Abstract
Teleost fishes constitute the most species-rich vertebrate clade and exhibit extensive genetic and phenotypic variation, including diverse immune defense strategies. The genomic basis of a particularly aberrant strategy is exemplified by Atlantic cod, in which a loss of major histocompatibility complex (MHC) II functionality coincides with a marked expansion of MHC I genes. Through low-coverage genome sequencing (9–39×), assembly and comparative analyses for 66 teleost species, we show here that MHC II is missing in the entire Gadiformes lineage and thus was lost once in their common ancestor. In contrast, we find that MHC I gene expansions have occurred multiple times, both inside and outside this clade. Moreover, we identify an association between high MHC I copy number and elevated speciation rates using trait-dependent diversification models. Our results extend current understanding of the plasticity of the adaptive immune system and suggest an important role for immune-related genes in animal diversification.
Similar content being viewed by others
Main
With over 32,000 extant species1, teleost fishes comprise the majority of vertebrate species. Their taxonomic diversity is matched by extensive genetic and phenotypic variation, including novel immunological strategies. Although the functionality of the adaptive immune system has been considered to be conserved since its emergence in the ancestor of all jawed vertebrates2,3, fundamental modifications of the immune gene repertoire have recently been reported in teleosts4,5,6,7. One of the most dramatic changes has occurred in Atlantic cod (Gadus morhua), involving complete loss of the MHC II pathway that is otherwise responsible for the detection of bacterial pathogens in vertebrates4. Moreover, this loss is accompanied by a substantially enlarged repertoire of MHC I genes, which normally encode molecules for protection against viral pathogens. It has thus been hypothesized that the expanded MHC I repertoire of cod evolved as a compensatory mechanism, whereby broader MHC I functionality makes up for the initial loss of MHC II (refs. 4,6). However, the questions of how and when MHC II was lost relative to the MHC I expansion, and whether these genomic modifications are causally related, have so far remained unresolved.
As key components of the vertebrate adaptive immune system, the complex MHC pathways and their functionality are now well characterized8,9,10, but less is known about the causes of MHC copy number variation, which poses an immunological tradeoff11,12. Although an increase in the number of MHC genes facilitates pathogen detection, it will also decrease the number of circulating T cells13,14,15,16, resulting in an immune system that can detect a large number of pathogens at the expense of being less efficient in removing them. The evolution of MHC copy numbers is therefore likely driven toward intermediate optima determined by a tradeoff between detection and elimination of pathogens—as suggested by selection for 5–10 copies inferred in case studies of fish17,18 and birds19. Because pathogen load and the associated selective pressures vary between habitats, the optimal number of MHC copies depends on the environment20,21,22. As a result, interbreeding between different locally adapted populations is expected to produce hybrids with excess (above optimal) MHC diversity that are characterized by T cell deprivation and low fitness. This process would introduce postzygotic reproductive isolation and promote reinforcement of premating isolation between the populations. Consequently, MHC genes have been suggested to have an important role in speciation22,23, but, to our knowledge, this role has never been tested comparatively in a macroevolutionary context.
Here we report comparative analyses of 76 teleost species, of which 66 were sequenced to produce partial draft genome assemblies, including 27 representatives of cod-like fishes within the order Gadiformes. First, we use phylogenomic analysis to resolve standing controversy regarding early-teleost divergences and to firmly establish the relationships of gadiform fishes. Second, by analyzing the presence and absence of key genes in the vertebrate adaptive immune system, we place the loss of MHC II functionality in the common ancestor of all Gadiformes; by time calibrating the teleost phylogeny on the basis of fossil sampling rate estimation, we further show that this loss occurred 105–85 million years ago. Third, we demonstrate that expansions of the MHC I gene repertoire are not limited to Gadiformes but are also found in three other clades, including the exceptionally species-rich group of percomorph fishes (Percomorphaceae). By applying phylogenetic comparative methods and ancestral state reconstruction, we trace the evolutionary history of MHC I gene expansions and infer distinct copy number optima in each of them. Finally, using trait-dependent diversification models, we identify a positive association between MHC I gene copy number and speciation rate. Our results highlight the plasticity of the vertebrate adaptive immune system and support the role of MHC genes as 'speciation genes', promoting rapid diversification in teleost fishes.
Results
Sequencing and draft assembly of 66 teleost genomes
To investigate the evolution of the immune gene repertoire of teleost fishes, we selected representatives of all major lineages in the Neoteleostei24 for low-coverage genome sequencing. As both loss and expansion of MHC genes have been reported in the Gadiformes4, we sampled this order more densely with 27 species representing most gadiform families and subfamilies (Supplementary Table 1)25. The sequencing strategy for de novo assembly of the 66 genomes was designed on the basis of gene space completeness and assembly statistics of simulated genomes (Supplementary Note). For each species, a single paired-end library was sequenced to 9–39× coverage on the Illumina HiSeq 2000 platform (Supplementary Table 2 and Supplementary Note). The genome sequences were assembled with the Celera Assembler26, resulting in N50 scaffold sizes between 3.2 and 71 kb (Supplementary Table 3 and Supplementary Note). For most species, we recovered more than 75% of the conserved eukaryotic genes included in the CEGMA27 analysis, and on average we recovered 74% of the conserved genes of the Actinopterygii data set included in the BUSCO28 analysis. Collectively, this is indicative of a sufficiently high degree of gene space completeness for gene detection in our partial draft genome assemblies (Supplementary Table 3 and Supplementary Note).
A genome-scale phylogeny of teleost fishes
To firmly establish teleost and gadiform relationships, we performed phylogenomic analyses using a stringently filtered data set of 567 exon orthologs from 111 genes, identified in the 66 new draft genome assemblies as well as in 10 previously published genomes. After filtering (Supplementary Note), the phylogenomic data set included 71,418 bp, which was used for time calibration and coalescent-based species tree analyses. To establish a timeline of teleost diversification and to determine the timing of immune gene repertoire alterations, we performed Bayesian phylogenetic inference with the concatenated phylogenomic data set, using a relaxed-clock model and 17 fossil constraints (Supplementary Figs. 1 and 2, Supplementary Tables 4 and 5, and Supplementary Note).
The time-calibrated species tree was used to test for incomplete lineage sorting among the species sampled for our phylogeny. Following Jarvis et al.29, we mapped genomic insertions and deletions on the rooted topology and found no correlation between branch lengths and uniquely mapped indels. This lack of correlation indicates that incomplete lineage sorting did not substantially affect divergences between the teleost lineages included in our phylogeny29 and supports concatenation as the most appropriate strategy for species phylogeny estimation with our data (Supplementary Note).
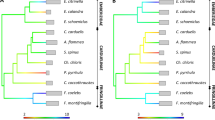
The resulting time tree (Fig. 1) supports a monophyletic clade including the orders Gadiformes, Stylephoriformes, Zeiformes, Polymixiiformes and Percopsiformes, thus corroborating the Paracanthopterygii sensu Miya et al.30. We further confirm the placement of Stylephorus chordatus as the closest extant relative of the Gadiformes30,31 and estimate the crown age of this order at approximately 85 million years ago. Within Gadiformes, we find support for a sister group relationship between Bregmacerotidae and all other gadiform families, which began to radiate around 70 million years ago.
Dashed branches correspond to species for which genome data were already available in Ensembl. All nodes are supported by Bayesian posterior probability of 1.0 except where noted (Supplementary Table 5). Clade names to the right and on branches are given according to Betancur-R et al.24. Branch colors represent taxonomy at the level of orders. Teleost illustrations are by G. Holm.
Loss of MHC II pathway genes
To assess the origin and extent of the MHC II pathway gene loss observed in Atlantic cod, we investigated the immune gene repertoires of the teleost genomes through a comparative gene mining pipeline comprising BLAST searches, prediction of ORFs and annotation (Supplementary Note). We explicitly investigated the presence of 27 genes chosen for their central role in MHC class I, MHC class II and cross-presentation pathways8,10. In addition, three highly conserved genes were included as a control (Fig. 2). As query sequences, we used orthologs from zebrafish (Danio rerio), medaka (Oryzias latipes), spotted green pufferfish (Tetraodon nigroviridis), fugu (Takifugu rubripes), three-spined stickleback (Gasterosteus aculeatus), Nile tilapia (Oreochromis niloticus), Amazon molly (Poecilia formosa), platyfish (Xiphophorus maculatus), cavefish (Astyanax mexicanus) and Atlantic cod (G. morhua) (Supplementary Table 6).
(a) Presence of key genes in MHC I, MHC II and cross-presentation pathways for recognition of pathogen-derived antigens. Genes not detected are indicated by white squares. Colors reflect order-level taxonomy, and the cladistic representation follows that in Figure 1. (b) Copy number estimates for non-classical MHC I Z-lineage (light gray) and U-lineage (dark gray) genes. Error bars, 95% confidence intervals based on 1,000 bootstrap replicates (Supplementary Note).
The genome-based phylogeny allowed us to place the immune gene characterization into an evolutionary perspective. Whereas the three control genes were identified in all species, we found that genes associated with the MHC II pathway were consistently missing in all inspected Gadiformes draft genome assemblies, as no orthologs could be identified for invariant chain (also known as cd74), cd4 and the MHC II α and β chains (Fig. 2a and Supplementary Table 7). This finding reiterates the previous observation in the Atlantic cod genome, where lack of these genes was confirmed through qPCR and synteny analyses4. Although individual genes could not be detected in a subset of species outside the Gadiformes, these occurrences comprise a minority (∼3%) of the total number of comparisons. Notably, these losses did not show any phylogenetic pattern and hence are more likely to have resulted from incomplete genome assembly (Supplementary Note). As MHC II pathway genes are otherwise ubiquitous among vertebrates, the observed pattern shows that these genes were lost collectively in the common ancestor of all Gadiformes, following its divergence from Stylephoriformes at approximately 105 million years ago. This result implies that MHC II gene loss is shared by all 616 extant species of gadiform fishes. These taxa inhabit a wide variety of habitats, showing that their alternative immune system is highly versatile and not restricted to specific ecological niches.
MHC I copy number variation
MHC I molecules exist as five distinct lineages—L, S, P, U and Z—with the latter two comprising predominantly 'classical' (peptide-binding) MHC I molecules involved in antigen presentation32,33,34,35. The Atlantic cod genome has been shown to harbor 80–100 copies of MHC I (ref. 4), which is 15 times higher than the copy number determined to be optimal in the three-spined stickleback14,17. To investigate the evolutionary origin of the gadiform MHC I expansion and potentially detect other expansions, we estimated the MHC I copy numbers of the two peptide-binding lineages (U and Z) in the 66 sequenced species. MHC I copy number estimates were calculated on the basis of Illumina raw sequencing reads relative to a set of 19 putatively single-copy reference regions. For both U- and Z-lineage genes, we determined the number of matching raw reads by aligning them to conserved regions of the MHC I genes (α3 domain) and the set of reference genes. The uncertainties of all copy number estimates were assessed with a double-bootstrapping procedure (Supplementary Tables 3 and 8, 9, 10, 11, and Supplementary Note). Our copy number estimates were cross-validated with previous results for Atlantic cod based on qPCR4 and transcribed MHC I U-lineage genes6. The copy number estimation procedure was further validated by estimating the relatively conserved number of Hox gene copies for all species. Estimated Hox gene copy numbers (mean = 50.3, s.d. = 11.4) were in agreement with expectations for teleost fishes36 and were uncorrelated with MHC I gene copy numbers (R2 = 0.002) (Supplementary Fig. 3, Supplementary Table 3 and Supplementary Note).
Mean copy number estimates for all species are shown in Figure 2b. Extremely high copy numbers of MHC I U-lineage genes were detected in Gadiformes, where several species had around 100 copies, followed by species within Percomorphaceae with up to 80 copies. Within Gadiformes, high copy numbers above 40 were observed in as many as 12 species. Although we also observed low copy numbers in a limited number of gadiform species (for example, five copies in Trisopterus minutus), nearly all representatives of this order appear to share an expanded MHC I gene repertoire with 15–30 copies of the U-lineage genes. Such gene family expansions may promote biological diversification by introducing new raw genetic material, potentially resulting in sub- or neofunctionalization and thus novel immunological pathways. In this regard, we identified the same cytoplasmic sorting motifs presumed to be part of a novel pathway enhancing cross-presentation functionality in Atlantic cod6,37,38 also in five additional species of the Gadinae. Because the origin of these signaling motifs only dates back to about 15 million years ago, it is likely that the enhanced cross-presentation functionality was not part of a preexisting machinery that could have favored the loss of MHC class II (Supplementary Fig. 4, Supplementary Table 12 and Supplementary Note).
Two hypotheses have been put forward as possible explanations for the loss of MHC II: a metabolic gain by not maintaining a costly system or a functional shift in the immune gene repertoire, rendering it obsolete39. Whether the loss occurred before or following the expansion of MHC I genes is key in discriminating between these hypotheses. In contrast to all other Gadiformes, the most basal gadiform lineage, represented by Bregmaceros cantori, is characterized by a complete absence of MHC I U-lineage genes, in addition to MHC II loss. The only antigen-presenting molecules detected in Bregmacerotidae are encoded by the MHC I Z-linage genes, of which 2–3 copies are still present. With minor exceptions, the Z-lineage genes were found in low copy numbers in all species, and, contrary to the U-lineage genes, they did not show a pattern of clade-specific expansions. Interestingly, the myctophiform Benthosema glaciale showed convergent loss of the MHC I U-lineage genes, and both Bregmaceros and Benthosema have experienced an additional loss of two genes involved in MHC I interactions with T cells (cd8a and cd8b), rendering the traditional pathway for endogenously derived pathogens non-functional as well (Fig. 2). The lack of MHC I U-lineage genes in B. cantori indicates that MHC I expansions in Gadiformes occurred after the divergence of Bregmacerotidae and thus subsequent to the loss of MHC II.
Rate shifts in MHC I copy number evolution
It has previously been shown that MHC copy number and molecular diversity are linked to the habitat that a species or population occupies21,22. Our data show that the species-rich clades Gadiformes and Percomorphaceae, which are both distributed across a wide variety of habitats, contain the highest numbers of MHC I gene copies among teleosts. We therefore used phylogenetic comparative methods to test for putative adaptive evolution of MHC I copy numbers in all groups included in our phylogeny. We used a Markov chain Monte Carlo (MCMC)-based reversible-jump Bayesian approach to fit multiple-regime Ornstein–Uhlenbeck models in which trait evolution is directed toward different optimal values in different parts of the tree40. This approach identified the probability and direction of rate shifts in MHC I copy number optima on branches in the phylogeny (Supplementary Fig. 5 and Supplementary Note). On the basis of the inferred branch-specific rate shift probabilities and the presence of copy number outliers, we tested explicit hypotheses for rate shift combinations in a likelihood framework41,42. We found that the overall best fitting model included five shifts of the MHC copy number optimum (Fig. 3a, Supplementary Table 13 and Supplementary Note). This model is characterized by a phylogenetic half-life of 23 million years, a stationary variance of 0.38 squared log copy numbers and an optimum of 6.8 MHC I copies for basal branches of the phylogeny, which is in concordance with the hypothesized MHC I repertoire of early gnathostomes43. In contrast, substantially higher optimal MHC I copy numbers of up to 58 copies were inferred within gadiform and percomorph fishes. The phylogenetic half-life of 23 million years shows that the MHC optima are approached slowly and that there can be considerable lag in adaptation. The multiple-optima model vastly outperformed alternatives such as Brownian motion, white noise, single-peaked Ornstein–Uhlenbeck and early-burst models. This finding corroborates the hypothesis that MHC I copy number evolution is characterized by selection toward intermediate optima, resulting from a tradeoff between detection and elimination of pathogens.
(a) 3D representation of teleost phylogeny (Fig. 1) and MHC I copy number evolution, based on ancestral state reconstruction with the best fitting Ornstein–Uhlenbeck model. This model supports five shifts toward elevated copy number optima, marked by colored dots: 25.2 copies in Percomorphaceae excluding Ophidiiformes, 41.5 in Berycoidei, 22.1 in Polymixiiformes and Percopsiformes, 19.8 in Gadiformes excluding Bregmacerotidae, and 57.5 in Gadiformes excluding Bregmacerotidae and Gadidae. Loss of MHC II in the common ancestor of Gadiformes is represented by a black dot. The ordering of species is identical to that in Figure 2. Gray shading visualizes reconstructed MHC I copy numbers along branches. Steps of 20 copies are indicated by light gray lines on the 3D reconstruction. (b) Per-species MHC I copy numbers and mean net diversification rate estimates, connected according to phylogenetic relationships and colored as in a. (c) Diversification rate estimates for lineages with high and low MHC I copy numbers, based on BiSSE analyses with incrementing thresholds between high and low copy numbers. Black lines represent mean estimates and gray shading represents standard deviation from 25 replicate analyses. (d) Improvement in log-transformed likelihood when speciation rates are unlinked for lineages with high and low copy numbers, in the analyses performed in c. The black line corresponds to the median and gray shading represents 0.05 and 0.95 quantiles from replicate analyses.
Diversification rate analyses
The Gadiformes comprise 616 extant species1 and are thus the most species-rich order within Paracanthopterygii, where Stylephoriformes are represented by a single species and Zeiformes, Percopsiformes and Polymixiformes collectively include 56 species. To investigate whether this pattern of diversity is the result of an elevated speciation rate in Gadiformes, we tested for differences in diversification among all clades included in our phylogenetic data set using a Bayesian framework (Supplementary Figs. 6, 7, 8 and Supplementary Note). For this analysis, we identified a set of 37 mutually exclusive clades that represent almost the entire extant diversity of the teleost supercohort Clupeocephala24. Assuming constant speciation and extinction rates within specific shift regimes, we identified two major shifts in diversification rates, one at the base of Gadiformes and a second within the taxonomically diverse Percomorphaceae24, that is, in the two clades featuring particularly high MHC I copy numbers (Fig. 3a). A comparison of per-species MHC I copy numbers and mean net diversification rates (Fig. 3b) indicates a relationship between the two measures. To quantify the association between MHC I copy numbers and rates of diversification, we carried out binary state speciation and extinction (BiSSE) analysis to estimate differences in diversification rate between lineages with high and low MHC I copy numbers (Supplementary Note). We found that diversification rates differed most when the threshold was placed between 20 and 25 copies (Fig. 3c). With a threshold in this range, the model with two separate speciation rates for lineages with high and low copy numbers was better supported than a model with a single speciation rate parameter (ΔAIC > 37.1; Fig. 3d and Supplementary Note). These results suggest that the influence of MHC I genes on speciation rates is stronger in species that have already evolved at least 20 copies.
Discussion
Teleost fishes are characterized by striking differences in species richness across different lineages, with some groups, such as cyprinids and cichlids, containing thousands of species, while others include only a single or very few taxa. Although these differences can in part be explained by ecological opportunity44 or key innovations45, a large proportion of lineage-specific variation in taxonomic diversity remains unexplained. It has previously been suggested that MHC genes can influence speciation rates through selection against hybrids with higher than optimal MHC copy numbers and consequent reinforcement22,23. For species with more MHC copies, the extra number of alleles in hybrids will be comparatively higher, and these species should therefore experience greater fitness reduction and stronger assortative mating to maintain co-adapted genes. The proposed role of MHC genes as speciation genes23, promoting diversification and the maintenance of recently diverged species, is therefore expected to be more pronounced in species exhibiting high copy numbers. The identification of an MHC I copy number threshold above which speciation rates are accelerated expands on this conceptual framework. Our analyses identify this threshold at 20–25 MHC I copies, suggesting that the effect of T cell depletion on hybrid fitness becomes more pronounced in this range and that this might affect mate choice in species with copy numbers above this threshold, promoting inbreeding and reinforcement.
By pinpointing the loss of MHC II pathway genes to the common ancestor of Gadiformes and by reconstructing MHC I copy number evolution, we show that loss of MHC II predated the MHC I expansions that occurred within this group. This implies that MHC II functionality was not outcompeted by a preexisting alternative immunological strategy such as an expanded MHC I repertoire but may instead have been lost as a result of the metabolic costs associated with its maintenance39. The temporal relationship between the loss and expansions in Gadiformes and the identification of putative cross-presentation signaling motifs in Atlantic cod6, as well as in several other gadiform species, indicate that a number of genes in the expanded MHC I repertoire evolved to compensate for loss of MHC II. Nevertheless, marked expansions of MHC I genes were also observed outside of Gadiformes in groups with intact MHC II pathways. These independent expansions clearly show that loss of MHC II is not the only potential trigger for MHC I expansion but that MHC copy number evolution has also been driven by other factors, highlighting the extensive plasticity of the teleost adaptive immune system.
The 66 new teleost draft genome assemblies provide unprecedented opportunities for comparative analyses of the largest clade of vertebrates. We have used this genomic information to unravel the evolutionary history of key immune genes and to show how MHC gene composition in teleost fishes has influenced diversification rates in this diverse vertebrate lineage.
Methods
Tissues, sequencing and assembly.
Genomic DNA was obtained from various tissues of the different species in this study. Most tissue samples were provided by museums and other collections, while some come from commercially caught fish in collaboration with local fishermen (see Supplementary Table 1 for a full list of tissues and contributors). A single paired-end library, with an insert size of ∼400 bp, was created for each species, using the Illumina TruSeq Sample Prep v2 Low-Throughput protocol. All species were sequenced (2 × 150 bp) to >9× coverage on the Illumina HiSeq 2000 platform, and sequences were assembled using the Celera Assembler26 (Supplementary Note). Draft genome assembly quality, in terms of gene space completeness, was assessed using CEGMA27 and BUSCO28 (Supplementary Table 3 and Supplementary Note).
Phylogenetic inference.
Strict filtering criteria were applied for the identification of suitable orthologous phylogenetic markers. For the 33,737 annotated zebrafish genes in release 78 of the Ensembl database46, we selected the longest transcript if it had at least five stop-codon-free exons of 150 bp or greater in length. We removed genes that could not be assigned to an Ensembl gene tree and genes for which teleost fishes did not form a monophyletic group in the gene tree. We further excluded genes for which the Ensembl gene tree indicated gene duplications among teleosts or did not include all ten teleost species of Ensembl v.78 (Supplementary Note). The ten teleost reference genomes of Ensembl were used to calculate TBLASTN bitscores for each of the zebrafish exons, using the BLAST+ v.2.2.29 suite of tools47. Exon-specific bitscore thresholds for putative orthologs were defined on the basis of this bitscore information, and exons for which two or more of the known orthologs had bitscores lower than this threshold were excluded. Genes with fewer than five remaining exons were discarded, which resulted in a set of 2,251 exon sequences of 302 zebrafish genes that were then used as queries in TBLASTN searches against each of the 66 new teleost draft genome assemblies, the 10 Ensembl teleost genomes and the genome sequence of salmon48. For each species, the best hits were accepted as putative orthologs if their TBLASTN bitscores were above the exon-specific bitscore threshold (Supplementary Note). Alignments of TBLASTN hits for the 2,251 exons were further filtered on the basis of ratios of nonsynonymous to synonymous substitutions (dN/dS) determined with the software codeml of the PAML 4 package49 to exclude exons putatively under positive selection. Unreliably aligned sites were identified with the software BMGE v.1.0 (ref. 50) and excluded from the alignments. We further removed all third codon positions from the alignments and excluded exons with high variation in GC content (Supplementary Note). For each gene, the concordance of individual exon trees was assessed with the software Concaterpillar v.1.7.2 (ref. 51), and we removed either individual exon alignments or all exon alignments for a given gene from the data set, depending on the number of congruent exon trees (Supplementary Note). To filter for genes with clock-like evolution, we estimated the coefficient of variation of rates of each gene with the software BEAST v.2.2.0 (ref. 52) and removed the genes with the highest estimated rate variation coefficients (Supplementary Note). After this step, our strictly filtered data set used for phylogenetic inference contained 567 exons from 111 genes, with a total alignment length of 71,418 bp and 7.3% missing data. To assess the consequences of strict filtering on phylogenetic inference, we compared maximum-likelihood phylogenies based on the strictly filtered data set (111 genes, 71,418 bp, 7.3% missing data) with phylogenies based on a data set that was substantially larger but less strictly filtered (302 genes, 252,442 bp, 18.2% missing data). Maximum-likelihood phylogenies for both data sets were inferred with the software RAxML v. 8.1.17 (ref. 53) (Supplementary Fig. 1 and Supplementary Note). The strictly filtered data set was further used to estimate the timeline of teleost diversification with the software BEAST v.2.2 (ref. 52). Calibration densities for time calibration were calculated with the BEAST add-on CladeAge54, on the basis of estimates of diversification rates and the fossil sampling rate. The earliest fossil occurrences of 17 clades in our phylogeny were identified and used to constrain the ages of these clades with CladeAge calibration densities, taking into account the uncertainties in the ages of these fossils (Supplementary Note). We further used coalescent-based species tree estimation to test for potentially misleading phylogenetic signal due to incomplete lineage sorting. This analysis was conducted both with individual gene trees and with trees based on alignments binned according to the binning approach of Mirarab et al.55. Maximum-likelihood trees produced with RAxML and maximum-clade-credibility trees resulting from BEAST analyses of binned and unbinned genes were used for species tree inference with the software ASTRAL v.4.7.8 (ref. 56) (Supplementary Fig. 2, Supplementary Table 5 and Supplementary Note). To test for incomplete lineage sorting among the taxa included in our phylogeny, we compared branch lengths and the proportion of synapomorphic indels supporting each branch, following Jarvis et al.29 (Supplementary Note).
Gene mining of draft genome assemblies.
All draft genome assemblies were mined for genetic content on the unitig (UTG) assembly level, as assembly parameters are stricter for UTGs than for contigs or scaffolds. The presence or absence of each gene was determined through an automated pipeline, using full-length amino acid sequences for 27 immune-related genes and 3 control genes, from ten teleost reference genomes (Ensembl gene identifiers are listed in Supplementary Table 6). Potential orthologs were detected using TBLASTN with an acceptance level of e value = 1 × 10−10 and evaluated through identification of ORFs predicted by the software Genescan57. All ORFs were then aligned to the UniProt database (Supplementary Note), and reciprocal TBLASTN hits were recorded as potentially correct if their e value was below 1 × 10−10. All recorded annotations for each gene were then manually inspected, and the best hit is reported (see the Supplementary Note for details and Supplementary Table 7 for the location of each identified ortholog).
Copy number estimation of MHC I genes.
High sequence similarity and conserved regions make the different MHC I genes difficult to assemble correctly. To estimate the number of copies of these genes in each of the sequenced genomes, we applied a new method for copy number estimation, based on a comparison of raw read counts for target and reference sequences. For MHC I U- and Z-lineage genes, we used 270 bp of the conserved α3 domain as the target and equivalently sized fragments from 14 single- or low-copy genes as references (see Supplementary Table 9 for a full overview of all reference regions). MHC I target sequences were prepared through consensus by majority for all hits detected in the individual draft genome assemblies with TBLASTN (e-value cutoff set to 1 × 10−5) using U- and Z-lineage MHC I α3-domain sequences from ten teleost reference genomes as queries. The number of copies of each of the target genes was determined on the basis of the number of unique sequencing reads mapping to this region, relative to the number of reads matching each of the reference gene regions. The copy numbers of each of the reference gene regions were estimated first, using an iterative method and four different BLAST stringencies. Not all reference regions fulfilled our criteria, and some references were discarded for some species (see the Supplementary Note for details and Supplementary Table 11 for a full list of the references used for each species). Copy numbers for both MHC I lineages were then estimated by comparing the number of raw reads matching both the target and reference sequences and taking estimated genome size, coverage variation and total number of reads into account. The uncertainties of all copy number estimates were assessed with a double-bootstrapping procedure (Supplementary Note).
Rate shifts in MHC I copy number evolution.
Phylogenetic signal in MHC I copy number evolution was assessed with Blomberg's K statistic58, calculated using the phylosignal function of the picante R package v.1.6-2 (ref. 59), and with Pagel's lambda60, calculated with function phylosig of the phytools R package v.0.4-45 (ref. 61) (Supplementary Note). The fits of four general models of trait evolution were compared on the basis of their sample-size-corrected Akaike information criterion (AICc), using the function fitContinuous of the geiger R package v.2.0.3 (ref. 62): a white noise model, a Brownian motion model, an early-burst model63 and a single-peak Ornstein–Uhlenbeck model40,64 (Supplementary Note). The reversible-jump Bayesian approach of the bayou R package v.1.0.1 (ref. 65) was used to perform MCMC sampling of locations, magnitudes and numbers of shifts in multiple-optima Ornstein–Uhlenbeck models (Supplementary Fig. 5 and Supplementary Note). On the basis of the results of the bayou analysis, explicit hypotheses for shift combinations were tested in a likelihood framework, using the SLOUCH R package41,42. For each shift combination, the likelihood of the best fitting combination of optimum, half-life and stationary variance was recorded and used for model comparison based on AICc scores (Supplementary Table 13 and Supplementary Note). The ancestral states of log-transformed MHC I copy numbers were reconstructed for internal nodes of the time-calibrated phylogeny, on the basis of the best fitting Ornstein–Uhlenbeck model (Supplementary Note).
Diversification rate analyses.
Patterns of species diversification were analyzed with the Bayesian framework implemented in BAMM v.2.2.0 (ref. 66), on the basis of the time-calibrated phylogeny and counts of species richness in each of the 37 mutually exclusive clades of teleost fishes (Supplementary Table 14). The 'MEDUSA-like' model of diversification, assuming constant speciation and extinction rates within specific shift regimes67, was used for this analysis (Supplementary Fig. 8 and Supplementary Note). To test whether high MHC I copy numbers are associated with lineages that have high diversification rates, we carried out BiSSE analyses68 with the diversitree R package69. In these analyses, species were grouped into two categories for high and low MHC I copy numbers, on the basis of a given threshold value. Analyses were repeated for 26 equally spaced copy number threshold values between 10 and 60. As diversitree allows terminal clades with extant diversities of no more than 200 species, we used birth–death models of diversification in combination with the diversified sampling scheme of Höhna et al.70 to stochastically resolve subclades of all clades with more than 200 extant species, which was repeated 25 times. BiSSE analyses were conducted for each of the 25 resulting phylogenies and with each of the 26 copy number thresholds, assuming symmetric transition rates between high and low copy numbers and identical extinction rates in taxa with high and low copy numbers (Supplementary Note and Supplementary Data).
URLs.
GitHub repository for scripts used in this study, https://github.com/uio-cees/teleost_genomes_immune.
Accession codes.
All reads generated for this project have been deposited in the European Nucleotide Archive (ENA) under study accession PRJEB12469 (sample identifiers ERS1199874–ERS1199939). All new assemblies (unitigs and scaffolds) reported on here have been deposited in the Dryad repository under doi:10.5061/dryad.326r8.
Accession codes
References
Eschmeyer, W.N. & Fricke, R. Catalog of Fishes http://research.calacademy.org/research/ichthyology/catalog/fishcatmain.asp (2016).
Flajnik, M.F. & Kasahara, M. Origin and evolution of the adaptive immune system: genetic events and selective pressures. Nat. Rev. Genet. 11, 47–59 (2010).
Litman, G.W., Rast, J.P. & Fugmann, S.D. The origins of vertebrate adaptive immunity. Nat. Rev. Immunol. 10, 543–553 (2010).
Star, B. et al. The genome sequence of Atlantic cod reveals a unique immune system. Nature 477, 207–210 (2011).
Haase, D. et al. Absence of major histocompatibility complex class II mediated immunity in pipefish, Syngnathus typhle: evidence from deep transcriptome sequencing. Biol. Lett. 9, 20130044 (2013).
Malmstrøm, M., Jentoft, S., Gregers, T.F. & Jakobsen, K.S. Unraveling the evolution of the Atlantic cod's (Gadus morhua L.) alternative immune strategy. PLoS One 8, e74004 (2013).
Buonocore, F. & Gerdol, M. Alternative adaptive immunity strategies: coelacanth, cod and shark immunity. Mol. Immunol. 69, 157–169 (2016).
Neefjes, J., Jongsma, M.L.M., Paul, P. & Bakke, O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol. 11, 823–836 (2011).
Parham, P. Co-evolution of lymphocyte receptors with MHC class I. Immunol. Rev. 267, 1–5 (2015).
Roche, P.A. & Furuta, K. The ins and outs of MHC class II–mediated antigen processing and presentation. Nat. Rev. Immunol. 15, 203–216 (2015).
Kaufman, J., Völk, H. & Wallny, H.J.A. A “minimal essential Mhc” and an “unrecognized Mhc”: two extremes in selection for polymorphism. Immunol. Rev. 143, 63–88 (1995).
Woelfing, B., Traulsen, A., Milinski, M. & Boehm, T. Does intra-individual major histocompatibility complex diversity keep a golden mean? Phil. Trans. R. Soc. B 364, 117–128 (2009).
Nowak, M.A., Tarczy-Hornoch, K. & Austyn, J.M. The optimal number of major histocompatibility complex molecules in an individual. Proc. Natl. Acad. Sci. USA 89, 10896–10899 (1992).
Reusch, T.B., Häberli, M.A., Aeschlimann, P.B. & Milinski, M. Female sticklebacks count alleles in a strategy of sexual selection explaining MHC polymorphism. Nature 414, 300–302 (2001).
Milinski, M. The major histocompatibility complex, sexual selection, and mate choice. Annu. Rev. Ecol. Evol. Syst. 37, 159–186 (2006).
Klein, L., Kyewski, B., Allen, P.M. & Hogquist, K.A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don't see). Nat. Rev. Immunol. 14, 377–391 (2014).
Kalbe, M. et al. Lifetime reproductive success is maximized with optimal major histocompatibility complex diversity. Proc. Biol. Sci 276, 925–934 (2009).
Aeschlimann, P.B., Häberli, M.A. & Reusch, T. Female sticklebacks Gasterosteus aculeatus use self-reference to optimize MHC allele number during mate selection. Behav. Ecol. Sociobiol. 54, 119–126 (2003).
Bonneaud, C., Chastel, O., Federici, P., Westerdahl, H. & Sorci, G. Complex Mhc-based mate choice in a wild passerine. Proc. Biol. Sci 273, 1111–1116 (2006).
Eizaguirre, C. & Lenz, T.L. Major histocompatibility complex polymorphism: dynamics and consequences of parasite-mediated local adaptation in fishes. J. Fish Biol. 77, 2023–2047 (2010).
Eizaguirre, C., Lenz, T.L., Kalbe, M. & Milinski, M. Divergent selection on locally adapted major histocompatibility complex immune genes experimentally proven in the field. Ecol. Lett. 15, 723–731 (2012).
Karvonen, A. & Seehausen, O. The role of parasitism in adaptive radiations—when might parasites promote and when might they constrain ecological speciation? Int. J. Ecol. 2012, 1–20 (2012).
Eizaguirre, C., Lenz, T.L., Traulsen, A. & Milinski, M. Speciation accelerated and stabilized by pleiotropic major histocompatibility complex immunogenes. Ecol. Lett. 12, 5–12 (2009).
Betancur-R, R. et al. The tree of life and a new classification of bony fishes. PLoS Curr. 5, 1–45 (2013).
Roa-Varón, A. & Ortí, G. Phylogenetic relationships among families of Gadiformes (Teleostei, Paracanthopterygii) based on nuclear and mitochondrial data. Mol. Phylogenet. Evol. 52, 688–704 (2009).
Miller, J.R. et al. Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 24, 2818–2824 (2008).
Parra, G., Bradnam, K., Ning, Z., Keane, T. & Korf, I. Assessing the gene space in draft genomes. Nucleic Acids Res. 37, 289–297 (2009).
Simão, F.A., Waterhouse, R.M., Ioannidis, P., Kriventseva, E.V. & Zdobnov, E.M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Jarvis, E.D. et al. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014).
Miya, M. et al. Mitochondrial genome and a nuclear gene indicate a novel phylogenetic position of deep-sea tube-eye fish (Stylephoridae). Ichthyol. Res. 54, 323–332 (2007).
Chen, W.-J. et al. New insights on early evolution of spiny-rayed fishes (Teleostei: Acanthomorpha). Front. Mater. Sci. 1, e215 (2014).
Kruiswijk, C.P., Hermsen, T.T., Westphal, A.H., Savelkoul, H.F.J. & Stet, R.J.M. A novel functional class I lineage in zebrafish (Danio rerio), carp (Cyprinus carpio), and large barbus (Barbus intermedius) showing an unusual conservation of the peptide binding domains. J. Immunol. 169, 1936–1947 (2002).
Stet, R.J.M., Kruiswijk, C.P. & Dixon, B. Major histocompatibility lineages and immune gene function in teleost fishes: the road not taken. Crit. Rev. Immunol. 23, 441–471 (2003).
Dijkstra, J.M. et al. A third broad lineage of major histocompatibility complex (MHC) class I in teleost fish; MHC class II linkage and processed genes. Immunogenetics 59, 305–321 (2007).
Grimholt, U. et al. A comprehensive analysis of teleost MHC class I sequences. BMC Evol. Biol. 15, 32 (2015).
Pascual-Anaya, J., D'Aniello, S., Kuratani, S. & Garcia-Fernàndez, J. Evolution of Hox gene clusters in deuterostomes. BMC Dev. Biol. 13, 26 (2013).
Bonifacino, J.S. & Traub, L.M. Signals for sorting of transmembrane proteins to endosomes and lysosomes. Annu. Rev. Biochem. 72, 395–447 (2003).
Lizée, G., Basha, G. & Jefferies, W.A. Tails of wonder: endocytic-sorting motifs key for exogenous antigen presentation. Trends Immunol. 26, 141–149 (2005).
Star, B. & Jentoft, S. Why does the immune system of Atlantic cod lack MHC II? BioEssays 34, 648–651 (2012).
Hansen, T.F. Stabilizing selection and the comparative analysis of adaptation. Evolution 51, 1341–1351 (1997).
Hansen, T.F., Pienaar, J. & Orzack, S.H. A comparative method for studying adaptation to a randomly evolving environment. Evolution 62, 1965–1977 (2008).
Labra, A., Pienaar, J. & Hansen, T.F. Evolution of thermal physiology in Liolaemus lizards: adaptation, phylogenetic inertia, and niche tracking. Am. Nat. 174, 204–220 (2009).
Ohta, Y. & Flajnik, M.F. Coevolution of MHC genes (LMP/TAP/class Ia, NKT-class Ib, NKp30-B7H6): lessons from cold-blooded vertebrates. Immunol. Rev. 267, 6–15 (2015).
Wagner, C.E., Harmon, L.J. & Seehausen, O. Ecological opportunity and sexual selection together predict adaptive radiation. Nature 487, 366–369 (2012).
Wainwright, P.C. et al. The evolution of pharyngognathy: a phylogenetic and functional appraisal of the pharyngeal jaw key innovation in labroid fishes and beyond. Syst. Biol. 61, 1001–1027 (2012).
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–D669 (2015).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Davidson, W.S. et al. Sequencing the genome of the Atlantic salmon (Salmo salar). Genome Biol. 11, 403 (2010).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Leigh, J.W., Susko, E., Baumgartner, M. & Roger, A.J. Testing congruence in phylogenomic analysis. Syst. Biol. 57, 104–115 (2008).
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Matschiner, M. et al. Bayesian node dating based on probabilities of fossil sampling supports trans-Atlantic dispersal of cichlid fishes. Preprint at bioRxiv http://dx.doi.org/10.1101/038455 (2016).
Mirarab, S., Bayzid, M.S., Boussau, B. & Warnow, T. Statistical binning enables an accurate coalescent-based estimation of the avian tree. Science 346, 1250463 (2014).
Mirarab, S. et al. ASTRAL: genome-scale coalescent-based species tree estimation. Bioinformatics 30, i541–i548 (2014).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Blomberg, S.P. & Garland, T. Tempo and mode in evolution: phylogenetic inertia, adaptation and comparative methods. J. Evol. Biol. 15, 899–910 (2002).
Kembel, S.W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Pagel, M. Inferring the historical patterns of biological evolution. Nature 401, 877–884 (1999).
Revell, L.J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2011).
Pennell, M.W. et al. geiger v2.0: an expanded suite of methods for fitting macroevolutionary models to phylogenetic trees. Bioinformatics 30, 2216–2218 (2014).
Harmon, L.J. et al. Early bursts of body size and shape evolution are rare in comparative data. Evolution 64, 2385–2396 (2010).
Butler, M.A. & King, A.A. Phylogenetic comparative analysis: a modeling approach for adaptive evolution. Am. Nat. 164, 683–695 (2004).
Uyeda, J.C. & Harmon, L.J. A novel Bayesian method for inferring and interpreting the dynamics of adaptive landscapes from phylogenetic comparative data. Syst. Biol. 63, 902–918 (2014).
Rabosky, D.L. et al. BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees. Methods Ecol. Evol. 5, 701–707 (2014).
Alfaro, M.E. et al. Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proc. Natl. Acad. Sci. USA 106, 13410–13414 (2009).
Maddison, W.P., Midford, P.E. & Otto, S.P. Estimating a binary character's effect on speciation and extinction. Syst. Biol. 56, 701–710 (2007).
FitzJohn, R.G. Diversitree: comparative phylogenetic analyses of diversification in R. Methods Ecol. Evol. 3, 1084–1092 (2012).
Höhna, S., Stadler, T., Ronquist, F. & Britton, T. Inferring speciation and extinction rates under different sampling schemes. Mol. Biol. Evol. 28, 2577–2589 (2011).
Acknowledgements
All computational work was performed on the Abel Supercomputing Cluster (Norwegian Metacenter for High-Performance Computing (NOTUR) and the University of Oslo), operated by the Research Computing Services group at USIT, the University of Oslo IT Department. Sequencing library creation and high-throughput sequencing were carried out at the Norwegian Sequencing Centre (NSC), University of Oslo, Norway, and the McGill University and Génome Queéec Innovation Centre, Canada. We are grateful for the kindness of the following researchers for providing tissue samples for this study: K. Østbye (University of Oslo), F. Cortesi (University of Queensland), J.Y. Poulsen (Greenland Institute of Natural Resources), M. Miya (Natural History Museum and Institute, Chiba University), A. Bentley (University of Kansas Biodiversity Institute), I. Kilde (Norwegian University of Science and Technology, Trondheim), I.R. Bradbury (Memorial University), C. Pampoulie (Marine Research Institute of Iceland) and L. Rüber (Natural History Museum, Bern). This work was funded by grants from the Research Council of Norway (RCN grants 199806 and 222378) to K.S.J. M. Matschiner acknowledges funding from the Swiss National Science Foundation (SNF grant PBBSP3-138680), R.H. acknowledges funding from the German Federal Ministry of Food and Agriculture (grant AutoMAT, 2819100712) and W.S. acknowledges funding from the European Research Council (ERC). We thank G. Holm (University of Oslo) for the teleost illustrations, and B. Walenz and J.R. Miller for assistance with the Celera Assembler.
Author information
Authors and Affiliations
Contributions
M. Malmstrøm, K.S.J. and S.J. conceived the project. M. Malmstrøm, M. Matschiner, W.S., N.C.S., K.S.J. and S.J. planned and oversaw the project. M. Malmstrøm, R.H., W.S. and S.J. provided samples for genome sequencing. M. Malmstrøm prepared DNA and libraries for genome sequencing. M. Malmstrøm and S.J. were in charge of genome sequencing management. M. Malmstrøm, O.K.T., H.T.B. and A.J.N. performed genome assembly. M. Matschiner performed phylogenomic, diversification rate and comparative analyses. T.F.H. contributed to comparative analyses. M. Malmstrøm analyzed gene presence and absence. M. Malmstrøm and L.G.S. performed copy number estimation. M. Malmstrøm and M. Matschiner prepared the figures. M. Malmstrøm, M. Matschiner and B.S. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Maximum-likelihood teleost phylogenies.
Phylogenies are based on two different marker sets, the strictly filtered marker set including 567 exons from 111 genes (left) and the less strictly filtered marker set including 1,938 exons from 302 genes (right). All nodes are supported by 100% of bootstrap replicates, except where indicated.
Supplementary Figure 2 Time-calibrated teleost phylogeny with node numbers.
Age estimates and support values for each node are given in Supplementary Table 5.
Supplementary Figure 3 Mean copy number estimates of MHC I and Hox genes.
Shown are mean estimates for both numbers (MHC I and Hox), obtained with our copy number-estimation procedure. Estimates for these numbers are not correlated (linear model; P = 0.692; R2 = 0.002).
Supplementary Figure 4 Peptide sequence alignment of MHC I cytoplasmic tails containing putative signal motifs associated with MHC cross-presentation.
Alignment of MHC I cytoplasmic tail sequences from the sequenced species representing Gadinae, Lotinae and Phycinae. Amino acid positions range from 334 to 354. Dots indicate identity with the Phycis phycis sequence, and shaded areas show sequences with either ExxxLL/A or YxxL signaling motifs. The tree shows the topology of included species, with triangles connecting different sequences from the same species.
Supplementary Figure 5 Identification of optima shifts in MHC I copy number.
The sizes of circles indicate the marginal probability for shifts in the theta parameter on each branch in the time-calibrated phylogeny, while circle color represents the direction of the shift. The analysis was performed with the bayou R package v.1.0.1.
Supplementary Figure 6 BAMM phylorate plot for the most credible configurations of diversification rate shifts.
The eight most credible shift configurations are shown together with their posterior probability f. For each shift configuration, the locations of rate shifts are shown as gray circles, with circle size proportional to marginal shift probabilities. All of the eight most credible shift configurations include at least one shift at or near the base of Percomorphaceae, and six of the shift configurations include a shift on the branch leading to Gadiformes excluding Bregmacerotidae.
Supplementary Figure 7 Macroevolutionary cohort matrices for diversification.
Cell color indicates the pairwise probability that two species share the same diversification rate regime. The time-calibrated phylogeny is shown to the left and at the top of the cohort matrix. Similar rate regimes are inferred for Gadiformes excluding Bregmacerotidae, with more divergent rates for the gadiform clades Merlucciidae and Macrourinae. In addition, all members of Percomorphaceae share diversification rates with high pairwise probabilities, with slightly lower probabilities for shared rate regimes between Ophidiiformes or Batrachoidiformes and other members of Percomorphaceae.
Supplementary Figure 8 Branch-specific Bayes factor support for diversification rate shifts inferred with BAMM.
Shown is the teleost phylogeny with branch lengths proportional to Bayes factors (BF) supporting diversification rate shifts on branches. Rate shifts on branches leading to Percomorphaceae and Gadiformes excluding Bregmacerotidae are strongly supported with BF > 40, followed by additional rate shifts within these groups, on the stem branch of a clade combining all Percomorphaceae except Ophidiiformes and Batrachoidiformes (BF = 30.9) and on a branch leading to a clade combining Moridae, Bathygadinae and Macrourinae (BF = 12.6).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note. (PDF 3456 kb)
Supplementary Table 1
Sample information for all species sequenced for this study. (XLSX 55 kb)
Supplementary Table 2
Sequencing information for all species. (XLSX 49 kb)
Supplementary Table 3
Assembly statistics, gene space completeness metrics, and MHC and Hox copy number estimations for all species sequenced for this study. (XLSX 23 kb)
Supplementary Table 4
Phylogenetic marker information. (XLSX 118 kb)
Supplementary Table 5
Phylogenetic node support and age estimates. (XLSX 48 kb)
Supplementary Table 6
Orthologs used as queries for immune gene detection. (XLSX 53 kb)
Supplementary Table 7
UTG ID for each identified ortholog in all genomes investigated. (XLSX 78 kb)
Supplementary Table 8
MHC I reference sequences used for initial detection of MHC I variants in all genomes. (XLSX 44 kb)
Supplementary Table 9
Gene ID and contig ID for sequences used as references in MHC I and Hox copy number estimations. (XLSX 47 kb)
Supplementary Table 10
k-mer estimates of genome sizes for all sequenced species. (XLSX 60 kb)
Supplementary Table 11
References used for each of the species during MHC and Hox copy number estimation. (XLSX 53 kb)
Supplementary Table 12
UTG ID for the location of cytoplasmic signaling motifs. (XLSX 51 kb)
Supplementary Table 13
Model selection for shifts in MHC I copy number optima. (XLSX 88 kb)
Supplementary Table 14
Per-clade species richness used in the BAMM analysis. (XLSX 53 kb)
Supplementary Data: Background data for BiSSE analysis.
This folder contains 700 subfolders with individual data sets and R scripts used for the BiSSE analyses. (ZIP 15505 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Malmstrøm, M., Matschiner, M., Tørresen, O. et al. Evolution of the immune system influences speciation rates in teleost fishes. Nat Genet 48, 1204–1210 (2016). https://doi.org/10.1038/ng.3645
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3645
This article is cited by
-
Teleost genomic repeat landscapes in light of diversification rates and ecology
Mobile DNA (2023)
-
Comparative genomics of the T cell receptor μ locus in marsupials and monotremes
Immunogenetics (2023)
-
Divergent and non-parallel evolution of MHC IIB in the Neotropical Midas cichlid species complex
BMC Ecology and Evolution (2022)
-
Ancestral reconstruction reveals catalytic inactivation of activation-induced cytidine deaminase concomitant with cold water adaption in the Gadiformes bony fish
BMC Biology (2022)
-
False gene and chromosome losses in genome assemblies caused by GC content variation and repeats
Genome Biology (2022)