Abstract
The glycolytic enzyme lactate dehydrogenase A (LDHA) is frequently overexpressed in cancer, which promotes glycolysis and cancer. The oncogenic effect of LDHA has been attributed to its glycolytic enzyme activity. Here we report an unexpected noncanonical oncogenic mechanism of LDHA; LDHA activates small GTPase Rac1 to promote cancer independently of its glycolytic enzyme activity. Mechanistically, LDHA interacts with the active form of Rac1, Rac1–GTP, to inhibit Rac1–GTP interaction with its negative regulator, GTPase-activating proteins, leading to Rac1 activation in cancer cells and mouse tissues. In clinical breast cancer specimens, LDHA overexpression is associated with higher Rac1 activity. Rac1 inhibition suppresses the oncogenic effect of LDHA. Combination inhibition of LDHA enzyme activity and Rac1 activity by small-molecule inhibitors displays a synergistic inhibitory effect on breast cancers with LDHA overexpression. These results reveal a critical oncogenic mechanism of LDHA and suggest a promising therapeutic strategy for breast cancers with LDHA overexpression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Publicly available datasets used in this study are TGCA (https://portal.gdc.cancer.gov/), Oncomine (https://www.oncomine.com/) and Cancer Cell Line Encyclopedia (https://sites.broadinstitute.org/ccle/). All data supporting the present study are available within the article and supplementary information files. Source data are provided with this paper.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Wolpaw, A. J. & Dang, C. V. Exploiting metabolic vulnerabilities of cancer with precision and accuracy. Trends Cell Biol. 28, 201–212 (2018).
Zhu, J. & Thompson, C. B. Metabolic regulation of cell growth and proliferation. Nat. Rev. Mol. Cell Biol. 20, 436–450 (2019).
Girgis, H. et al. Lactate dehydrogenase A is a potential prognostic marker in clear cell renal cell carcinoma. Mol. Cancer 13, 101 (2014).
Huang, X. et al. High expressions of LDHA and AMPK as prognostic biomarkers for breast cancer. Breast 30, 39–46 (2016).
Fantin, V. R., St-Pierre, J. & Leder, P. Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 9, 425–434 (2006).
Cui, J. et al. FOXM1 promotes the Warburg effect and pancreatic cancer progression via transactivation of LDHA expression. Clin. Cancer Res. 20, 2595–2606 (2014).
Zhao, D. et al. Lysine-5 acetylation negatively regulates lactate dehydrogenase A and is decreased in pancreatic cancer. Cancer Cell 23, 464–476 (2013).
Koukourakis, M. I. et al. Lactate dehydrogenase 5 expression in operable colorectal cancer: strong association with survival and activated vascular endothelial growth factor pathway–a report of the Tumour Angiogenesis Research Group. J. Clin. Oncol. 24, 4301–4308 (2006).
Lv, J. et al. Prognostic value of lactate dehydrogenase expression in different cancers: a meta-analysis. Am. J. Med. Sci. 358, 412–421 (2019).
Shim, H. et al. c-Myc transactivation of LDH-A: implications for tumor metabolism and growth. Proc. Natl Acad. Sci. USA 94, 6658–6663 (1997).
Le, A. et al. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl Acad. Sci. USA 107, 2037–2042 (2010).
Wang, Z. Y. et al. LDH-A silencing suppresses breast cancer tumorigenicity through induction of oxidative stress mediated mitochondrial pathway apoptosis. Breast Cancer Res. Treat. 131, 791–800 (2012).
Doherty, J. R. & Cleveland, J. L. Targeting lactate metabolism for cancer therapeutics. J. Clin. Investig. 123, 3685–3692 (2013).
Yeung, C. et al. Targeting glycolysis through inhibition of lactate dehydrogenase impairs tumor growth in preclinical models of Ewing sarcoma. Cancer Res. 79, 5060–5073 (2019).
Rizwan, A. et al. Relationships between LDH-A, lactate, and metastases in 4T1 breast tumors. Clin. Cancer Res. 19, 5158–5169 (2013).
Xie, H. et al. Targeting lactate dehydrogenase-A inhibits tumorigenesis and tumor progression in mouse models of lung cancer and impacts tumor-initiating cells. Cell Metab. 19, 795–809 (2014).
Martinez-Ordonez, A. et al. POU1F1 transcription factor induces metabolic reprogramming and breast cancer progression via LDHA regulation. Oncogene 40, 2725–2740 (2021).
Semenza, G. L. et al. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 271, 32529–32537 (1996).
Zhang, D. G., Zheng, J. N. & Pei, D. S. P53/microRNA-34-induced metabolic regulation: new opportunities in anticancer therapy. Mol. Cancer 13, 115 (2014).
Valvona, C. J., Fillmore, H. L., Nunn, P. B. & Pilkington, G. J. The regulation and function of lactate dehydrogenase A: therapeutic potential in brain tumor. Brain Pathol. 26, 3–17 (2016).
Feng, Y. et al. Lactate dehydrogenase A: a key player in carcinogenesis and potential target in cancer therapy. Cancer Med. 7, 6124–6136 (2018).
Brand, A. et al. LDHA-associated lactic acid production blunts tumor immunosurveillance by T and NK cells. Cell Metab. 24, 657–671 (2016).
Woodford, M. R., Chen, V. Z., Backe, S. J., Bratslavsky, G. & Mollapour, M. Structural and functional regulation of lactate dehydrogenase-A in cancer. Future Med. Chem. 12, 439–455 (2020).
Pan, C., Li, B. & Simon, M. C. Moonlighting functions of metabolic enzymes and metabolites in cancer. Mol. Cell 81, 3760–3774 (2021).
Dasgupta, S. et al. Metabolic enzyme PFKFB4 activates transcriptional coactivator SRC-3 to drive breast cancer. Nature 556, 249–254 (2018).
Enzo, E. et al. Aerobic glycolysis tunes YAP/TAZ transcriptional activity. EMBO J. 34, 1349–1370 (2015).
Hodge, R. G. & Ridley, A. J. Regulating Rho GTPases and their regulators. Nat. Rev. Mol. Cell Biol. 17, 496–510 (2016).
Kazanietz, M. G. & Caloca, M. J. The Rac GTPase in cancer: from old concepts to new paradigms. Cancer Res. 77, 5445–5451 (2017).
Liang, J. et al. Rac1, a potential target for tumor therapy. Front Oncol. 11, 674426 (2021).
Yue, X. et al. Gain-of-function mutant p53 activates small GTPase Rac1 through SUMOylation to promote tumor progression. Genes Dev. 31, 1641–1654 (2017).
Rhodes, D. R. et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 9, 166–180 (2007).
Gyorffy, B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct. Biotechnol. J. 19, 4101–4109 (2021).
Soderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Zhang, C. et al. Glutaminase 2 is a novel negative regulator of small GTPase Rac1 and mediates p53 function in suppressing metastasis. eLife 5, e10727 (2016).
Hayashi-Takagi, A. et al. Disrupted-in-schizophrenia 1 (DISC1) regulates spines of the glutamate synapse via Rac1. Nat. Neurosci. 13, 327–332 (2010).
Cardama, G. A. et al. Relevance of small GTPase Rac1 pathway in drug and radio-resistance mechanisms: opportunities in cancer therapeutics. Crit. Rev. Oncol. Hematol. 124, 29–36 (2018).
Fukata, M. et al. Rac1 and Cdc42 capture microtubules through IQGAP1 and CLIP-170. Cell 109, 873–885 (2002).
Feig, L. A. Tools of the trade: use of dominant-inhibitory mutants of Ras-family GTPases. Nat. Cell Biol. 1, E25–E27 (1999).
Joneson, T., White, M. A., Wigler, M. H. & Bar-Sagi, D. Stimulation of membrane ruffling and MAP kinase activation by distinct effectors of RAS. Science 271, 810–812 (1996).
Keely, P. J., Westwick, J. K., Whitehead, I. P., Der, C. J. & Parise, L. V. Cdc42 and Rac1 induce integrin-mediated cell motility and invasiveness through PI(3)K. Nature 390, 632–636 (1997).
Murphy, D. A. & Courtneidge, S. A. The ‘ins’ and ‘outs’ of podosomes and invadopodia: characteristics, formation and function. Nat. Rev. Mol. Cell Biol. 12, 413–426 (2011).
Augoff, K., Hryniewicz-Jankowska, A. & Tabola, R. Invadopodia: clearing the way for cancer cell invasion. Ann. Transl. Med. 8, 902 (2020).
Moshfegh, Y., Bravo-Cordero, J. J., Miskolci, V., Condeelis, J. & Hodgson, L. A trio-Rac1-Pak1 signalling axis drives invadopodia disassembly. Nat. Cell Biol. 16, 574–586 (2014).
Donnelly, S. K. et al. Rac3 regulates breast cancer invasion and metastasis by controlling adhesion and matrix degradation. J. Cell Biol. 216, 4331–4349 (2017).
Seals, D. F. et al. The adaptor protein Tks5/Fish is required for podosome formation and function, and for the protease-driven invasion of cancer cells. Cancer Cell 7, 155–165 (2005).
Yamaguchi, H. et al. Lipid rafts and caveolin-1 are required for invadopodia formation and extracellular matrix degradation by human breast cancer cells. Cancer Res. 69, 8594–8602 (2009).
Yamaguchi, H. et al. Phosphoinositide 3-kinase signaling pathway mediated by p110α regulates invadopodia formation. J. Cell Biol. 193, 1275–1288 (2011).
Attanasio, F. et al. Novel invadopodia components revealed by differential proteomic analysis. Eur. J. Cell Biol. 90, 115–127 (2011).
Gao, Y., Dickerson, J. B., Guo, F., Zheng, J. & Zheng, Y. Rational design and characterization of a Rac GTPase-specific small molecule inhibitor. Proc. Natl Acad. Sci. USA 101, 7618–7623 (2004).
Pajak, B. et al. 2-Deoxy-d-glucose and its analogs: from diagnostic to therapeutic agents. Int. J. Mol. Sci. 21, 234 (2019).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Wang, F. et al. Glycolytic stimulation is not a requirement for M2 macrophage differentiation. Cell Metab. 28, 463–475 (2018).
Zhang, C. et al. Tumour-associated mutant p53 drives the Warburg effect. Nat. Commun. 4, 2935 (2013).
Pulaski, B. A. & Ostrand-Rosenberg, S. Mouse 4T1 breast tumor model. Curr. Protoc. Immunol. 20, Unit 20.2 (2001).
Wan, L., Pantel, K. & Kang, Y. Tumor metastasis: moving new biological insights into the clinic. Nat. Med. 19, 1450–1464 (2013).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Pioli, P. A., Hamilton, B. J., Connolly, J. E., Brewer, G. & Rigby, W. F. Lactate dehydrogenase is an AU-rich element-binding protein that directly interacts with AUF1. J. Biol. Chem. 277, 35738–35745 (2002).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Liu, J. et al. Parkin ubiquitinates phosphoglycerate dehydrogenase to suppress serine synthesis and tumor progression. J. Clin. Invest. 130, 3253–3269 (2020).
Liu, J. et al. Parkin targets HIF-1α for ubiquitination and degradation to inhibit breast tumor progression. Nat. Commun. 8, 1823 (2017).
Chang, C. Y. et al. Tumor suppressor p53 regulates intestinal type 2 immunity. Nat. Commun. 12, 3371 (2021).
Pellegrin, S. & Mellor, H. Rho GTPase activation assays. Curr. Protoc. Cell Biol. 14, Unit 14.18 (2008).
Dawson, N. J., Bell, R. A. & Storey, K. B. Purification and properties of white muscle lactate dehydrogenase from the anoxia-tolerant turtle, the red-eared slider, trachemys scripta elegans. Enzym. Res. 2013, 784973 (2013).
Javed, M. U., Yousuf, F. A., Hussain, A. N., Ishaq, M. & Waqar, M. A. Purification and properties of lactate dehydrogenase from liver of Uromastix hardwickii. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 111, 27–34 (1995).
Chou, T. C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010).
Chou, T. C. & Talalay, P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv. Enzym. Regul. 22, 27–55 (1984).
Baik, M. et al. Identification of invadopodia by TKS5 staining in human cancer lines and patient tumor samples. MethodsX 6, 718–726 (2019).
Wang, Y. H. et al. Cell-state-specific metabolic dependency in hematopoiesis and leukemogenesis. Cell 158, 1309–1323 (2014).
Zhen, C. et al. Gankyrin promotes breast cancer cell metastasis by regulating Rac1 activity. Oncogene 32, 3452–3460 (2013).
Acknowledgements
LC–MS/MS proteomic analysis was performed at the Biological Mass Spectrometry facility of Rutgers University. This work was supported in part by grants from the National Institutes of Health (R01CA227912 and R01CA214746 to Z.F., as well as R01CA203965 and R01CA260837 to W.H.) and Congressionally Directed Medical Research Programs (CA214746 to Z.F.). T.Z. and C.C. were supported by the postdoctoral fellowship from New Jersey Commission on Cancer Research.
Author information
Authors and Affiliations
Contributions
J.L., C.Z., T.Z., C.C., J.W. and L.B. performed the experiments and analyzed data. L.Z. and B.G.H. analyzed data and contributed important materials. W.H. and Z.F. conceived and supervised the study. J.L., W.H. and Z.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks William J. Muller, Taro Hitosugi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Giménez-Cassina, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LDHA overexpression and its association with clinical outcomes in different subtypes of breast cancers classified by the status of ER, PR or HER2.
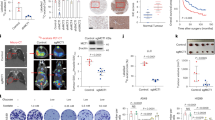
a-c, The increased LDHA mRNA levels in different subtypes of breast cancers compared with matched adjacent non-tumor breast tissues. The data were obtained from TCGA and the P-value was analyzed by two-tailed paired Student’s t-test. d-f, High LDHA mRNA expression is associated with poor relapse-free survival in patients with different subtypes of breast cancers. The data were obtained from Kaplan-Meier plotter (http://kmplot.com) and analyzed by the log-rank (Mantel–Cox) test. ER: estrogen receptor; PR: progesterone receptor.
Extended Data Fig. 2 LDHA displayed a much weaker interaction with Rac1-T35S mutant compared with WT Rac1 in cells.
Hs578T cells with ectopic expression of LDHA-Flag and WT Myc-Rac1, Myc-Rac1-T17N, or Myc-Rac1-T35S were employed for co-IP assays followed by western-blot assays. Data represent three repeats with similar results.
Extended Data Fig. 3 The dominant negative Rac1-T17N mutant reduces the promoting effect of WT and L4 LDHA on colony formation, migration and invasion of breast cancer cells.
a-c, Hs578T and SK-BR3 cells with ectopic expression of WT or L4 LDHA were transduced with control or Rac1-T17N expression vectors for colony formation (a), migration (b) and invasion (c) assays. Data represent mean ± s.d. (n = 6 independent experiments), two-way ANOVA followed by Tukey’s or Bonferroni’s test. *: P < 0.0001.
Extended Data Fig. 4 LDHA but not Rac1 is localized in invadopodia in breast cancer cell lines.
HS578T and MDA-MB231 cells seeded on gelatin-coated coverslips were labeled with anti-LDHA or Rac1 in far-red, anti-Tks5 in green and phalloidin (to stain F-actin) in red. The co-localization of Tks5 and F-actin in a punctate manner in cells was used as an indication of invadopodia formation. Arrows indicate invadopodia in cells. Scale bars: 20 μm. Data represent three repeats with similar results.
Extended Data Fig. 5 The potential role of different Rho family proteins in mediating the oncogenic effect of LDHA in breast cancer cells.
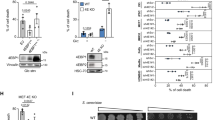
a, The interaction between LDHA with different Rho family proteins analyzed by co-IP assays. Hs578T cells expressing LDHA-Flag and Myc-Rac1, Myc-Cdc42, Myc-Rac3, or Myc-RhoA were used for co-IP and western-blot assays. b, LDHA activated Rac3 and Cdc42, but at a much less extent compared with its effect on Rac1. Hs578T and SK-BR3 cells expressing LDHA-Flag were used for PAK-PBD pull-down assays to measure the levels of GTP-bound Rac1, Rac3 and Cdc42. Left panels: represented results. Right panels: relative Rac1, Rac3 and Cdc42 activities analyzed by comparing the levels of GTP-bound Rac1, Rac3 and Cdc42 to the levels of total Rac1, Rac3 and Cdc42 proteins, respectively, in cells. Data represent n = 3 independent experiments, two-tailed unpaired Student’s t-test. c, Knockdown of Rac3 or Cdc42 displayed a much less pronounced effect on colony formation, migration and invasion of breast cancer cells with WT or L4 LDHA expression compared with Rac1 knockdown. Data represent mean ± s.d. (n = 6 independent experiments), two-way ANOVA followed by Tukey’s test or Dunnett’s test. d, Knockdown of Rac3 and Cdc42 in cells was confirmed by Taqman real-time PCR assays. Their mRNA levels were normalized with Actin. Data represent n = 3 independent experiments, one-way ANOVA followed by Dunnett’s test. *: P < 0.0001.
Extended Data Fig. 6 The relative expression levels of Rac1, Rac3 and Cdc42 in breast cancer cells.
Rac1 has a much higher expression level than Rac3 and Cdc42 in majority of breast cancer cell lines, including cell lines used in this study (labeled in red) as shown by the RNA-Seq data from Cancer Cell Line Encyclopedia (CCLE; https://sites.broadinstitute.org/ccle/).
Extended Data Fig. 7 LDHA and Rac1 small-molecule inhibitors display a much more pronounced inhibitory effect on colony formation, migration and invasion of breast cancer cells.
a, The combination treatment displayed a much more pronounced inhibitory effect on colony formation of breast cancer cells. BT-549, MCF7 and ZR-75-1 cells were treated with indicated concentrations of FX11 and/or NSC23766 for 4 days before colony formation assays. Combo: FX11 + NSC23766. b, c, The combination treatment displayed a much more pronounced inhibitory effect on migration (b) and invasion (c) of breast cancer cells than the single inhibitor treatment as analyzed by Transwell assays. In b, c, BT-549, MCF7 and ZR-75-1 cells were treated with the indicated concentrations of FX11 and/or NSC23766 for 24 h. NSC: NSC23766. Data represent mean ± s.d. (n = 6 independent experiments), one-way ANOVA followed by Tukey’s test. *: P < 0.0001.
Extended Data Fig. 8 The effect of small-molecule inhibitor treatments on the body weights of tumor-bearing mice.
a, Small-molecule inhibitor treatments did not significantly affect the body weights of female nude mice bearing orthotopic tumors formed by WT Hs578T cells. b, Small-molecule inhibitor treatments did not significantly affect the body weights of female BALB/c mice bearing orthotopic tumors formed by 4T1 cells transduced with the control lentiviral shRNA vector. Mice were treated with FX11 (1 mg/kg/day; i.p.; once/day) and/or NSC23766 (1.5 mg/kg/day, i.p.; once/day), or vehicle (−). Mice with Hs578T tumors were treated for 3 weeks and mice with 4T1 tumors were treated for 2 weeks. The body weights of mice were measured and recorded at the days indicated. Data represent mean ± s.d. n=8 mice/group. Statistical differences were determined by two-way ANOVA followed by Tukey’s test.
Extended Data Fig. 9 The effect of FX11 and/or NSC23766 on the growth and lung metastasis of mammary tumors in MMTV-PyMT mice.
a, The effects of FX11 and/or NSC23766 on the growth and metastasis of mammary tumors in 10-week-old MMTV-PyMT mice. The 10-week-old female MMTV-PyMT mice that developed late-stage tumors were treated with FX11 (1 mg/kg/day; i.p.; once/day) and/or NSC23766 (1.5 mg/kg/day, i.p.; once/day) for 3 weeks before they were sacrificed for analysis. Data represent mean ± s.d.; n=6 mice/group. One-way ANOVA followed by Student’s t-test. Con: vehicle; NSC: NSC23766; combo: FX11 + NSC. b, The interaction between endogenous LDHA and Rac1 in both non-tumor and tumor mammary tissues of MMTV-PyMT mice detected by co-IP and western-blot assays. Non-tumor mammary tissues were obtained from 5-week-old MMTV-PyMT mice and mammary tumor tissues were obtained from 10-week-old MMTV-PyMT mice. Normal mammary tissues from 5-week-old LDHA-deficient mice (the R26-Cre-ERT2, LDHAflox/flox mice treated with Tamoxifen to delete LDHA as presented in Fig. 2h) were used as negative controls for assays. NC: negative control.
Extended Data Fig. 10 LDHA expression is not linked to any specific subtypes of breast cancer.
LDHA protein levels in two breast cancer TMAs, including TMA-RCINJ (a-c; n=200) and TMA-BR2082a (d-f; n=120), were analyzed by IHC staining and were compared in different cancer subtypes classified by the status of ER (a, d), PR (b, e), or HER2 (c, f). Statistical studies were performed by using χ2 test. The TMA-BR2161 does not have information on the status of ER, PR or HER2.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Tables 1 and 2 and unprocessed blots for Supplementary Figs.
Supplementary Data 1
Supplementary statistical source data.
Source data
Source Data Fig. 1
Unprocessed western blots for Fig. 1.
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Unprocessed western blots for Fig. 2.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Unprocessed western blots for Fig. 3.
Source Data Fig. 4
Unprocessed western blots for Fig. 4.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 5
Unprocessed western blots for Fig. 5.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Fig. 6
Statistical source data for Fig. 6.
Source Data Fig. 7
Statistical source data for Fig. 7.
Source Data Extended Data Fig. 2
Unprocessed western blots for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 5
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Unprocessed western blots for Extended Data Fig. 9.
Source Data Extended Data Fig. 9
Statistical source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Zhang, C., Zhang, T. et al. Metabolic enzyme LDHA activates Rac1 GTPase as a noncanonical mechanism to promote cancer. Nat Metab 4, 1830–1846 (2022). https://doi.org/10.1038/s42255-022-00708-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00708-4
This article is cited by
-
Targeting small GTPases: emerging grasps on previously untamable targets, pioneered by KRAS
Signal Transduction and Targeted Therapy (2023)
-
Beyond Warburg: LDHA activates RAC for tumour growth
Nature Metabolism (2022)