNAs in the process.
Given its widely reported involvement in multiple oncogenic processes including metastasis39, we focused our subsequent studies on miR-21. The upregulation of mature miR-21 expression, confirmed by qRT-PCR analysis, was observed in an independent set of primary tumours in association with metastasis and spindle morphology, but not with the genotype of the tumour (Fig 3d).
The miR-21 is known to target and down-regulate the expression of multiple tumor suppressors such as Sprouty2 (Spry2)40 and Pten41,42, which lead to the upregulation of Erk/MAPK and PI3K/Akt axis respectively. In squamous cell carcinomas, the expression of Spry2 and Pten was exclusively found in the more differentiated areas (Fig 3e, g), being almost undetectable in spindle tumours (Fig 3e’, g’). Accordingly, reduced expression of phosphorylated, active Akt and Erk was observed in SCC (Fig 3f, h), whilst in SpCC generalized expression of phosphorylated Akt and Erk was detected (Fig 3f’, h’). Biochemical analysis by western blot (Fig 3i) and qRT-PCR (Fig 3j) of different tumour samples corresponding to SCCs, SpCCs and tumours that histologically showed a mixture of both phenotypes, demonstrated a clear correlation between decreased Pten and Spry2 expression, activation of Akt and Erk (Fig 3i) and increased miR-21 (Fig 3j) expression in undifferentiated tumors.
miR-21 expression confers tumour aggressiveness and metastatic potential
To further confirm the possible role of miR-21 in EMT and metastasis, we first monitored the effect of increased miR-21 expression in the immortalized HaCaT keratinocyte cell line by transfection experiments. The increased expression of miR-21 (Supp Fig 2a) did not affect the cell morphology (Supp Fig 2b, b’), in spite of reducing Spry2 and Pten expression with a concomitant increase in Akt and Erk phosphorylation (Supp Fig 2c). Furthermore, no significant changes were detected in the expression of Snail, FoxC2 or Twist (Supp Fig 2c) and neither decrease of E-cadherin nor increase in Vimentin were observed in transfected cells compared with controls (Supp Fig 2d, d’, e, e’). These results indicate that the augmented expression of miR-21 is insufficient to drive EMT in non-transformed cells.
Next, we explored whether the increased expression of miR-21 conferred more aggressive properties to the PB mouse transformed keratinocyte cell line. These cells are derived from a chemically induced mouse skin papilloma43 and develop well differentiated SCC upon subcutaneous injection into nude mice flanks44. Remarkably, the overexpression of Akt in these cells conferred more aggressive characteristics demonstrated by reduced tumour differentiation, increased angiogenesis and activation of specific biochemical pathways44,45,46. In addition, we also generated cells overexpressing Snail and both Snail and miR-21, for comparison. Transfection of Snail in PB cells (Fig 4a) produced a moderate increase in miR-21 expression (Fig 4b), below the levels obtained upon co-transfection of miR-21 and Snail. On the other hand, the forced expression of miR-21, even though not reaching the levels observed in undifferentiated mouse tumours, was able to induce Snai1 expression (Fig 4a, g). Next, we explored the effects of the different transfections in the expression and localization of the epidermal markers keratin K5 and E-cadherin by immunofluorescence. The data revealed that expression of miR-21 (Fig 4d), Snail (Fig 4e), or co-expression of Snail and miR-21 (Fig 4f), reduced the levels of these two proteins in cultured cells compared with control (Fig 4c). Biochemical analyses by western blot (Fig 4g) corroborated the immunofluorescence findings on E-cadherin downregulation, although the effects were more evident upon expression of Snail (Fig 4g). In addition, western blot also showed that miR-21 promoted a moderate increase in vimentin, Snail and very mild in Foxc2, when compared to cells transfected with Snail or Snail and miR-21 (Fig 4g). On the contrary, the induction of Twist, the decrease in Pten and Spry2 and the activation of Akt and ERK were similar in cells transfected with Snail, miR-21 and Snail, or miR-21 alone (Fig 4g). These data indicate that miR-21 can be sufficient to drive some characteristics of EMT and that miR-21 can cooperate with Snail in the process.
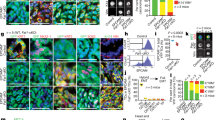
miR-21 expression promotes invasive properties in tumor keratinocytes.
(a, b) Summary of qRT-PCR analyses showing the expression of Snai1 gene (a) and miR-21 (b) in PB keratinocytes and their derivatives upon transfection with the quoted coding plasmids. c-f) Immunofluorescence examples showing the expression of E-cadherin (red) and keratin K5 (green) in PB cells (c) or derivatives transfected with miR-21 (d), Snail (e), or co-transfected with Snail and miR-21 (f). (g) Western blot analysis showing the expression of the quoted proteins in PB cells and transfected derivatives. (h) Quantitative analysis using boyden chamber assays of the migratory (left panel), invasive (mid panel) capacities and percentage of invasion (right panel) of parental PB keratinocytes or transfected derivatives (* denotes p ≤ 0.05; ** denotes p ≤ 0.01). i, i’) Examples of tumours (H&E staining) produced upon subcutaneous injection of mirVec (empty vector; i) or miR-21 transfected PB keratinocytes; (j) summary of the different phenotypes of the tumours upon subcutaneous injection (n = 8 in each case). k) Example of lung metastasis (H&E) observed in mice bearing subcutaneous injection of tumours in miR-21 transfected PB tumour keratinocytes. Inset shows a higher magnification of the metastasis.
Since EMT affects the migratory and invasive potential of transformed epidermal cells, we monitored these properties in cells expressing the above commented constructs in vitro. Our data (Fig 4h) demonstrated that the expression of miR-21 or Snail increased the migratory and invasive properties of PB keratinocytes. However, the co-expression of miR-21 and Snail did not further increase migration and invasion (Fig 4h).
Next we explored whether the miR-21 mediated changes observed in vitro could also affect the tumourigenic properties of the PB cells in vivo upon subcutaneous injection in nude mice. Although we did not observe a significant difference in the appearance or growth rate of tumours between PBmir-21 and PBmirVec cells (not shown), we found that the expression of miR-21 dramatically changed the histopathology of the tumors, from a well differentiated SCC characteristics (Fig 4i, j) to a more undifferentiated with SpCC morphology (Fig 4i’, j). Further, mice injected with PBmiR-21 cells, but not those bearing PBmiRVec cells, showed metastatic signs in the proximal ganglia and in the lungs (3 out of 5 and 0 out of 5 injected mice, respectively). Collectively these data demonstrate that the expression of miR-21 conferred increased aggressiveness and metastatic properties to tumour keratinocytes.
miR-21 expression is required for metastasis in Trp53-deficient tumor cells
The above described results indicate that miR-21 contributes to EMT and metastasis. To analyze whether these processes are dependent on miR-21 in fully transformed cells, we established a cell line (named 940T) from a spontaneous primary SpCC from a RbΔEC; p53ΔEC mouse with lung metastasis. The blockade of miR-21 in these cells using miR-ZIP-21 constructs (Fig 5a) produced increased expression of Spry2 and Pten and the concomitant reduced activation of Erk and Akt (Fig 5b). This also resulted in increased E-cadherin expression (Fig 5b, c, c’) and reduced proliferation in vitro (Fig 5d). The blockade on miR-21 in the RbΔEC; p53ΔEC tumour derived cells also resulted in reduced migration and invasiveness in vitro (Fig 5e).
miR-21 blockade inhibits metastasis development.
(a) qRT-PCR analysis showing the expression levels of miR-21 and the reporter marker (COP-GFP) in 940T tumor cells transfected with miR ZIPVec (empty vector) or miR-ZIP-21. (b) Western blot analysis showing the expression of the quoted proteins in miR ZIPVec and miR-ZIP21-transfected 940T tumor keratinocytes. c, c’) Immunofluorescence examples showing the expression of E-cadherin (red) in miR-ZIPVec (c) and miR-ZIP-21-transfected (c’) 940T tumour keratinocytes. (d) Immunofluorescence examples and summary of three independent experiments showing BrdU incorporation in miR-ZIPVec and miR-ZIP-21-transfected 940T tumor keratinocytes. Bar = 25 µm. (e) Quantitative analysis using boyden chamber assays of the migratory (upper panel), invasive (mid panel) capacities and percentage of invasion (lower panel) of miR-ZIPVec and miR-ZIP-21-transfected 940T tumour keratinocytes. (f) Growth of tumours produced by subcutaneous injection of miR-ZIPVec (black squares) and miR-ZIP-21-transfected (red circles) 940T tumour keratinocytes (n = 6). g, g’) Examples of lung sections (H&E stained) upon intravenous injection of miR-ZIPVec (g’) and miR-ZIP-21-transfected (g’) 940T tumour keratinocytes. Bar = 1 cm.
Of note, the subcutaneous injection of miR-ZIPVec or miR-ZIP-21 transfected cells in immunodeficient mice did not produce major changes in tumour aggressiveness or histopathology (not shown), although a partial reduction in tumour growth was detected at early stages (Fig 5f). Finally, we monitored whether the miR-21 blockade could affect metastatic behaviour of highly transformed cells. To this, miR-ZIPVec or miR-ZIP-21 cells were injected in the tail vein and the formation of lung metastasis was analyzed. We observed that, in contrast with the control cells (Fig 5g) that produced the development of a large number of lung metastasis (48±10 per lung) in all the injected animals (8/8), miR-ZIP-21 transfected cells only produced one small micrometastasis in one of the injected mice (1/8). These results clearly demonstrated that miR-21 is necessary to confer metastatic behaviour to p53-deficient mouse skin tumours.
miR-21 expression depends on mTOR and Stat3 activity
Numerous reports have indicated that p53 can modulate miRNA expression17,47,48. However, miR-21 has not been reported to be transcriptionally modulated by p53. Thus it is conceivable that the observed miR-21 increased expression in p53-deficient tumours is attributable to a signalling pathway induced by p53 loss. Among them, mTOR and Stat3 are induced by p53 deficiency49,50,51,52 and can modulate miR-21 expression53,54,55. We thus analyzed the possible correlation of mTOR and Stat3 activity and miR-21 expression in a series of spontaneous tumours derived from p53ΔEC and RbΔEC; p53ΔEC mice. We found that the increased activity of mTOR, analyzed by phosphorylated S6 ribosomal protein and active Stat3, determined by tyrosine-phophorylated Stat3, were found predominantly in SpCC (Fig 6a) in close correlation with increased miR-21 expression (Fig 6b). We thus used the tumour derived 940T cell line to analyze the effects of mTOR and Stat3 signalling inhibitors. Compared with an immortalized mouse keratinocyte cell line (CoCa56), the levels of miR-21 were significantly upregulated in 940T cells (Fig 6c). These upregulated levels were decreased by the treatment with mTOR or Stat3 inhibitors and significantly dropped by the combined treatment of 940T cells with both inhibitors.
The expression of miR-21 is modulated by mTOR and Stat3 in spontaneous tumors.
(a) Western blot showing the expression of phosphorylated S6, phosphorylated Stat3 and phosphorylated Akt in skin and tumours samples of the quoted genotype and phenotype. (b) qRT-PCR showing the expression levels of miR21 in the same tumors showed in a. (c) qRT-PCR analysis of miR-21 expression levels in non-transformed mouse immortalized keratinocytes (CoCa) and in 940T tumour keratinocytes untreated or treated with the quoted inhibitors for 48h.
miR-21 expression is increased in human metastatic lung cancer
The above explained results prompted us to analyze if similar process could also take place in human cancer samples. We have previously demonstrated that p53ΔEC and RbΔEC; p53ΔEC derived tumours displayed a significant overlap with multiple human tumours characterized by augmented aggressiveness and metastatic development26. These included multiple epithelial cancers from different tissues of origin, such as breast and lung26. In particular, we observed that in non small cell lung cancer (NSCLC) a 20-gene signature derived from mouse p53-deficient tumours26, can stratify human patients and predict overall survival from a external dataset of human lung adenocarcinoma samples (Supp Fig 3A)57. Moreover miR-21 has been considered an oncogene in human lung cancer58,59,60. In agreement, in two external datasets of lung cancer61,62 the increased expression of miR-21 is also predictive of overall survival (Supp Fig 3b, c).
These connections prompted us to analyze whether the findings relative to miR-21 in mouse samples could be extensive to human lung cancers. To this, we studied the expression of miR-21 in a set of human lung tumours of known p53 status as well as other clinical and molecular characteristics, including gene expression profiling63. We observed a significant association between increased miR-21 and TP53 gene mutations (Fig 7a), but not with the presence of KRAS mutations (Fig 7b). Notably, there was a significant association between such increased miR-21 levels and the presence of distant metastasis in these human tumours (Fig 7c) and a significant correlation between miR-21 and the expression levels of human SPRY2 and PTEN genes (Fig 7d).
miR-21 expression in human lung cancers.
(a-c) Box plots showing the miR-21 expression in human lung carcinomas63 according to TP53 gene status (a), KRAS gene status (b) or the presence or absence of distant metastasis (c). (d) Correlation between miR-21 levels and the expression of SPRY2 or PTEN genes in human lung cancer samples63.
Discussion
The tumour suppressor p53, plays a pivotal role in the onset and development of a widely range of human cancers affecting more than 50% of human patients. This makes the p53 pathway one of the main and most significant targets for the chemotherapeutic drug design. In most cases, p53 mutations are missense, which have been considered to provide a gain of function, conferring a more aggressive behaviour to the tumours. This is, in part, exemplified by knock out models: p53 deficient mice display a restricted pattern of tumors and they are rarely metastatic, whereas knock in models reproducing human missense mutants display broader types of tumours and metastasis10. However, truncating mutations in TP53, leading to complete loss of p53, are profuse in certain aggressive human tumours, such in breast cancer bearing mutated BRCA1 gene64.
In multiple solid tumours, metastasis is preceded by EMT events, which allow the cells to repress epithelial characteristics acquiring mesenchymal ones and conferring migratory and invasive properties65. We have characterized a close association of such EMT events and the metastatic properties in our mouse models. Indeed, we found that metastatic potential is increased in tumours that have undergone EMT processes. The molecular events that govern the process are similar in our and in other reported models and lie on an aberrant TGFβ signalling that leads to the expression of master regulatory genes such as Snai1, Twist, Zeb1, Zeb2 and FoxC22. The connection between EMT and p53 has been widely reported and has been attributed to multiple signalling pathways, such as crosstalk with Smads66,67. Remarkably this may help to explain the poor prognosis displayed by p53 mutant tumours6,7. More recently it has been shown that p53 interferes with EMT processes through specific miRNA expression, in particular by the induction of miR-34 or miR-200 families68,69,70. Of note, in our microarray experiments miR-34 did not discriminate between control and tumour samples, probably due to a reduced expression of this miRNA in normal skin samples (Supp Fig 1 and data not shown). On the other hand, although no unique miR-200 family member discriminated between SCC and SpCC (Supp Fig 1), the overall expression of these miRNAs were decreased in metastasis prone samples. Thus, our data reinforce the role of p53 as a potential regulator of this miRNA family.
We found an important deregulation of miRNAs in tumours affecting those involved in epidermal development32,33 and metastasis71,72. However, upon highly restrictive analyses, few miRNAs were associated with the spindle morphology of the tumours. Among them, we observed a good correlation with increased miR-21 expression in undifferentiated SpCC samples and increased metastatic potential. This is in agreement with the reported increased miR-21 expression in human tumours bearing mutated TP53 and displaying distant metastasis39. Our data are also in agreement with the reduced susceptibility to chemical carcinogenesis in skin due to in vivo loss of miR-2173. The functions of miR-21 in oncogenesis and metastasis have been associated with its ability to target multiple tumour suppressors, including PTEN and Spry1/2 leading to increased Akt and MAPK activities41,42,73. In agreement, we could correlate such increased Akt and MAPK activities with the increased expression of miR-21 and EMT processes (Fig 2). Notably, increased expression of miR-21 was unable to promote transformation or EMT in immortalized keratinocytes (Supp Fig 2), but contributed to the acquisition of metastatic properties in otherwise poorly metastatic transformed PB cells and promoted poorly differentiated tumours. This finding is in agreement with our previous data showing that increased Akt activity in these cells confers tumor aggressiveness and angiogenesis44,45. In agreement, the in vivo elimination of miR-21 does not promote tumour development but confers increased sensitivity to chemical carcinogenesis protocols73.
In summary, it is conceivable that other mechanisms could contribute to the metastatic capacities of p53-deficient tumours. TGFβ and p53 have been suggested as potential mechanisms for the deregulation of miRNAs18,74 and these small non coding RNAs have been widely involved in oncogenesis and in metastasis promotion28,71,72. Notably miR-21 can be induced by TGFβ and contributes to increased migration in HaCaT cells during wound healing75. In this regard, we found that overexpression of Snail, which is a target of TGFβ-mediated EMT76, also induced miR-21 although to a moderate level. Of note, we also found that S6 phosphorylation, a well known effect of mTOR activation, was increased in the undifferentiated, metastasis-prone, primary tumours. This could be attributed to the loss of p53 functions49 and can contribute to increased TGFβ activity in tumours77,78,79. The possible cooperative roles between TGFβ signalling and miR-21 in EMT as well as the possible roles of TGFβ in inducing this miRNA would deserve future research.
Regarding the possible mechanisms leading to the upregulation of miR-21 expression, it has been shown that miR-21 expression is induced by activated Stat353 and by PI3K- and Erk-dependent signalling80. Notably, Stat3 and mTOR pathways are activated by p53 loss49,50,51,52,81 and are also functionally interconnected82,83. We observed that tumours showing increased activity of mTOR and/or Stat3 also displayed increased miR-21 levels. The use of inhibitors against these two pathways indicated that the induction of miR-21 could be attributed, in this system, to their coordinated function. This observation opens the possibility that these inhibitors could be considered of particular relevance to prevent metastatic spreading in the mouse tumours, which undoubtedly merits future investigation.
Our findings, demonstrating that miR-21 expression was elevated in human lung tumours bearing mutated TP53 and displaying distant metastasis, are of a particular relevance. This observation is in agreement with previous reports indicating its role in lung cancer58,59. Here, using a series of molecularly characterized human tumours we could establish that such increased expression is correlated with decreased expression of SPRY2 and PTEN. Therefore the oncogenic and metastatic-prone activities of miR-21 are not exclusively associated with SCCs, but could be a more generalized tumoural process. The validation of the molecular events contributing to the increased expression of miR-21 in vivo are also of a particular relevance in the design of potential therapies against metastatic spreading of primary human lung tumours showing increased miR-21 expression.
Methods
Mouse and Histological procedures
Mouse models have been previously described20. Histological analyses were performed as previously reported in formalin fixed paraffin embedded samples20,24,84. For immunohistochemistry, high temperature antigen unmasking technique (10-minute microwaving of slides in 0.01M citrate buffer) was used after deparaffinization to enhance the staining. Sections were then incubated with 5% horse serum for 30 minutes to block the Fc receptor in tissue and then washed three times with sterile PBS (pH 7.5) prior to incubation with the appropriate primary antibodies diluted in PBS/BSA. Antibodies are listed on Supplementary Table 2. Biotin-conjugated secondary antibodies were purchased from Jackson ImmunoResearch and used at 1/5000 dilution. For IHC, signal was amplified using avidin-peroxidase (ABC elite kit Vector) and peroxidase was visualized using diaminobenzidine as a substrate (DAB kit Vector). Control slides were obtained by replacing primary antibodies with PBS (data not shown). All the animal experiments were approved by the Animal Ethical Committee (CEEA) and conducted in compliance with Centro de Investigaciones Energéticas, Medioambientales y Tecnológicas (CIEMAT) guidelines.
Cell culture and Chemicals
HaCaT and PB cells were cultured in DMEM containing 10% FBS. Transfection experiments were performed as described85 using plasmids coding for human miR-21 (NKI library86) and Snail87. The selection of the transfected cells was performed for at least 15 days in appropriate antibiotic containing medium and 40-60 pooled clones were used. Migration and invasiveness were monitored using boyden chambers (BD-Biosciences) following the manufacturer’s recommendations. Tumour cells were obtained from a primary tumour of an RbΔEC; p53ΔEC mouse bearing lung metastasis upon trypsin digestion and disaggregation. Tumour cells were grown as reported for wild type mouse epidermal keratinocytes22. The non transformed immortalized CoCa keratinocyte cell line56 was cultured in CnT7 (CellnTech Genycell). miR-ZIP-21 was purchased from System Bioscience and transfected in the tumour cells using Fugene (Roche). Indirect immunofluorescence including BrdU detection was performed as previously described85,88,89. Pharmacological inhibition was performed incubating cultures for 48 hours in the presence of Rapamycin (50 nM) and/or Tyrphostin AG940 (100 µM) (Sigma).
Positron Emission Tomography (PET)
In vivo imaging by positron emission tomography using 2-[18F]Fluoro-2-deoxy-d-glucose (FDG), as tracer was performed as previously reported84. Images were taken in an axial plane avoiding brain and heart to eliminate the possible background due to the high incorporation of FDG in these organs.
External microarray datasets of human cancer
To stratify human lung cancer patients according the miR-21 expression, two different external datasets61,62 were used. To determine the possible genomic similarities between human lung and mouse p53-deficient epidermal tumours, we followed our previously reported approach26. In particular, we determined whether a 20-gene signature characteristic of mouse p53-deficient tumours, which can identify human tumours with poor outcome from breast cancer, astrocytoma and multiple myeloma26, can also help to determine the prognosis of human lung cancer patients. To this, we obtained the corresponding 20-gene score in each human tumour sample from a comprehensive genomic study of human lung adenocarcinomas downloaded from GEO57. This allows the classification of the samples in three groups depending on this score26 and analyze the association of these clusters of patients with disease specific survival.
Gene expression of human lung tumors used to monitor microRNA expression have been previously reported63. Tissues were provided by the CNIO Tumour Bank Network, in collaboration with different Spanish hospitals. The study was approved by the corresponding institutional review boards and ethics committees and informed consent was obtained from each patient.
Microarray analyses
The microarray analysis using Affymetrix mouse gene chip 430A of total RNA extracted from tumours has been previously reported26. The microRNA analysis was performed using miRCURY LNATM v 11.0 platform. This contains more than 1700 capture probes, covering all human, mouse and rat miRNAs annotated in miRBase 11.0, as well as all viral microRNAs. In addition, this array contains capture probes for 435 new miRPlus™ human miRNAs. The probes allowing classification between normal and tumoural samples was performed by sequential SAM (FDR q<0.01) and t Test (Bonferroni corrected p<0.01) analyses. The supervised analysis discriminating SCC and SpCC was performed in a similar manner using the previous dataset.
Western blot
Ground mortar obtained extracts from tumour or skin samples, or pelleted keratinocytes were lysed by freeze-thawing cycles in lysis buffer (200 mM HEPES pH 7’9, 25% glycerol, 400 mM NaCl, 1 mM EDTA, 1 mM EGTA, 1 μg/mL aprotinin, 1 μg/mL leupeptin, 1 mM PMSF, 20 mM NaF, 1 mM NaPPi, 1 mM Na3VO4, 2.5 mM DTT) and centrifuged to obtain supernatant containing total protein. 35 µg protein per sample were resolved in SDS-PAGE gels and transferred to nitrocellulose membranes (Amersham). Membranes were blocked with 5% non-fat milk diluted in TBS and incubated with the appropriate antibodies diluted in TBS-T 0.5% BSA. Secondary antibodies were purchased from Jackson ImmunoResearch and used at 1/5000. Super Signal West Pico Chemiluminscence Substrate (Pierce) was used according to the manufacturer’s recommendations to visualize the bands. In all cases Actin was used for loading control. The antibodies are as follows Actin (Santa Cruz sc-1616 diluted 1/500), Akt (Santa Cruz sc-1619 diluted 1/500), E-cadherin (Transduction Lab 610182, diluted 1/2500) Erk (Santa Cruz sc-154, diluted 1/500) FOX C2 (AbCam 5060 diluted 1/500) Keratin K5 (Covance PRB-160P, diluted1/500), p53 (NovoCastra NCL-p53-CM5p, diluted 1/500), P-Akt (ser 473) (Cell Signaling 4058S, diluted 1/200) P-Erk (Santa Cruz sc-7383, diluted 1/200), P-S6 (Cell Signaling 2211, diluted 1/250), P-Stat3(Tyr 705) (Cell Signaling 9131, diluted 1/250), PTEN (Santa Cruz sc-6818, diluted 1/500), Snail (AbCam 17732, diluted 1/1000), Sprouty 2 (Santa Cruz sc-30049, diluted 1/500) Stat 3 (Cell Signaling 4904, diluted 1/500), Twist (AbCam ab50581, diluted 1/5000 and Abnova H00007291-M01, diluted 1/300) Vimentin (Biogenex, diluted 1/200), Zeb1 (Santa Cruz 10572, diluted 1/100), anti BrdU (AbCam ab6326, diluted 1/100; and Roche 1170376, diluted 1/100), Dicer (AbCam ab14201, diluted 1/100), Drosha (AbCam ab12286, diluted 1/100) and DGCR8 (AbCam ab36865, diluted 1/200).
Quantitative real-time PCR from mRNA and miRNA
Purification of total RNA including miRNA was done using the miRNeasy Mini Kit (Qiagen). RNA integrity was tested using Bioanalyzer (Agilent). For gene expression, reverse transcription was done with the Omniscript® Reverse Transcription kit (Qiagen) using oligo-dT primers. Real-time PCR was performed using gene specific primers (Supplementary Table 4) and the SYBR Green system (Applied Biosystems). β- glucuronidase (GUSB) was used as normalizing, housekeeping gene. TaqMan® MicroRNA Assays (Applied Biosystems) were used to quantify miRNAs in samples according to the manufacturer instructions with the TaqMan® Universal PCR Master Mix reagent kit (Applied Biosystems). Normalization was performed using U6 as housekeeping small RNA. qRT-PCR for mRNA and miRNA was performed in an ABI 7500fast Real-Time PCR System.
The primers used for qPCR are:
GUS B:
Forward 5′… GAGGATCAACAGTGCCCATT…3′
Reverse 5′… CAGCCTCAAAGGGGAGGT…3′
Murine Snai1:
Forward 5′…CACCTCCAGACCCACTCAGAT…3′
Reverse 5′…CCTGAGTGGGGTGGGAGC…3′
COP-GFP:
Forward 5′…CGGCTTCTACCACTTCGGC…3
Reverse 5′…TTGTTGATGGCGTGCAGGA…3′
References
Moreno-Bueno, G., Portillo, F. & Cano, A. Transcriptional regulation of cell polarity in EMT and cancer. Oncogene 27, 6958–6969 (2008).
Peinado, H., Olmeda, D. & Cano, A. Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 7, 415–428 (2007).
Yang, M. H. et al. Bmi1 is essential in Twist1-induced epithelial-mesenchymal transition. Nat Cell Biol 12, 982–992 (2010).
Egeblad, M., Nakasone, E. S. & Werb, Z. Tumors as organs: complex tissues that interface with the entire organism. Dev Cell 18, 884–901 (2010).
Chiang, A. C. & Massague, J. Molecular basis of metastasis. N Engl J Med 359, 2814–2823 (2008).
Petitjean, A. et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat 28, 622–629 (2007).
Toledo, F. & Wahl, G. M. Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nat Rev Cancer 6, 909–923 (2006).
Muller, P. A. et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 139, 1327–1341 (2009).
Adorno, M. et al. A Mutant-p53/Smad complex opposes p63 to empower TGFbeta-induced metastasis. Cell 137, 87–98 (2009).
Lozano, G. The oncogenic roles of p53 mutants in mouse models. Current opinion in genetics & development 17, 66–70 (2007).
Flores, E. R. et al. Tumor predisposition in mice mutant for p63 and p73: evidence for broader tumor suppressor functions for the p53 family. Cancer Cell 7, 363–373 (2005).
Wang, Z. et al. Targeting miRNAs involved in cancer stem cell and EMT regulation: An emerging concept in overcoming drug resistance. Drug Resist Updat 13, 109–118 (2010).
Liu, C. & Tang, D. G. MicroRNA Regulation of Cancer Stem Cells. Cancer Res 71, 5950–5954 (2011).
White, N. M. et al. Metastamirs: a stepping stone towards improved cancer management. Nat Rev Clin Oncol 8, 75–84 (2011).
He, X., He, L. & Hannon, G. J. The guardian's little helper: microRNAs in the p53 tumor suppressor network. Cancer Res 67, 11099–11101 (2007).
Raver-Shapira, N. & Oren, M. Tiny actors, great roles: microRNAs in p53's service. Cell Cycle 6, 2656–2661 (2007).
He, L., He, X., Lowe, S. W. & Hannon, G. J. microRNAs join the p53 network--another piece in the tumour-suppression puzzle. Nat Rev Cancer 7, 819–822 (2007).
Suzuki, H. I. & Miyazono, K. Dynamics of microRNA biogenesis: crosstalk between p53 network and microRNA processing pathway. J Mol Med (2010).
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature genetics 29, 418–425 (2001).
Martinez-Cruz, A. B. et al. Spontaneous squamous cell carcinoma induced by the somatic inactivation of retinoblastoma and Trp53 tumor suppressors. Cancer Res 68, 683–692 (2008).
Martinez-Cruz, A. B. et al. Spontaneous tumor formation in Trp53-deficient epidermis mediated by chromosomal instability and inflammation. Anticancer research 29, 3035–3042 (2009).
Ruiz, S. et al. Unique and overlapping functions of pRb and p107 in the control of proliferation and differentiation in epidermis. Development 131, 2737–2748 (2004).
Ruiz, S., Santos, M. & Paramio, J. M. Is the loss of pRb essential for the mouse skin carcinogenesis? Cell Cycle 5, 625–629 (2006).
Ruiz, S. et al. Unexpected roles for pRb in mouse skin carcinogenesis. Cancer Res 65, 9678–9686 (2005).
Lara, M. F. et al. p107 acts as a tumor suppressor in pRb-deficient epidermis. Mol Carcinog 47, 105–113 (2008).
Garcia-Escudero, R. et al. Gene expression profiling of mouse p53-deficient epidermal carcinoma defines molecular determinants of human cancer malignancy. Molecular cancer 9, 193 (2010).
Taube, J. H. et al. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proceedings of the National Academy of Sciences of the United States of America 107, 15449–15454 (2010).
Cano, A. & Nieto, M. A. Non-coding RNAs take centre stage in epithelial-to-mesenchymal transition. Trends Cell Biol 18, 357–359 (2008).
Ma, L. & Weinberg, R. A. MicroRNAs in malignant progression. Cell Cycle 7, 570–572 (2008).
Gregory, P. A., Bracken, C. P., Bert, A. G. & Goodall, G. J. MicroRNAs as regulators of epithelial-mesenchymal transition. Cell Cycle 7, 3112–3118 (2008).
Inui, M., Martello, G. & Piccolo, S. MicroRNA control of signal transduction. Nat Rev Mol Cell Biol 11, 252–263 (2010).
Andl, T. et al. The miRNA-Processing Enzyme Dicer Is Essential for the Morphogenesis and Maintenance of Hair Follicles. Curr Biol 16, 1041–1049 (2006).
Yi, R. et al. Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nature genetics 38, 356–362 (2006).
Bracken, C. P., Gregory, P. A., Khew-Goodall, Y. & Goodall, G. J. The role of microRNAs in metastasis and epithelial-mesenchymal transition. Cell Mol Life Sci 66, 1682–1699 (2009).
Bracken, C. P. et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 68, 7846–7854 (2008).
Kumar, M. S., Lu, J., Mercer, K. L., Golub, T. R. & Jacks, T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nature genetics 39, 673–677 (2007).
Su, X. et al. TAp63 suppresses metastasis through coordinate regulation of Dicer and miRNAs. Nature 467, 986–990 (2010).
Megraw, M., Sethupathy, P., Corda, B. & Hatzigeorgiou, A. G. miRGen: a database for the study of animal microRNA genomic organization and function. Nucleic acids research 35, D149–155 (2007).
Selcuklu, S. D., Donoghue, M. T. & Spillane, C. miR-21 as a key regulator of oncogenic processes. Biochem Soc Trans 37, 918–925 (2009).
Sayed, D. et al. MicroRNA-21 targets Sprouty2 and promotes cellular outgrowths. Molecular biology of the cell 19, 3272–3282 (2008).
Pezzolesi, M. G., Platzer, P., Waite, K. A. & Eng, C. Differential expression of PTEN-targeting microRNAs miR-19a and miR-21 in Cowden syndrome. Am J Hum Genet 82, 1141–1149 (2008).
Meng, F. et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133, 647–658 (2007).
Yuspa, S. H. et al. Cultivation and characterization of cells derived from mouse skin papillomas induced by an initiation-promotion protocol. Carcinogenesis 7, 949–958 (1986).
Segrelles, C. et al. Functional roles of Akt signaling in mouse skin tumorigenesis. Oncogene 21, 53–64 (2002).
Segrelles, C. et al. Akt mediates an angiogenic switch in transformed keratinocytes. Carcinogenesis 25, 1137–1147 (2004).
Segrelles, C. et al. Molecular determinants of Akt-induced keratinocyte transformation. Oncogene 25, 1174–1185 (2006).
Hermeking, H. p53 enters the microRNA world. Cancer Cell 12, 414–418 (2007).
Boominathan, L. The guardians of the genome (p53, TA-p73 and TA-p63) are regulators of tumor suppressor miRNAs network. Cancer metastasis reviews 29, 613–639 (2010).
Feng, Z. & Levine, A. J. The regulation of energy metabolism and the IGF-1/mTOR pathways by the p53 protein. Trends Cell Biol (2010).
Feng, Z., Zhang, H., Levine, A. J. & Jin, S. The coordinate regulation of the p53 and mTOR pathways in cells. Proceedings of the National Academy of Sciences of the United States of America 102, 8204–8209 (2005).
Lin, J., Tang, H., Jin, X., Jia, G. & Hsieh, J. T. p53 regulates Stat3 phosphorylation and DNA binding activity in human prostate cancer cells expressing constitutively active Stat3. Oncogene 21, 3082–3088 (2002).
Lin, J. et al. Modulation of signal transducer and activator of transcription 3 activities by p53 tumor suppressor in breast cancer cells. Cancer Res 62, 376–380 (2002).
Loffler, D. et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 110, 1330–1333 (2007).
Darido, C. et al. Targeting of the Tumor Suppressor GRHL3 by a miR-21-Dependent Proto-Oncogenic Network Results in PTEN Loss and Tumorigenesis. Cancer Cell 20, 635–648 (2011).
Kim, Y. J. et al. PTEN Modulates miR-21 Processing via RNA-Regulatory Protein RNH1. PLoS One 6, e28308 (2011).
Segrelles, C. et al. Establishment of a murine epidermal cell line suitable for in vitro and in vivo skin modelling. BMC dermatology 11, 9 (2011).
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med 14, 822–827 (2008).
Volinia, S. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proceedings of the National Academy of Sciences of the United States of America 103, 2257–2261 (2006).
Hatley, M. E. et al. Modulation of K-Ras-dependent lung tumorigenesis by MicroRNA-21. Cancer Cell 18, 282–293 (2010).
Seike, M. et al. MiR-21 is an EGFR-regulated anti-apoptotic factor in lung cancer in never-smokers. Proceedings of the National Academy of Sciences of the United States of America 106, 12085–12090 (2009).
Yu, S. L. et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell 13, 48–57 (2008).
Raponi, M. et al. MicroRNA classifiers for predicting prognosis of squamous cell lung cancer. Cancer Res 69, 5776–5783 (2009).
Angulo, B. et al. Expression signatures in lung cancer reveal a profile for EGFR-mutant tumours and identify selective PIK3CA overexpression by gene amplification. The Journal of pathology 214, 347–356 (2008).
Holstege, H. et al. High incidence of protein-truncating TP53 mutations in BRCA1-related breast cancer. Cancer Res 69, 3625–3633 (2009).
Peinado, H., Portillo, F. & Cano, A. Transcriptional regulation of cadherins during development and carcinogenesis. Int J Dev Biol 48, 365–375 (2004).
Piccolo, S. p53 regulation orchestrates the TGF-beta response. Cell 133, 767–769 (2008).
Cordenonsi, M. et al. Links between tumor suppressors: p53 is required for TGF-beta gene responses by cooperating with Smads. Cell 113, 301–314 (2003).
Chang, C. J. et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol 13, 317–323 (2011).
Kim, N. H. et al. A p53/miRNA-34 axis regulates Snail1-dependent cancer cell epithelial--mesenchymal transition. J Cell Biol 195, 417–433 (2011).
Kim, T. et al. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J Exp Med 208, 875–883 (2011).
Dykxhoorn, D. M. MicroRNAs and metastasis: little RNAs go a long way. Cancer Res 70, 6401–6406 (2010).
Zhang, H., Li, Y. & Lai, M. The microRNA network and tumor metastasis. Oncogene 29, 937–948 (2010).
Ma, X. et al. Loss of the miR-21 allele elevates the expression of its target genes and reduces tumorigenesis. Proceedings of the National Academy of Sciences of the United States of America 108, 10144–10149 (2011).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Yang, X. et al. miR-21 promotes keratinocyte migration and re-epithelialization during wound healing. Int J Biol Sci 7, 685–690 (2011).
Peinado, H., Quintanilla, M. & Cano, A. Transforming growth factor beta-1 induces snail transcription factor in epithelial cell lines: mechanisms for epithelial mesenchymal transitions. J Biol Chem 278, 21113–21123, 10.1074/jbc.M211304200 M211304200 [pii] (2003).
Lamouille, S. & Derynck, R. Cell size and invasion in TGF-beta-induced epithelial to mesenchymal transition is regulated by activation of the mTOR pathway. J Cell Biol 178, 437–451 (2007).
Wu, L. & Derynck, R. Essential role of TGF-beta signaling in glucose-induced cell hypertrophy. Dev Cell 17, 35–48 (2009).
Wempe, F. et al. Inactivation of sestrin 2 induces TGF-beta signaling and partially rescues pulmonary emphysema in a mouse model of COPD. Dis Model Mech 3, 246–253 (2010).
Frezzetti, D. et al. Upregulation of miR-21 by Ras in vivo and its role in tumor growth. Oncogene 30, 275–286 (2011).
Feng, Z. et al. The regulation of AMPK beta1, TSC2 and PTEN expression by p53: stress, cell and tissue specificity and the role of these gene products in modulating the IGF-1-AKT-mTOR pathways. Cancer Res 67, 3043–3053 (2007).
Onda, H. et al. Tsc2 null murine neuroepithelial cells are a model for human tuber giant cells and show activation of an mTOR pathway. Mol Cell Neurosci 21, 561–574 (2002).
Ma, J. et al. Mammalian target of rapamycin regulates murine and human cell differentiation through STAT3/p63/Jagged/Notch cascade. J Clin Invest 120, 103–114 (2010).
Moral, M. et al. Akt activation synergizes with Trp53 loss in oral epithelium to produce a novel mouse model for head and neck squamous cell carcinoma. Cancer Res 69, 1099–1108 (2009).
Paramio, J. M., Segrelles, C., Ruiz, S. & Jorcano, J. L. Inhibition of protein kinase B (PKB) and PKCzeta mediates keratin K10-induced cell cycle arrest. Mol Cell Biol 21, 7449–7459 (2001).
Voorhoeve, P. M. et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 124, 1169–1181 (2006).
Peinado, H. et al. Snail and E47 repressors of E-cadherin induce distinct invasive and angiogenic properties in vivo. J Cell Sci 117, 2827–2839 (2004).
Paramio, J. M. & Jorcano, J. L. Assembly dynamics of epidermal keratins K1 and K10 in transfected cells. Exp Cell Res 215, 319–331 (1994).
Paramio, J. M. et al. Modulation of cell proliferation by cytokeratins K10 and K16. Mol Cell Biol 19, 3086–3094 (1999).
Acknowledgements
Grant support: Ministerio de Ciencia e Innovación (MICINN) grants SAF2008-0121 and SAF2011-26122-C02-01, Comunidad Autónoma de Madrid Oncocycle Program Grants S2006/BIO-0232 and S2010/BMD-2470, Ministerio de Sanidad y Consumo grant ISCIII-RETIC RD06/0020/0029 and from Fundación Sandra Ibarra to JMP, RETIC RD06/0020/0062 (to MSC) and RD06/0020/0111 (to FP). The excellent technical support by Pilar Hernández in histology and the personnel of the CIEMAT Animal Facility are specially acknowledged. We also acknowledge the human lung tumor collection provided by the CNIO Tumour Bank Unit. Plasmids coding for human and mouse Snail are a generous gift form Dr. A Cano (Universidad Autónoma de Madrid, Spain)
Author information
Authors and Affiliations
Contributions
J.M.P directed all aspects of the p53 metastasis project. M.S and J.M.P. designed the experiments, R.G-E and J.M.P. analyzed the data and J.M.P. wrote the manuscript. O. B., A.B. M-C., C.C., C.S., C.L., A.B., C.S-L., X.A., A.M., B.P., J.M.A. and M.S. performed the experiments, T. G., Performed the PET-CT analyses. R.G-E. and M.D. supervised the gene array data collection and pre-analysis processing, F.P., M.S., MS-C. Analyzed the results and provided reagents.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplymentary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Bornachea, O., Santos, M., Martínez-Cruz, A. et al. EMT and induction of miR-21 mediate metastasis development in Trp53-deficient tumours. Sci Rep 2, 434 (2012). https://doi.org/10.1038/srep00434
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00434
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.