Abstract
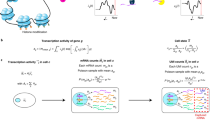
Most tissue samples are composed of different cell types. Differential expression analysis without accounting for cell-type composition cannot separate the changes due to cell-type composition or cell type-specific expression. We propose a computational framework to address these limitations: CARseq (cell-type-aware analysis of RNA-seq). CARseq employs a negative binomial distribution that appropriately models the count data from RNA-seq experiments. Simulation studies show that CARseq has substantially higher power than a linear model-based approach and it also provides more accurate estimate of the rankings of differentially expressed genes. We have applied CARseq to compare gene expression of schizophrenia/autism subjects versus controls, and identified the cell types underlying the difference and similarities of these two neuron-developmental diseases. Our results are consistent with the results from differential expression analysis using single-cell RNA-seq data.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data used in this study are available in the following public repositories.
snRNA-seq data for CT-specific expression reference were generated by the Allen Institute for Brain Science. File ‘http://human_mtg_gene_expression_matrices_2018-06-14.zip/’ was downloaded from http://celltypes.brain-map.org/api/v2/well_known_file_download/694416044.
The gene expression and clinical data of schizophrenia patients and healthy controls were generated by Common Mind Consortium (CMC) and the relevant data were obtained from the following links. Data access is governed by the NIMH Repository and Genomics Resources. CMC gene expression data, https://www.synapse.org/#!Synapse:syn3346749; CMC gene expression meta data, https://www.synapse.org/#!Synapse:syn18103174; CMC clinical data, https://www.synapse.org/#!Synapse:syn3275213.
The gene expression and clinical data of autism patients and healthy controls were part of The PsychENCODE (PEC) Capstone Collection https://www.synapse.org/#!Synapse:syn12080241 and the relevant data were obtained from the following links. Data access is governed by the NIMH Repository and Genomics Resources. UCLA-ASD gene expression data, https://www.synapse.org/#!Synapse:syn8365527; UCLA-ASD gene expression meta data, https://www.synapse.org/#!Synapse:syn5602933; UCLA-ASD clinical data, https://www.synapse.org/#!Synapse:syn5602932. The list of SFARI ASD risk genes was downloaded from https://gene.sfari.org/database/human-gene/.
Source Data for Figures 2–5 are available with this manuscript.
Code availability
The codes for generating the CT-specific gene expression reference panel are included in GitHub repository scRNAseq_pipelines (https://github.com/Sun-lab/scRNAseq_pipelines). We analyzed three scRNA-seq datasets: MTG, dronc and psychENCODE, and the codes were saved in corresponding folders. The codes to compare different references and generate final references were saved in ‘_brain_cell_type’ folder. The codes for CARseq analyses (including simulation and analyses of SCZ and ASD datasets) were included in GitHub repository CARseq_pipelines (https://github.com/Sun-lab/CARseq_pipelines). The file ‘reproducible_figures.html’ has the code to generate most figures in this paper. The R package CARseq was deposited at GitHub repository CARseq (https://github.com/Sun-lab/CARseq). All codes were also deposited in a Zendo repository36.
References
Nowakowski, T. J. et al. Expression analysis highlights AXL as a candidate Zika virus entry receptor in neural stem cells. Cell Stem Cell 18, 591–596 (2016).
Zhang, T. et al. Cell-type specific eQTL of primary melanocytes facilitates identification of melanoma susceptibility genes. Genome Res. 28, 1621–1635 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Leng, N. et al. EBSeq: an empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29, 1035–1043 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Shen-Orr, S. S. et al. Cell type-specific gene expression differences in complex tissues. Nat. Methods 7, 287–289 (2010).
Li, Z., Wu, Z., Jin, P. & Wu, H. Dissecting differential signals in high-throughput data from complex tissues. Bioinformatics 35, 3898–3905 (2019).
Zheng, S. C., Breeze, C. E., Beck, S. & Teschendorff, A. E. Identification of differentially methylated cell types in epigenome-wide association studies. Nat. Methods 15, 1059–1066 (2018).
Luo, X., Yang, C. & Wei, Y. Detection of cell-type-specific risk-CpG sites in epigenome-wide association studies. Nat. Commun. 10, 3113 (2019).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Wilson, D. R., Jin, C., Ibrahim, J. G. & Sun, W. ICeD-T provides accurate estimates of immune cell abundance in tumor samples by allowing for aberrant gene expression patterns. J. Am. Stat. Assoc. 115, 1055–1065 (2019).
Zhong, Y. & Liu, Z. Gene expression deconvolution in linear space. Nat. Methods 9, 8–9 (2012).
Cattane, N., Richetto, J. & Cattaneo, A. Prenatal exposure to environmental insults and enhanced risk of developing schizophrenia and autism spectrum disorder: focus on biological pathways and epigenetic mechanisms. Neurosci. Biobehav. Rev. 117, 253–278 (2018).
Anttila, V. et al. Analysis of shared heritability in common disorders of the brain. Science 360, eaap8757 (2018).
Prata, J., Santos, S. G., Almeida, M. I., Coelho, R. & Barbosa, M. A. Bridging autism spectrum disorders and schizophrenia through inflammation and biomarkers-pre-clinical and clinical investigations. J. Neuroinflammation 14, 179 (2017).
Jardri, R. et al. Are hallucinations due to an imbalance between excitatory and inhibitory influences on the brain? Schizophrenia Bull. 42, 1124–1134 (2016).
Aitchison, J. & Egozcue, J. J. Compositional data analysis: where are we and where should we be heading? Math. Geol. 37, 829–850 (2005).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Lin, M. et al. Heat shock alters the expression of schizophrenia and autism candidate genes in an induced pluripotent stem cell model of the human telencephalon. PLoS ONE 9, e94968 (2014).
Parikshak, N. N. et al. Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism. Nature 540, 423–427 (2016).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Petrelli, F., Pucci, L. & Bezzi, P. Astrocytes and microglia and their potential link with autism spectrum disorders. Front. Cell. Neurosci. 10, 21 (2016).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584 (2020).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Raymond, L. J., Deth, R. C. & Ralston, N. V. Potential role of selenoenzymes and antioxidant metabolism in relation to autism etiology and pathology. Autism Res. Treat. 2014, 164938 (2014).
Greenhalgh, A. D., David, S. & Bennett, F. C. Immune cell regulation of glia during cns injury and disease. Nat. Rev. Neurosci. 21, 139–152 (2020).
Regev, A. et al. Science forum: The Human Cell Atlas. eLife 6, e27041 (2017).
Kehrer, C., Maziashvili, N., Dugladze, T. & Gloveli, T. Altered excitatory-inhibitory balance in the NMDA-hypofunction model of schizophrenia. Front. Mol. Neurosci. 1, 6 (2008).
Ajram, L. et al. Shifting brain inhibitory balance and connectivity of the prefrontal cortex of adults with autism spectrum disorder. Transl. Psychiatry 7, e1137 (2017).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at BioRxiv https://doi.org/10.1101/060012 (2021).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, 1–12 (2010).
Jin, C., Chen, M., Lin, D. Y. & Sun, W. CARseq (2021); https://doi.org/10.5281/zenodo.4592636
Acknowledgements
We acknowledge the following grants: NIH R01GM105785 to W.S. and C.J., NIH R21CA224026 to W.S., NIH R01GM126550 to W.S., NIH R01HG009974 to D.-Y.L., NIH P01CA142538 to D.-Y.L., NIH R01GM126553 to M.C., NSF 2016307 to M.C. and a Sloan Foundation Fellowship to M.C. We also appreciate helpful discussions with P. Little.
Author information
Authors and Affiliations
Contributions
W.S. and C.J. conceived the approach. C.J. implemented the methods and performed analysis, with input from W.S., M.C. and D.-Y.L.; W.S. and C.J. wrote the paper, with input from M.C. and D.-Y.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Computational Science thanks Ruibin Xi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ananya Rastogi was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary notes, results, Tables 1–6 and Figs. 1–60.
Source data
Source Data Fig. 2
Source Data for Fig. 2.
Source Data Fig. 3
Source Data for Fig. 3.
Source Data Fig. 4
Source Data for Fig. 4.
Source Data Fig. 5
Source Data for Fig. 5.
Rights and permissions
About this article
Cite this article
Jin, C., Chen, M., Lin, DY. et al. Cell-type-aware analysis of RNA-seq data. Nat Comput Sci 1, 253–261 (2021). https://doi.org/10.1038/s43588-021-00055-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-021-00055-6
This article is cited by
-
ISLET: individual-specific reference panel recovery improves cell-type-specific inference
Genome Biology (2023)
-
CeDAR: incorporating cell type hierarchy improves cell type-specific differential analyses in bulk omics data
Genome Biology (2023)
-
Cell-type-specific co-expression inference from single cell RNA-sequencing data
Nature Communications (2023)
-
SCADIE: simultaneous estimation of cell type proportions and cell type-specific gene expressions using SCAD-based iterative estimating procedure
Genome Biology (2022)
-
Extracting insights from heterogeneous tissues
Nature Computational Science (2021)