Abstract
Teleost fishes comprise more than half of all vertebrate species, yet genomic data are only available for 0.2% of their diversity. Here, we present whole genome sequencing data for 66 new species of teleosts, vastly expanding the availability of genomic data for this important vertebrate group. We report on de novo assemblies based on low-coverage (9–39×) sequencing and present detailed methodology for all analyses. To facilitate further utilization of this data set, we present statistical analyses of the gene space completeness and verify the expected phylogenetic position of the sequenced genomes in a large mitogenomic context. We further present a nuclear marker set used for phylogenetic inference and evaluate each gene tree in relation to the species tree to test for homogeneity in the phylogenetic signal. Collectively, these analyses illustrate the robustness of this highly diverse data set and enable extensive reuse of the selected phylogenetic markers and the genomic data in general. This data set covers all major teleost lineages and provides unprecedented opportunities for comparative studies of teleosts.
Design Type(s) | species comparison design |
Measurement Type(s) | genome assembly |
Technology Type(s) | DNA sequencing |
Factor Type(s) | organism |
Sample Characteristic(s) | Osmerus eperlanus • Borostomias antarcticus • Parasudis fraserbrunneri • Guentherus altivela • Benthosema glaciale • Polymixia japonica • Percopsis transmontana • Typhlichthys subterraneus • Zeus faber • Cyttopsis rosea • Stylephorus chordatus • Bregmaceros cantori • Merluccius polli • Merluccius merluccius • Merluccius capensis • Melanonus zugmayeri • Muraenolepis marmoratus • Trachyrincus scabrus • Trachyrincus murrayi • Mora moro • Laemonema laureysi • Bathygadus melanobranchus • Macrourus berglax • Malacocephalus occidentalis • Phycis blennoides • Phycis phycis • Lota lota • Molva molva • Brosme brosme • Trisopterus minutus • Gadiculus argenteus • Pollachius virens • Melanogrammus aeglefinus • Merlangius merlangus • Arctogadus glacialis • Boreogadus saida • Gadus chalcogrammus • Gadus morhua • Regalecus glesne • Lampris guttatus • Monocentris japonicus • Myripristis jacobus • Holocentrus rufus • Neoniphon sammara • Beryx splendens • Rondeletia loricata • Acanthochaenus luetkenii • Brotula barbata • Lamprogrammus exutus • Carapus acus • Chatrabus melanurus • Thunnus albacares • Lesueurigobius sanzi • Perca fluviatilis • Myoxocephalus scorpius • Sebastes norvegicus • Chaenocephalus aceratus • Symphodus melops • Spondyliosoma cantharus • Antennarius striatus • Selene dorsalis • Helostoma temminkii • Anabas testudineus • Parablennius parvicornis • Chromis chromis • Pseudochromis fuscus • fin • muscle organ • thymus • spleen • blood |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Fueled by recent advances in comparative genomics, teleost fishes are becoming increasingly important research objects in several scientific disciplines, ranging from ecology, physiology and evolution to medicine, cancer research and aquaculture1–7. Genome information from non-model organisms is highly important in these comparative genomic analyses as they represent specific phenotypes that aid in disentangling the common parts of gene sets from those that have evolved as adaptations to specific ecosystems. In a quest to identify the evolutionary origin of the MHC II pathway loss first observed in the Atlantic cod (Gadus morhua)8,9, we applied a single sequencing library procedure to cost-efficiently produce draft assemblies for 66 teleost species, representing all major lineages within teleost fishes10. Since the alternative immune system, characterized by both the lack of MHC II and an expansion of MHC I, has so far only been identified in the Atlantic cod, we sampled the cod-like fishes of the order Gadiformes more densely than other groups, including 27 species of this order. Based on these genome sequence data, we were able to reconstruct the evolutionary history of the sampled lineages, to pinpoint the loss of the MHC II pathway to the common ancestor of all Gadiformes, and to identify several independent expansions in MHC I copy number within and outside the order Gadiformes. While these analyses and results are reported in a companion paper (Malmstrøm et al.11), we here present in greater detail the underlying data sets used for these analyses, including samples, sequencing reads (Data Citation 1), draft assemblies (Data Citation 2), and both mitochondrial and nuclear phylogenetic markers. By providing these data and the applied methodology in a coherent manner we aim to supply the scientific community with a highly diverse, reliable, and easy-to-use genomic resource for future comparative studies.
Our sequencing strategy was chosen on the basis of several pseudo-replicates of the budgerigar (Melopsittacus undulatus) genome12 (Data Citation 3), comprising different combinations of read lengths and coverages to determine the most cost-effective manner to produce genome data of sufficient quality for a reliable determination of gene presence or absence. These budgerigar data sets were furthermore assembled with two of the most used assemblers, the de Bruijn graph based SOAPdenovo13 and the Overlap-Layout-Consensus based Celera Assembler14 to investigate which assembly algorithm performed best on the various data replicates. On the basis of these in silico experiments, all species were sequenced on the Illumina HiSeq2000 platform, aiming for ~15× coverage. The sequenced reads were then quality controlled, error corrected and trimmed before performing assembly with Celera Assembler. The continuity of the assemblies was subsequently assessed through N50 statistics and the assembly quality was evaluated on the basis of gene space completeness of highly conserved genes. The assemblies were further used to identify mitochondrial genome sequences, which we used in combination with previously available sequences of related teleosts to verify the phylogenetic positions of sampled taxa (Data Citations 4 to 124). By recovering all taxa in their expected positions, clustering with conspecific or congeneric sequences where such were available, our phylogenetic analysis corroborates the correct identification of all sampled taxa and the absence of DNA contamination.
Figure 1 illustrates the total workflow, and detailed information for each analysis step is further provided in the Methods section and in Tables 1–7 (available online only). The data sets presented here contain sequencing reads and assembled draft genomes for non-model species adapted to a wide variety of habitats, ranging from the deep sea and tropical coral reefs, to rivers and freshwater lakes. These data sets can be used individually or collectively, as resources for studies such as gene family evolution, adaptation to different habitats, phylogenetic inference of teleost orders, transposons and repeat content evolution as well as many other applications regarding gene and genome evolution in a comparative or model organism framework.
(1) A full overview of species, sample supplier and tissue used for DNA extraction is provided in Table 1 (available online only). (2) The DNA extraction method is also found in Table 1 (available online only). (3) All sequencing libraries were created using the Illumina TruSeq Sample Prep v2 Low-Throughput Protocol. Adaptor indexes are provided in Table 2 (available online only). (4) Sequencing statistics and insert sizes for all species are also listed in Table 2 (available online only). (5) FastQC and SGA PreQC analyses were performed for all read sets prior to assembly. (6) Estimated genome sizes, coverages and assembly statistics for all species are presented in Table 3 (available online only), and accession links are provided in Table 4 (available online only). (7) CEGMA and BUSCO statistics are reported in Table 5 (available online only). (8) GenBank accession numbers and UTG IDs for all mitochondrial genomes used in phylogenetic analyses are provided in Tables 6 and 7 (available online only). (9) The maximum-likelihood phylogeny based on mitochondrial genomes is presented in Fig. 3.
Methods
Sample acquisition and DNA extraction
The majority of samples were taken from validated species (mostly voucher specimens) and were provided by museums or university collections. Some samples were obtained from wild caught specimens, in collaboration with local fishermen. All samples were stored on either 96% ethanol or RNA-later (Ambion). The extraction of genomic DNA was carried out using either EZNA Tissue DNA Kit (Omega Bio-Tek), following the manufacturer’s instructions, or using the ‘High salt DNA extraction’ method as described by Phill Watts (https://www.liverpool.ac.uk/~kempsj/IsolationofDNA.pdf). Detailed information about all samples, including origin, voucher specimen ID and DNA extraction method is provided in Table 1 (available online only).
Fragmentation and library preparation
Genomic DNA samples were diluted to 120 μl (50 ng μl−1) with Qiagen Elution Buffer (Qiagen) if necessary and fragmented to lengths of ~400 bp by sonication using a Covaris S220 (Life Technologies) with the following settings: 200 cycles for 90 s with ω-peak at 105. All sequencing libraries were constructed following the Illumina TruSeq Sample Prep v2 Low-Throughput Protocol.
Sequencing and quality control
All sequencing was performed on an Illumina HiSeq 2000 platform with additional chemicals added to extend the number of cycles, yielding paired reads of 150 bp each. The read quality was then assessed using FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Prior to assembly we used SGA PreQC15 to estimate coverage, per-base error rates, level of heterozygosity, repeat content and genome size in order to assess whether more sequencing would be needed. Some samples were then subjected to a second round of sequencing of the same library. Sequencing statistics are presented in Table 2 (available online only).
Draft genome assembly
The methods used for genome assembly are also described in the Supplementary Note of Malmstrøm et al.11. We expand on these methods here, describing the different parameters and settings in greater detail in order to present a complete overview of our analyses.
All draft genomes were created using Celera Assembler, and the version used was downloaded from the CVS (Concurrent Version System, http://wgs-assembler.sourceforge.net/) repository on January 12th 2013. The program meryl, included in the Celera Assembler package was used to create a database of k-mers from the pairs of sequencing reads. Lower k-mer sizes might not resolve repetitive regions, while higher k-mer sizes might not overlap, leading to a loss of information required to correct the reads. Thus, an intermediate k-mer size of 22 was used for all assemblies. Meryl was run with the following options, where the sequences from the reads were concatenated into a file named ‘reads.fa’: meryl -B -v -m 22 -memory 55000 -threads 16 -C -s reads.fa -o reads
In this command, –B specifies that a k-mer database should be created, and that this should be done using the verbose setting (-v). The –m option denotes the ‘merSize’, while –C specifies that canonical reads (both strands) should be used for creating the k-mer database.
The options –threads and –memory specify the computational resources that meryl can utilize and only influence run-time.
Most of the computational time used by Celera Assembler is required to identify overlap between reads. To reduce analysis time and generate longer input sequences, overlapping paired reads were merged with the software FLASH v1.2 (ref. 16), executed with the following command, where –d denotes the path to the output directory (with the prefix given with the –o option), –r is the read length, –f is the insert size, and –s is the standard deviation of the insert size:flash input_1.fastq input_2.fastq -d. -r 150 -f 290 -s 50 –o output_prefix
Celera Assembler’s merTrim program (see Tørresen et al.17) was used to trim, error correct and remove adapters of all reads. The merTrim program estimates the coverage of the sequencing library by analysing the abundance of k-mers versus the number of k-mers at that abundance. By default, k-mers occurring at a frequency corresponding to at least one fourth of the coverage peak can be used to correct reads with k-mers that occur with a frequency of at most one third of the coverage peak. Reads were trimmed to the largest region containing k-mers with a frequency of more than one third of the coverage peak. The trimming of reads removes sequences not supported by other reads and reduces the possible fragmentation of the assembly. Adaptor sequences are not part of the genome and could lead to assembly fragmentation in the same way as repeated regions would. To remove adaptor sequences and other unsupported sequences from the read data, merTrim was executed with the following command:merTrim -F reads.fastq -m 22 -mc meryl_db -mCillumina -t 16 -o out.fastq
In this command, –F specifies the reads, –m the k-mer size, –mc the database of trusted k-mers, and –mCillumina specifies that Illumina type adapters should be removed. The –t option defines the number of threads and thus only influences run time.
Following correction and trimming, the files in frg format were created with the following commands, as implemented in Celera Assembler: fastqToCA -technology illumina -insertsize 500 50 -libraryname lib_name -mates read1_clean.fastq,read2_clean.fastq>paired_reads.frgfastqToCA -technology illumina-long -insertsize 500 50 -libraryname lib_name -reads merged_reads.fastq>merged_reads.frg
The frg files contain information about the sequencing data, such as the expected insert size, location of the fastq files and the prefix for determining the species. Providing this information in the form of frg files is a prerequisite for Celera Assembler. Celera Assembler was then used to assemble the sequencing reads, with the following command specifying the prefix (–p) and the directory for the output (–d):runCA -p prefix -d CA -s spec_file
The ‘spec_file’ contains a list of settings and run-options for Celera Assembler. Some of the settings and options are specific to the computing system used for the assembly (such as the number of parallel overlap processes, ‘ovlConcurrency’), but as mentioned above, k-mer size as specified with the option –m (‘merSize’) can have effects on the contiguity of the assembly. The option ‘doFragmentCorrection’ was set to 0 because the reads were corrected with merTrim. The content of this file was:
ovlConcurrency=4 ovlThreads=8 cnsConcurrency=32 merSize=22 merylMemory=50000 merylThreads=32 merThreshold=5000 doOBT=0 overlapper=ovl ovlRefBlockSize=6000000 ovlHashBits=24 ovlHashBlockLength=800000000 doFragmentCorrection=0 unitigger=bogart batMemory=55 batThreads=32 doExtendClearRanges=0 doToggle=0 paired_reads.frg merged_reads.frg
The output of Celera Assembler consists of a set of three fasta files with increasing continuity that contain unitigs, contigs and scaffolds, respectively. Unitigs are either a unique DNA sequence found in a genome or a repeat, and unique unitigs are used as seeds to create contigs and scaffolds. In cases where Celera Assembler was not able to place a unitig confidently in the assembly, this unitig was not included in the contigs and scaffolds, but output separately. As a result of this, some additional sequence information is available in the assembled unitig fasta file compared to the assembled scaffolds. These additional sequences can include repeated sequences like transposable elements and tandem repeats, but also repeated gene fragments, conserved gene family domains, and other sequences that conflict with the biological assumptions of the assembler. As multiple copies of the mitochondrial genome are present in each cell, it is sequenced to a much higher coverage than the nuclear genome, and may therefore also be excluded from contigs due to false classification as a repetitive region. For these reasons, unitigs instead of contigs were used for both the identification of fragmented genes (see Malmstrøm et al.11) and for the mitochondrial phylogeny analysis described below. Assembly statistics for all draft genomes are provided in Table 3 (available online only).
Code availability
The most crucial commands are implemented in the Methods section, while additional scripts (used in phylogenetic analyses) are available on the code repository on GitHub (https://github.com/uio-cees/teleost_genomes_data_descriptor).
Data Records
All raw sequencing reads have been deposited in the European Nucleotide Archive (ENA) with study accession number PRJEB12469 (Data Citation 1). Table 4 (available online only) list the sample identifiers for each species. Each read file is available as a compressed file in fastq format (with extension fastq.gz). For some of the species, more than one read set is available as these were sequenced in two rounds, aiming to increase coverage. Two versions of all assembled genomes, unitigs (utg) and scaffolds (scf), are deposited in the Dryad repository under digital object identifier (DOI): doi:10.5061/dryad.326r8. (Data Citation 2). See Table 4 (available online only) for specific DOI for each species and assembly type.
Technical Validation
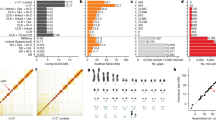
Both genome coverage and N50 lengths of contigs and scaffolds are considered important attributes for assessing a genome assembly. Assembly statistics for all species are reported in Table 2 (available online only). Another, and perhaps more crucial attribute, is the completeness of gene space, which is particularly important for the investigation of gene presence or absence. We used two different programs, CEGMA18 (Core Eukaryotic Genes Mapping Approach) v. 2.4.010312 and BUSCO19 (Benchmarking Universal Single-Copy Orthologs) v. 1.1b, to assess the gene-space completeness of our draft genome assemblies. CEGMA generates a list of ‘partial’ and ‘complete’ gene hits for the 248 most conserved genes, which were used as a validation of the assembly quality. BUSCO can be executed with several different reference data sets, optimized for different taxonomic groups. We used the ‘actinopterygii’ data set consisting of 3,698 highly conserved genes in acanthopterygian species (this specific data set is not publicly available yet—as of September 9th, 2016—but was provided by the developers of BUSCO upon request). BUSCO identifies and classifies these genes in the target genomes as either ‘Complete’, ‘Complete and duplicated’, ‘Fragmented’ or ‘Missing’. Table 5 (available online only) lists the CEGMA and BUSCO results for all assembled draft genomes, while Fig. 2a,b show the proportions of these conserved genes found (as partial hits) in relation to the read coverage and N50 scaffold length of all assemblies. In line with the results of our initial investigation of the budgerigar genome, we find no improvement in CEGMA or BUSCO gene set recovery when assembly coverage exceeds ~15× for the genomes included in this data set (linear regression of BUSCO versus coverage (>15×): R2=0.038, P=0.07; CEGMA versus coverage (>15×): R2=0.002, P=0.30) (Fig. 2a). When comparing the fractions of partial CEGMA and BUSCO genes recovered in each assembly with the N50 scaffold lengths of these assemblies, an initial steep increase is evident, clearly illustrating the sensitivity of these methods in relation to continuity (linear regression of BUSCO versus N50 scaffold length: R2=0.55, P<10–12; CEGMA versus N50 scaffold length: R2=0.30, P<10–5) (Fig. 2b). Finally, we find that the N50 scaffold length is largely uncorrelated with coverage (linear regression: R2=0.015, P=0.17), indicating that the specific sequencing strategy (insert size and read length) and the properties of the sequenced genomes (repeat content etc.) are more likely the limiting factors for N50 scaffold length (Fig. 2c). The observed lack of a correlation across all assemblies seems to be influenced by generally low N50 scaffold lengths for species of the order Gadiformes despite relatively high coverage for these genomes (mean coverage: 28×, mean N50 scaffold length: 6 kbp) compared to all other genomes (mean coverage: 20×, mean 50 scaffold length: 16 kbp). Thus, the species of the order Gadiformes appear more difficult to assemble which is likely explained by their high proportion of repetitive regions (see Tørresen et al.17). Collectively, these analyses illustrate that most of the variation in the recovery rate of the highly conserved genes is not due to low coverage, but rather reflects lineage-specific genomic features such as the amount and identity of repetitive elements that hamper the assembly of long continuous sequences.
(a) Scatterplot illustrating the correlation of gene space completeness (evaluated on the basis of BUSCO and CEGMA partially complete genes detected) and the read coverage (linear regression of BUSCO versus coverage (>15×): R2=0.038, P=0.07; CEGMA versus coverage (>15×): R2=0.002, P=0.30). (b) Scatterplot showing the correlation of BUSCO / CEGMA scores and N50 scaffold length (linear regression of BUSCO versus N50 scaffold length: R2=0.55, P<10–12 and CEGMA versus N50 scaffold length: R2=0.30, P<10–5) for all genome presented in the data set. (c) Scatterplot illustrating the correlation of coverage and N50 scaffold length (linear regression: R2=0.015, P=0.17). Species within the order Gadiformes are represented by triangles in all three plots. The lines shown are smooth LOESS curves, also referred to as local regressions, and the gray shaded areas represent 95% confidence interval in all three plots.
Phylogenetic analyses using mitochondrial genomes
To verify the correct identification of sampled species and the absence of contamination, we performed phylogenetic analyses of mitochondrial genomes extracted from all assemblies, in combination with previously available mitochondrial sequence data for sampled taxa and their close relatives. Mitochondrial genomes are particularly suitable for this comparison as the coverage of mitochondrial sequences is usually extremely high owing to the multiple copies of mitochondrial DNA (mtDNA) present in each mitochondrion and the large number of mitochondria per cell20. Furthermore, mitochondrial genomes are useful phylogenetic markers due to the very low frequency of recombination in animal mtDNA21 and the large number of mitochondrial genome sequences already available in GenBank22 (Data Citations 5 to 124).
We downloaded mitochondrial genome sequences for 120 species of which 14 species (Lampris guttatus, Polymixia japonica, Percopsis transmontana, Zeus faber, Stylephorus chordatus, Lota lota, Gadus morhua, Monocentris japonicus, Rondeletia loricata, Beryx splendens, Antennarius striatus, Anabas testudineus, Helostoma temminkii, and Perca fluviatilis) were also included in our set of 66 new teleost genome assemblies and an additional 8 species (Osmerus mordax, Polymixia lowei, Bregmaceros nectabanus, Beryx decadactylus, Myripristis berndti, Lamprogrammus niger, Carapus bermudensis, and Thunnus thynnus) were represented by a congener. GenBank accession numbers for the 120 downloaded genome sequences are given in Table 6 (available online only) (Data Citations 5 to 124). Protein-coding sequences for all mitochondrial genes except mt-ND6 (see Miya et al.23) were extracted from the 120 mitochondrial genomes, aligned with the software MAFFT24, v7.213 and translated to amino-acid sequences using AliView25 v.1.16.
To extract mitochondrial genomes from the 66 new unitig assemblies, we generated nucleotide BLAST databases for a subset of each assembly, consisting of all unitigs matched by at least 1,000 reads. This threshold was selected based on observed coverage distributions and the assumption that mitochondrial unitigs have particularly high coverage due to the relatively higher abundance of mitochondrial compared to nuclear DNA within each cell. The use of this threshold does not imply that all unitigs with higher coverage are mitochondrial, only that unitigs with lower coverage were ignored when mining for mitochondrial orthologs. For each mitochondrial gene, all 120 aligned amino-acid sequences were used as queries in searches with TBLASTN26 v.2.2.29 to identify unitigs with orthologous sequences in each of the 66 BLAST databases. For comparison, we also performed TBLASTN searches with the same queries against 10 additional BLAST databases generated for genome assemblies downloaded from ENSEMBL27 v.78 (Danio rerio, Astyanax mexicanus, Gadus morhua, Gasterosteus aculeatus, Oreochromis niloticus, Oryzias latipes, Takifugu rubripes, Tetraodon nigroviridis and Xiphophorus maculatus) and GenBank (Salmo salar; NCBI accession number AGKD00000000.3). For each of the 76 BLAST databases, the overall best TBLASTN hit for each mitochondrial gene was recorded and accepted as a homologous sequence if its e-value was below 1e–15. In cases where different unitigs matched different regions of the same gene (each with e-values below the threshold), these unitigs were jointly recorded as a single hit. Unitig identifiers for all hits are given in Table 7 (available online only). All hits were subsequently added to the untranslated mitochondrial gene alignments and realigned on the basis of amino-acid translations using TranslatorX28. Alignments were further analyzed with the software BMGE29 v.1.0 to determine unreliably aligned regions, and we excluded all codons that included sites with a gap rate above 0.2 or a smoothed entropy-like score (see Criscuolo & Gribaldo29) above 0.5. Finally, we concatenated the alignments of all mitochondrial genes, excluding two taxa (Parasudis fraserbrunneri and Acanthochaenus luetkenii) for which no homologs could be identified for eight or more genes. The final alignment used for phylogenetic inference included 9,303 bp.
Maximum-likelihood phylogenetic inference was performed with the software RAxML30 v.8.1.12, applying separate instances of the GTRCAT substitution model31 to three partitions corresponding to all first, second, and third codon positions. To assess the impact of potentially saturated third codon positions in the phylogenetic inference, we conducted two additional analyses in which these positions were either completely ignored or coded as ‘R’ and ‘Y’ so that only transversions would be counted as state changes. Phylogenetic node support was estimated through bootstrapping with an automatically determined number of bootstrap replicates (RAxML option ‘autoMRE’).
Topologies of the three resulting maximum-likelihood phylogenies based on different usage of third codon positions were highly congruent, however, basal branches appeared to be best resolved in the analysis based on the alignment with three equally coded partitions. This maximum-likelihood phylogeny (Fig. 3) also received the highest mean bootstrap support (81.6, compared to 76.7 and 80.1 for the analyses in which third codon positions were ignored or coded as ‘R’ and ‘Y’, respectively). All taxa sampled for new genome assemblies had phylogenetic positions according to the expectations; for the 14 species for which we included both a GenBank sequence and a mitochondrial genome extracted from new assembly data, the two sequences clustered monophyletically in each case and were connected by short branches (see e.g., Polymixia japonica; Fig. 3). In other cases, mitochondrial genomes extracted from new assemblies clustered monophyletically with their congeneric counterparts downloaded from GenBank (see e.g., the mitochondrial genomes of Osmerus eperlanus and Osmerus mordax; Fig. 3).
Sequences extracted from the new assemblies are marked in bold, all other mitochondrial genome sequences were previously available from the GenBank or ENSEMBL (where noted) databases. Black circles on nodes are sized proportional to bootstrap support, and the circle size corresponding to support values of 50, 75, and 100 are shown. Clade labels indicate taxonomic orders of all species as well as (with smaller font size) the (sub)family of gadiform species, following Betancur-R. et al.10 and Nelson48. Note that the orders Tetraodontiformes, Beloniformes, and Lophiiformes appear as non-monophyletic (marked with asterisks). For comparability, color code is identical to Fig. 1 in Malmstrøm et al.11. The tree file in Newick format has been deposited on Figshare under DOI: doi:10.6084/m9.figshare.4224234 (Data Citation 4).
It should be noted that basal phylogenetic nodes generally received relatively weak bootstrap support values, indicating that mitochondrial sequence data may not be sufficient to reliably resolve these ancient divergence events. Furthermore, three orders (Tetraodontiformes, Beloniformes, and Lophiiformes) appeared non-monophyletic, however, in all of these cases only weakly supported nodes separated two subgroups of the order. Thus, our results do not contradict the monophyly of these orders, which has been strongly supported in previous studies10,32,33. Most importantly, despite the not unexpected lower support values of basal nodes, our mitochondrial phylogeny corroborates the correct species identification and the absence of DNA contamination in the 66 new assemblies.
Phylogenetic analyses using nuclear markers
To reliably reconstruct the evolutionary history of the 66 sequenced teleost species, we further extracted a set of carefully selected phylogenetic markers from the nuclear genomes. Based on a strict filtering procedure (see Malmstrøm et al.11), we selected one-to-one orthologs for 567 exons of 111 genes from the 66 draft assemblies and from 10 genome assemblies available in the ENSEMBL database (Danio rerio, Astyanax mexicanus, Gasterosteus aculeatus, Oreochromis niloticus, Oryzias latipes, Takifugu rubripes, Tetraodon nigroviridis, Poecilia formosa and Xiphophorus maculatus) or GenBank (Salmo salar). The 111 selected genes were characterized by clock-like evolution, homogeneity in GC content among species, and no or only weak signals of selection and were therefore particularly well suited for the reconstruction of time-calibrated phylogenies. The 111 genes were distributed across all chromosomes of the zebrafish genome and included between 3 and 14 exons that were used in our analyses. Per gene, we concatenated sequences of these exons into a single alignment, which then included between 300 and 1,888 (mean: 643.4) bp, between 47 and 777 (mean: 240.5) variable sites and between 33 and 490 (mean: 157.5) parsimony-informative sites. As orthologous sequences for the 111 genes could be detected in almost all assemblies, the resulting 111 alignments contained only between 1.4 and 11.9% (mean: 7.3%) missing data (Table 8 (available online only)).
These alignments were used for an extensive set of phylogenetic analyses to reconstruct the species tree as well as individual gene trees, using both maximum-likelihood and Bayesian inference. Detailed descriptions of these analyses and a discussion of the resulting species tree can be found in Malmstrøm et al.11 In addition, we here present analyses of gene tree discordance in relation to the 66 new assemblies, as a heterogeneous phylogenetic signal among gene trees could, among other causes (e.g. Fontaine et al.34; Gante et al.35), result from assembly issues such as contamination. To quantify gene tree discordance, we compared each gene tree to the species tree based on their K-scores36 and using the Shimodaira-Hasegawa (SH) test37 implemented in PAUP* v.4.0a150 (http://paup.csit.fsu.edu). All gene trees used in this comparison were maximum-clade-credibility (MCC) trees inferred with the software BEAST38 v.2.2.0 for each of the 111 alignments. Similarly, we considered the MCC tree inferred with BEAST for a single concatenated alignment of all genes as the species tree (Fig. 1 in Malmstrøm et al.11) used in this comparisons. According to results of the SH test, all but four gene trees were significantly different (P<0.05) from the species tree (Table 8 (available online only)). K-scores were calculated for 91 of the 111 gene trees, but could not be calculated for the remaining 20 gene trees due to negative branch lengths. The resulting K-scores ranged from 217.5 to 478.6, indicating considerable gene tree discordance in agreement with the results of the SH test (even though individual K-scores and SH test P-values did not correlate; P=0.64). However, such tree discordance does not necessarily indicate assembly issues but can arise from multiple factors including incomplete lineage sorting39 or a lack of phylogenetic signal40. While high levels of incomplete lineage sorting have been shown to affect phylogenomic inference of rapidly radiating lineages like Neoavian birds41 or cichlid fishes42,43, its effect is expected to be limited in the analysis of ancient clades with long internode distances44 such as the teleost species tree inferred from our set of 111 nuclear markers11. We investigated the presence of incomplete lineage sorting in this species tree in Malmstrøm et al.11 by testing for a correlation of indel hemiplasy and branch length45. However, since no such correlation could be detected in our data set, we concluded that incomplete lineage sorting was weak or absent in the teleost species tree reported in Malmstrøm et al.11. In addition, we now tested whether instead of incomplete lineage sorting, a lack of phylogenetic signal in individual marker alignments could explain the observed gene tree discordance. To this end, we calculated the mean Bayesian posterior probability (BPP) of each gene tree as a measure of its phylogenetic signal and compared it to the K-score between this gene tree and the species tree. We find a highly significant negative correlation between the two measures (linear regression: R2=0.34, P<10–15), which is illustrated in Fig. 4. Furthermore, we also detected a highly significant correlation between the number of parsimony-informative sites per marker and the respective K-score (linear regression: R2=0.49, P<10–13) (Fig. 4). These tests show that low phylogenetic signal in individual marker alignments, rather than contamination in the assemblies, is responsible for the observed gene tree discordance. This lack of signal in individual alignments, however, is not exclusive to our phylogenomic data set, but is a feature that is commonly observed in nuclear markers40,44. As demonstrated by Malmstrøm et al.11 as well as other phylogenomic studies45–47 the combination of such stringently filtered exonic markers nevertheless allows an extremely reliable inference of ancient species trees that could not be achieved with faster-evolving sequence such as mitochondrial genomes, intronic regions, or genes under selection. We therefore recommend the reuse of the marker set presented here as a highly suitable resource for future analyses of the teleost species tree with extended taxon sets.
Topological distances are measured by the K-score between the gene trees and the species trees, and phylogenetic signal of the gene trees is measured as mean Bayesian posterior probability (BPP). Dots are colored according to the number of parsimonious-informative (PI) sites. The black line represents the linear regression (R2=0.34, P<10–15).
Usage Notes
Sequencing reads from all species can be downloaded from the European Nucleotide Archive (ENA), under the sample identifiers ERS1199874—ERS1199939. Unitig and scaffold level assemblies are available for download from the Dryad repository with individual assemblies found under DOI: doi:10.5061/dryad.326r8/1.—dryad.326r8/132. See Table 7 (available online only) for individual identifiers for both the raw sequencing read sets and the two assembly versions. The mitochondrial phylogeny (Fig. 3) can be downloaded as a tree file in Newick format from Figshare under DOI: doi:10.6084/m9.figshare.4224234 (Data Citation 4).
Additional information
How to cite: Malmstrøm, M. et al. Whole genome sequencing data and de novo draft assemblies for 66 teleost species. Sci. Data 4:160132 doi: doi:10.1038/sdata.2016.132 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Lieschke, G. J. & Currie, P. D. Animal models of human disease: Zebrafish swim into view. Nat. Rev. Genet. 8, 353–367 (2007).
Chang, N. et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res. 23, 465–472 (2013).
White, R., Rose, K. & Zon, L. Zebrafish cancer: The state of the art and the path forward. Nat. Rev. Cancer 13, 624–636 (2013).
Braasch, I. et al. A new model army: Emerging fish models to study the genomics of vertebrate Evo-Devo. J. Exp. Zool. 324, 316–341 (2014).
Ellegren, H. Genome sequencing and population genomics in non-model organisms. Trends Ecol. Evol. 29, 51–63 (2014).
MacRae, C. A. & Peterson, R. T. Zebrafish as tools for drug discovery. Nat. Rev. Drug. Discov. 14, 721–731 (2015).
Nakamura, T., Gehrke, A. R., Lemberg, J., Szymaszek, J. & Shubin, N. H. Digits and fin rays share common developmental histories. Nature 537, 225–228 (2016).
Star, B. et al. The genome sequence of Atlantic cod reveals a unique immune system. Nature 477, 207–210 (2011).
Malmstrøm, M., Jentoft, S., Gregers, T. F. & Jakobsen, K. S. Unraveling the evolution of the Atlantic cod’s (Gadus morhua L.) alternative immune strategy. PLoS ONE 8, e74004 (2013).
Betancur-R, R. et al. The tree of life and a new classification of bony fishes. PLoS Curr. 5, 1–45 (2013).
Malmstrøm, M. et al. Evolution of the immune system influences speciation rates in teleost fishes. Nat. Genet. 48, 1204–1210 (2016).
Bradnam, K. R. et al. Assemblathon 2: Evaluating de novo methods of genome assembly in three vertebrate species. GigaSci. 2, 545–531 (2013).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Miller, J. R. et al. Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 24, 2818–2824 (2008).
Simpson, J. T. Exploring genome characteristics and sequence quality without a reference. Bioinformatics 30, 1228–1235 (2014).
Magoc, T. & Salzberg, S. L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Tørresen, O. K. et al. An improved genome assembly uncovers prolific tandem repeats in Atlantic cod. bioRxiv 10.1101/060921, 1–33 (Cold Spring Harbor Labs Journals, 2016).
Parra, G., Bradnam, K., Ning, Z., Keane, T. & Korf, I. Assessing the gene space in draft genomes. Nucleic Acids Res. 37, 289–297 (2009).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Robin, E. D. & Wong, R. Mitochondrial DNA Molecules and Virtual Number of Mitochondria per Cell in Mammalian Cells. J. Cell. Physiol. 136, 507–513 (1968).
Rokas, A., Ladoukakis, E. & Zouros, E. Animal mitochondrial DNA recombination revisited. Trends Ecol. Evol. 18, 411–417 (2003).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 43, D30–D35 (2015).
Miya, M. et al. Mitochondrial genome and a nuclear gene indicate a novel phylogenetic position of deep-sea tube-eye fish (Stylephoridae). Ichthyol. Res. 54, 323–332 (2007).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30, 3276–3278 (2014).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinform. 10, 421 (2009).
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–D669 (2015).
Abascal, F., Zardoya, R. & Telford, M. J. TranslatorX: Multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res. 38, W7–W13 (2010).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): A new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Near, T. J. et al. Phylogeny and tempo of diversification in the superradiation of spiny-rayed fishes. Proc. Natl. Acad. Sci. 110, 12738–12743 (2013).
Matschiner, M. et al. Bayesian phylogenetic estimation of clade ages supports trans-Atlantic dispersal of cichlid fishes. Syst. Biol. 10.1093/sysbio/syw076, 1–59 (2016).
Fontaine, M. C. et al. Mosquito genomics. Extensive introgression in a malaria vector species complex revealed by phylogenomics. Science 347, 1258524–1258524 (2015).
Gante, H. F. et al. Genomics of speciation and introgression in Princess cichlid fishes from Lake Tanganyika. Mol. Ecol. 10.1111/mec.13767, 1–19 (2016).
Soria-Carrasco, V., Talavera, G., Igea, J. & Castresana, J. The K tree score: Quantification of differences in the relative branch length and topology of phylogenetic trees. Bioinformatics 23, 2954–2956 (2007).
Shimodaira, H. & Hasegawa, M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol. Biol. Evol. 16, 1114–1116 (1999).
Bouckaert, R. et al. BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Maddison, W. P. Gene trees in species trees. Syst. Biol. 46, 523–536 (1997).
Salichos, L. & Rokas, A. Inferring ancient divergences requires genes with strong phylogenetic signals. Nature 497, 327–331 (2013).
Suh, A., Smeds, L. & Ellegren, H. The dynamics of incomplete lineage sorting across the ancient adaptive radiation of Neoavian birds. PLoS Biol. 13, e1002224 (2015).
Brawand, D. et al. The genomic substrate for adaptive radiation in African cichlid fish. Nature 513, 375–381 (2014).
Meyer, B. S., Matschiner, M. & Salzburger, W. Disentangling incomplete lineage sorting and introgression to refine species-tree estimates for lake Tanganyika cichlid fishes. Syst. Biol. 10.1093/sysbio/syw069, 1–62 (2016).
Gatesy, J. & Springer, M. S. Phylogenetic analysis at deep timescales: Unreliable gene trees, bypassed hidden support, and the coalescence/concatalescence conundrum. Mol. Phylogenet. Evol. 80, 231–266 (2014).
Jarvis, E. D. et al. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014).
Misof, B. et al. Phylogenomics resolves the timing and pattern of insect evolution. Science 346, 763–767 (2014).
Prum, R. O. et al. A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Nature 526, 569–573 (2015).
Nelson, J. S. Fishes of the World. 4th edn (John Wiley & Sons, 2006).
Data Citations
ENA Sequence Read Archive PRJEB12469 (2016)
Malmstrøm, M. Dryad Data Repository https://doi.org/10.5061/dryad.326r8 (2016)
NCBI Sequence Read Archive ERP002324 (2013)
Malmstrøm, M. Figshare https://dx.doi.org/10.6084/m9.figshare.4224234 (2016)
GenBank NC_009064 (2007)
GenBank NC_004398 (2006)
GenBank NC_024752 (2014)
GenBank NC_008128 (2006)
GenBank NC_004391 (2010)
GenBank AB282828 (2013)
GenBank NC_004391 (2010)
GenBank NC_004372 (2008)
GenBank NC_008129 (2006)
GenBank NC_004413 (2016)
GenBank NC_008130 (2006)
GenBank NC_003178 (2006)
GenBank NC_002674 (2010)
GenBank NC_004374 (2006)
GenBank AP006738 (2016)
GenBank NC_004393 (2010)
GenBank NC_003188 (2006)
GenBank NC_008124 (2010)
GenBank NC_004405 (2006)
GenBank NC_004406 (2006)
GenBank NC_004373 (2007)
GenBank NC_004375 (2006)
GenBank NC_004383 (2016)
GenBank NC_004389 (2010)
GenBank NC_018340 (2012)
GenBank NC_003159 (2010)
GenBank NC_004381 (2006)
GenBank NC_004382 (2006)
GenBank NC_003160 (2006)
GenBank NC_003169 (2011)
GenBank NC_003183 (2006)
GenBank NC_002646 (2010)
GenBank NC_004404 (2006)
GenBank NC_001727 (2012)
GenBank NC_001606 (2010)
GenBank NC_003194 (2006)
GenBank NC_004402 (2006)
GenBank NC_003185 (2006)
GenBank NC_003164 (2010)
GenBank NC_004376 (2006)
GenBank AB034825 (2012)
GenBank NC_008126 (2006)
GenBank NC_008127 (2006)
GenBank NC_004415 (2006)
GenBank NC_004407 (2009)
GenBank NC_005254 (2010)
GenBank NC_003184 (2006)
GenBank NC_002081 (2010)
GenBank NC_004388 (2010)
GenBank AP002944 (2016)
GenBank AP005977 (2013)
GenBank NC_003161 (2006)
GenBank NC_003195 (2006)
GenBank NC_022728 (2013)
GenBank AB282829 (2013)
GenBank NC_003187 (2006)
GenBank NC_004386 (2006)
GenBank NC_004400 (2006)
GenBank NC_003179 (2006)
GenBank NC_004401 (2006)
GenBank NC_003176 (2011)
GenBank NC_009054 (2012)
GenBank NC_003165 (2006)
GenBank NC_004378 (2006)
GenBank NC_008125 (2010)
GenBank NC_004380 (2010)
GenBank NC_004379 (2006)
GenBank NC_004409 (2006)
GenBank NC_003193 (2006)
GenBank NC_004384 (2006)
GenBank NC_004385 (2006)
GenBank NC_004392 (2010)
GenBank NC_003192 (2006)
GenBank NC_003182 (2006)
GenBank NC_003163 (2010)
GenBank NC_003189 (2006)
GenBank NC_004399 (2006)
GenBank NC_009062 (2007)
GenBank NC_003180 (2006)
GenBank NC_009061 (2007)
GenBank NC_001717 (2010)
GenBank NC_004387 (2008)
GenBank NC_015246 (2011)
GenBank NC_004394 (2010)
GenBank NC_002386 (2010)
GenBank NC_004396 (2010)
GenBank NC_026313 (2015)
GenBank NC_003168 (2006)
GenBank NC_004411 (2006)
GenBank NC_004410 (2006)
GenBank NC_004377 (2006)
GenBank NC_002648 (2010)
GenBank NC_003181 (2006)
GenBank NC_006920 (2010)
GenBank NC_003172 (2006)
GenBank NC_004408 (2006)
GenBank NC_004414 (2006)
GenBank NC_003186 (2006)
GenBank AP004451 (2016)
GenBank NC_002616 (2010)
GenBank NC_004395 (2010)
GenBank NC_004403 (2012)
GenBank NC_003162 (2010)
GenBank NC_011936 (2009)
GenBank NC_013723 (2010)
GenBank NC_003171 (2010)
GenBank NC_002574 (2011)
GenBank NC_008123 (2010)
GenBank NC_024236 (2016)
GenBank NC_003177 (2006)
GenBank NC_009948 (2010)
GenBank NC_004416 (2011)
GenBank AP004439 (2016)
GenBank NC_014052 (2016)
GenBank NC_003166 (2010)
GenBank AB282835 (2013)
GenBank NC_004397 (2006)
GenBank NC_003173 (2010)
GenBank NC_003190 (2010)
GenBank NC_003167 (2006)
Acknowledgements
All computational work was performed on the Abel Supercomputing Cluster (Norwegian metacenter for High Performance Computing (NOTUR) and the University of Oslo) operated by the Research Computing Services group at USIT, the University of Oslo IT-department (http://www.hpc.uio.no/). Sequencing library creation and high-throughput sequencing were carried out at the Norwegian Sequencing Centre (NSC), University of Oslo, Norway, and McGill University and Genome Quebec Innovation Centre, Canada. We are grateful for the kindness of the following researchers for providing tissue samples for this study: K. Østbye (University of Oslo), F. Cortesi (University of Queensland), J. Y. Poulsen (Greenland Institute of Natural Resources), Reinhold Hanel (Thünen-Institute of Fisheries Ecology) M. Miya (Natural History Museum & Institute, Chiba University), A. Bentley (University of Kansas Biodiversity Institute), I. Kilde (Norwegian University of Science and Technology in Trondheim), I. R. Bradbury (Memorial University), C. Pampoulie (Marine Research Institute of Iceland), and L. Rüber (Natural History Museum in Bern). This work was funded by grants from the Research Council of Norway (RCN grants 199806 and 222378) to K.S.J. Mi.M. acknowledges funding from the Swiss National Science Foundation (SNF grant PBBSP3-138680. We thank Brian Walenz and Jason R. Miller for assistance with Celera Assembler.
Author information
Authors and Affiliations
Contributions
M. Malmstrøm, K.S.J. and S.J. conceived the project. M. Malmstrøm, M. Matschiner, K.S.J. and S.J. planned and oversaw the project. M. Malmstrøm and S.J. provided samples for genome sequencing. M. Malmstrøm prepared DNA and libraries for genome sequencing. M. Malmstrøm and S.J. were in charge of genome sequencing management. M. Malmstrøm, and O.K.T. performed quality control and assembled the genomes. O.K.T. performed gene-space completeness assessments. M. Matschiner performed mitochondrial and nuclear marker based phylogenetic analyses. M. Malmstrøm, M. Matschiner and O.K.T. prepared the figures. M. Malmstrøm and M. Matschiner wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Malmstrøm, M., Matschiner, M., Tørresen, O. et al. Whole genome sequencing data and de novo draft assemblies for 66 teleost species. Sci Data 4, 160132 (2017). https://doi.org/10.1038/sdata.2016.132
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.132
This article is cited by
-
Mitochondrial genome structure and composition in 70 fishes: a key resource for fisheries management in the South Atlantic
BMC Genomics (2024)
-
A chromosome-level genome of the striated frogfish (Antennarius striatus)
Scientific Data (2024)
-
Multi-genome comparisons reveal gain-and-loss evolution of anti-Mullerian hormone receptor type 2 as a candidate master sex-determining gene in Percidae
BMC Biology (2024)
-
Teleost genomic repeat landscapes in light of diversification rates and ecology
Mobile DNA (2023)
-
First paleoproteome study of fossil fish otoliths and the pristine preservation of the biomineral crystal host
Scientific Reports (2023)