Abstract
The study of developmental processes in Rhodnius prolixus has recently advanced with the sequencing of the genome. In this work, we analyze the maternal gene expression driving oogenesis and early embryogenesis in R. prolixus. We examined the transcriptional profile of mRNAs to establish the genes expressed across the ovary, unfertilized eggs and different embryonic stages of R. prolixus until the formation of the germ band anlage (0, 12, 24, and 48 h post egg laying). We identified 81 putative maternal and ovary-related genes and validated their expression by qRT-PCR. We validate the function of the ortholog gene Bicaudal-D (Rp-BicD) by in situ hybridization and parental RNAi. Consistent with a role in oogenesis and early development of R. prolixus, we show that lack of Rp-BicD does not significantly affect oogenesis but impairs the formation of the blastoderm. Based on our findings, we propose three times of action for maternal genes during oogenesis and embryogenesis in R. prolixus.
Similar content being viewed by others
Introduction
During insect embryogenesis, a sequential series of dynamic processes that include cell division, growth and fate specification take place to establish the necessary components to give rise to a complete organism, playing a fundamental role to support the developmental process of the whole life cycle1,2,3.
There are three modes of insect embryogenesis: long, intermediate, and short germ embryogenesis4. Long germ embryogenesis is defined by the simultaneous establishment of all segmental fates at the blastoderm stage. This is a derived mode of embryogenesis, found in scattered species among the Holometabola, such as Drosophila melanogaster. These insects have polytrophic meroistic ovaries. In short or intermediate germ insects only the most anterior segments are specified before gastrulation, while the more posterior segments are generated and patterned progressively from a posterior region called the growth zone. This represents an ancestral type of insect embryogenesis, described in insect models such Oncopeltus fasciatus, Rhodnius prolixus, Bombyx mori, Tribolium castaneum. It corresponds to insects with telotrophic or panoistic ovaries4,5,6,7,8. A common feature across the different modes of embryogenesis is the loading of maternal mRNA transcripts and proteins in the egg during oogenesis9.
In the last 20 years the rise of new models for comparative insect development provided a framework to understand the genetic basis of development and evolution10,11,12,13,14,15,16,17. The different mechanisms of insect embryogenesis are determined by specific spatiotemporal gene expression patterns derived from common genetic programs, suggesting that the mechanisms are much more conserved than the diversity of germ types might suggest4,18,19,20,21,22. In addition, a detailed study of cell flow during germ band extension and the fate map of T. castaneum embryo led to the idea that short and long germ bands share many more common features than thought23. In the last decade the expanse of genomics and transcriptomic analysis provided an insight of the transcriptional basis of the embryonic development in non-model insect species24. However, the complete repertory of genes involved in oogenesis and early embryogenesis has been reported in detail only in D. melanogaster25,26,27,28,29 and T. castaneum15,30,31, remaining an open question in other model organisms.
The blood-feeding insect Rhodnius prolixus is one vector of Trypanosoma cruzi, the etiologic agent of Chagas disease32,33. In addition to its medical interest, it has been a classical model for physiology and biochemistry34,35,36,37. The embryonic development of R. prolixus has been described from fertilization to hatching7, and the process of oogenesis studied in detail38,39,40,41,42,43,44. The genome was recently sequenced45 and since then, R. prolixus is an emerging model for developmental biology46,47,48,49,50. Several transcriptome analyses were reported, focusing in the gene expression of the follicular epithelium, the early previtellogenic stage of oogenesis; as well, in the impact of the nutritional state on regulatory pathways associated with reproductive performance51,52,53. Very recently, a thorough study on previtellogenic ovaries and unfertilized eggs discovered a large number of unannotated genes in the R. prolixus genome and unveiled a large set of maternal genes54. With all this knowledge in place, we have moved forward to the understanding of the genetic and molecular mechanisms driving oogenesis and the maternal contribution to embryo patterning in R. prolixus. Here, we present a transcriptome profiling approach to identify the genetic basis underlying oogenesis and early embryogenesis of R. prolixus until the onset of gastrulation, with a focus on genes related to embryonic patterning and egg formation. We provide novel insight into the molecular basis of early embryo formation and show the dynamic of mRNA expression during early embryo development in R. prolixus. Our study provides maternal and early embryonic transcriptomes of this hemimetabolous insect. We present a comprehensive qualitative data about genes related to segmentation, dorsal ventral axis and oogenesis, validate gene expression by qRT-PCR and show the phenotype of Bicaudal D (BicD) homolog likely related to early steps of the maternal cascade that leads to patterning.
Materials and methods
Insect rearing
A colony of R. prolixus was maintained in our laboratory in a 12:12 h light/dark period at 28 °C and 80% relative humidity in controlled environment incubators. In these conditions, embryogenesis takes 14 ± 1 days. Insects were regularly fed on chickens, which were housed, cared, fed and handled in accordance with resolution 1047/2005 (Consejo Nacional de Investigaciones Científicas y Técnicas, CONICET) regarding the bioethical framework for biomedical research with laboratory, farm, and nature collected/wild animals. This framework is in accordance with international standard procedures. Biosecurity considerations agree with CONICET resolution 1619/2008, which is in accordance with the WHO Biosecurity Handbook (ISBN 92 4 354 6503).
Sample collection, RNA isolation and sequencing
Adult mated insects 6th days after the feeding regimen were used to collect fertilized eggs at specific points in developmental time—0 (zygote), 12 (blastoderm), 24 (cellular blastoderm) and 48 (onset of germ band formation) hours post egg laying (hPL). Virgin female adults were used to collect unfertilized eggs, which were immediately frozen and stored in liquid nitrogen. At the same time, female ovaries were dissected in the vitellogenic stage and placed in a cryotube containing Trizol (Invitrogen), flash frozen and stored in liquid nitrogen until use.
For the transcriptome profiling, RNA was extracted from 150 embryos for each developmental time and 30 vitellogenic ovaries. For qRT-PCR analysis, independent experiments were carried out using 75 embryos from each specific time and 10 vitellogenic ovaries. Total RNA was isolated using Trizol (Invitrogen) as recommended by the manufacturer. RNA integrity was determined by agarose electrophoresis and concentration measured using Qubit RNA Assay Kit in a Qubit 2.0 Fluorometer (Life Technologies, Invitrogen). cDNA libraries were synthetized from 1 µg of total RNA and sequenced using a HiSeq-3000 platform (Illumina) to obtain the 50 base pairs (bp) (single-end) or 150 bp (paired-end) reads. The RNA-seq data has been submitted to the NCBI SRA database, available under accession code PRJNA694974.
Quality control, alignment and transcriptome assembly
Raw data were processed with FASTX-toolkit software (http://hannonlab.cshl.edu/fastx_toolkit/), to remove adapter sequences, reads with unknown bases and reads with quality scores lower than Q30, showed by the FastQC report (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). To avoid contaminants, the presence of adaptor sequences was ruled out using BLASTn55 and the UniVec database (ftp://ftp.ncbi.nlm.nih.gov/pub/UniVec/) from NCBI. Additionally, to remove rRNA sequences the SILVA database was used56. The remaining reads were defined as clean reads and used for subsequent bioinformatics analyses. TopHat257 was used to map clean reads to the ab initio annotations of R. prolixus, genome dataset version RproC3.345,58. The mapping statistics by the RNA-seq reads were calculated by using bam_stat.py implemented in the RSeQC package59 and the advanced statistics of coverage analysis were performed by the Qualimap application60. After TopHat alignment, transcripts were assembled using Cufflinks61,62. Assembly quality was assessed for each assembly using BUSCO analysis63, with the reference gene set of arthropods (2676 proteins) with default parameters. Fasta Statistics was used to display summary statistics from each transcriptome generated64. The eggNOG 5.0 database was used for functional annotation of the transcripts with common denominators or functional categories (i.e., derived from the original COG categories). Also, predicted protein-coding transcripts65 were functionally annotated. For each protein sequence protein signatures were assigned, using InterProScan search Version 5.0.066, through the PfamA and SuperFamily databases. Proteins annotated by signatures were assigned into GO (Gene Ontology) categories, including biological processes (BP), molecular functions (MF) and cellular components (CC). To statistically analyze GO-term enrichment, topGO package67 was implemented, using Fisher’s exact test and the false discovery rate (FDR) adjusted method. A q-value smaller than 0.05 were considered as significant. The reference set of gene-to-GO mappings was available from VectorBase (https://www.vectorbase.org/).
Oogenesis and early embryogenesis gene identification
Gene identification was performed using local BLAST55 on the six transcriptome assemblies. The BLAST algorithm used was BLASTx. The search was limited to 84 protein sequences derived from FlyBase (Version FB2020_03, https://flybase.org/), comprising genes related to oogenesis and early embryogenesis, with an e-value threshold of 10−5. Transcript with blast hit to Drosophila were then manually checked by BLASTx against all Arthropoda protein sequences (NCBI non-redundant protein (nr) database, assessed January 2018) to confirm sequence identity. BLAST results were classified into the known D. melanogaster developmental process. For the maternal gene search, a database was generated from different resources containing 10,277 specific protein sequences (Additional file 1)15,25,26,27,28,29.
Quantitative real-time PCR
Total RNA was isolated using Trizol reagent (Invitrogen) and treated with DNAse (QIAGEN). cDNA was synthesized using SuperScript™ VILO™ MasterMix kit (Invitrogen) following the manufacturer’s instructions. PCR was performed in technical triplicates (3 wells/cDNA sample), in a 10 μl final volume as follows: (i) 95 °C for 10 min; (ii) 95 °C for 15 s; (iii) 55 °C for 30 s; (iv) 72 °C for 45 s; (v) steps (ii) to (iv) for 35 cycles. Gene expression level was quantified using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad) in an Applied Biosystems 7500 Real-Time PCR System (Thermo Fisher Scientific). A control without a template was included in all batches and α-tubulin was used as reference, after a screen of several housekeeping gene candidates, as it provided consistent results on the embryonic stages analyzed. All primer pairs (Additional file 2) were tested for dimerization, efficiency, and amplification of a single product. The Ct value was averaged for the technical triplicate experiments and subtracted from the average Ct of the reference gene, to yield the expression difference (dCt) for each biological replicate. The results were analyzed according to68. To test whether the expression of a given gene was significantly different across developmental times, a one-way ANOVA was carried out followed by post-hoc test using GraphPad Prism v6.0 software (GraphPad Software, CA, USA, www.graphpad.com).
In situ Hybridization and Parental RNAi
DNA templates used to synthesize in situ hybridization probes were obtained by PCR using oligonucleotides carrying T7 promoter sequences at the 5'-end. The templates, bearing T7 promoter either in the sense or antisense direction, were in vitro transcribed using DIG RNA labeling kit (Roche) to produce sense and antisense probes. In situ hybridization was performed as described in Pascual et al.50. Female ovaries (n = 11) were dissected during the vitellogenic stage and eggs (n = 40) collected at specific developmental times.
For parental RNAi, dsRNA was produced by simultaneous transcription with T7 RNA polymerase (New England Biolabs) on templates containing T7 promoter sequences at both ends. The amplicons were sequenced to confirm identity (Macrogen Inc.). dsRNABicD was quantitated by fluorescence and injected into virgin females as described in Lavore et al.69. Two days after injection, the females were fed to induce oogenesis and mated. After mating, eggs were collected and ovaries fixed as described50. Phenotypic analysis of the ovaries was always performed at the vitellogenic stage. A negative control was performed injecting virgin females with dsRNA corresponding to the β-lactamase gene (dsRNAβ-lac) of E. coli69. Oligonucleotides used in this study are listed in Additional file 2.
Results and discussion
Assembling the ovarian and early embryonic transcriptomes of R. prolixus: characterization and completeness analysis
The RNA-seq output comprises six R. prolixus samples that cover late oogenesis to the beginning of germ band extension (48 hPL). Statistics on the sequencing and mapping are reported in Table 1A. According to completeness analysis, the coverage metrics obtained indicate that the assembled transcriptomes are sufficient for a meaningful analysis (Table 1B). As the genomic reference has a reasonable number of positions that are not called, transcriptomes assembled by mapping genomic predictions (ab initio) were used for subsequent analyses. In this respect, a review of zygotic genes14 and of the regulatory pathways involved in egg production52 has been reported based on these genomic annotations with a robust gene identification.
In order to conduct a transcriptome-composition representation analysis, eggNOG analysis was performed. A total of 25 eggNOG categories were detected (Fig. 1 and Additional file 3), in which the category “function unknown” was dominant followed by “Signal transduction mechanisms” and “Post-translational modification, protein turnover, and chaperones” in all the analyzed developmental times. To obtain information of the predicted proteins, InterproScan searches were performed to identify functional domains, repeats, sites and protein families conserved in the protein-coding transcripts (Table 2 and Additional file 4). For all of the characterized transcripts, statistically over-represented GO terms were identified using the FDR adjusted relative to a reference set of 11,947 genes. These statistically highlighted GO terms were summarized to generic GO categories for each developmental time studied (Table 2 and Additional file 5). GO analysis showed that mainly metabolic processes were enriched during embryo development, such as cellular macromolecule metabolic process (GO: 0044260), nucleobase-containing compound metabolic process (GO: 0006139), organonitrogen compound biosynthetic process (GO: 1901566), gene expression (GO: 0010467). This enrichment is in agreement with the requirements of the embryo during the transitions between the different embryonic stages with rapidly changing of the anabolic and catabolic demands70,71.
Classification of eggNOG annotations in the R. prolixus transcriptomes. The capital letters on the X-axis represent different eggNOG categories. Y-axis shows the number of transcripts in each eggNOG category. A: “RNA processing and modification”, B: “Chromatin structure and dynamics”, C: “Energy production and conversion”, D: “Cell cycle control, cell division, chromosome partitioning”, E: “Amino acid transport and metabolism”, F: “Nucleotide transport and metabolism”, G: “Carbohydrate transport and metabolism”, H: “Coenzyme transport and metabolism”, I: “Lipid transport and metabolism”, J: “Translation, ribosomal structure and biogenesis”, K: “Transcription”, L: “Replication, recombination and repair”, O: “Post-translational modification, protein turnover, and chaperones”, P: “Inorganic ion transport and metabolism”, Q: “Secondary metabolites biosynthesis, transport, and catabolism”, R: “General function prediction only”, S: “Function unknown”, T: “Signal transduction mechanisms”, U: “Intracellular trafficking, secretion, and vesicular transport”, V: “Defense mechanisms”, W: “Extracellular structures”, Y: “Nuclear structure”, Z: “Cytoskeleton”.
A total of 1192 annotated transcripts common to all developmental stages studied were further examined to determine GO enrichment. The represented GO terms (Fig. 2 and Additional file 6) were categorized in two main groups: cellular components and biological processes. The main terms of cellular components are cell structures related to protein synthesis, while the terms of biological processes are related to lipid, carbohydrate, nucleic acid and protein metabolism, all significantly over-represented. As expected in the enrichment observed for each developmental time, these biological processes play key roles in the embryonic development of R. prolixus. Energy is supplied during embryogenesis by the breakdown of biomolecules stored in the yolk72. This, in turn, drives the anabolic pathways such as protein and nucleotide biosynthesis to meet the needs of the developing embryo70. These also agree with the increment of lipid, protein and carbohydrate biosynthesis reported during embryonic stages of T. castaneum, Boophilus microplus and Aedes aegypti73,74,75, and in fed females of R. prolixus and other triatomines76,77,78,79.
Gene identification for developmental processes
In order to study genes involved in early embryogenesis and oogenesis, a list of 84 developmental genes (37 segmentation-related genes, 16 of the dorsal patterning pathway and 31 linked to oogenesis) related to these processes in D. melanogaster was used to identify ortholog sequences in the R. prolixus transcriptomes (Table 3A-C). The approach identified 81 expressed genes conserved in R. prolixus. We cannot rule out that the absence of a transcript is due to an incomplete transcriptome coverage rather than evolutionary divergence. It is plausible to consider that some genes are expressed at very low levels or in a small subset of cells in the developmental stages analyzed and that enrichment analysis may be necessary for their identification. The genes gurken (grk), bicoid (bcd), oskar (osk), were not included in the search because they have been reported absent in triatomines14. We observed in some cases, as we previously had50, that the reads that correspond to a single assembled transcript correspond to two different gene ID. Rather than corresponding to duplicated genes (paralogs), manual curation and assembly of the sequences revealed that both gene ID corresponded to a single transcriptional unit. Our work and the remarkable progress in the annotation of early genes by Coelho et al.54 suggest that the current version of the genome and the annotated gene IDs need to be revised or that the genome needs long read resequencing to fill gaps in the assembled sequence.
One would expect that genes related to oogenesis should not be expressed in embryonated eggs at gastrulation and germ band formation (48 hPL), as well as zygotic expression genes should not be expressed in unfertilized eggs. However, our analysis of different developmental stages revealed a number of genes expressed throughout early embryogenesis. Thus, all genes showed maternal expression. These results, although surprising, would agree with the role of the maternally active genome as director of most of early animal development15,80,81,82,83. In D. melanogaster and T. castaneum a high percentage of maternal transcripts are deposited during oogenesis, and so were detected (approximately 58–65%) in the course of the first hours of embryo development, while zygotic transcription was not detected15,25,27,83. These maternal mRNAs have been reported to have several functions, including the establishment of polarization gradients84,85,86, segregation of cell-fate determinants87,88,89,90,91,92,93,94,95, and targeting of protein synthesis to specialized organelles or cellular domains96,97,98,99,100.
To gain insight into this maternal contribution, ovary, unfertilized and 0 hPL transcriptomes were used to examine the expression of R. prolixus orthologues of genes reported maternal in D. melanogaster. The dataset was comprised of 10,277 sequences, of which the 54% had a R. prolixus ortholog to the D. melanogaster genes (Fig. 3 and Additional file 7). 40.7% were expressed during the three stages, oogenesis, unfertilized and 0 hPL eggs, 1.46% of the maternal genes had R. prolixus orthologs which were expressed only during oogenesis but not in deposited eggs, 1.72% had R. prolixus orthologs that were only maternally loaded into unfertilized eggs, and 2.03% only showed expression in the first hours of fertilized eggs. A similar analysis was reported with unfertilized eggs of T. castaneum with respect to D. melanogaster maternal genes. Here we identify in R. prolixus 306 maternal orthologs in addition to the ones reported in T. castaneum15. Out of 81 developmental genes investigated, eight were not reported to be maternal neither in D. melanogaster nor T. castaneum (Table 4). These eight genes were identified in the transcriptome of unfertilized eggs (gooseberry: RPRC008887, odd paired: RPRC013047, odd skipped: RPRC011812, rhomboid: RPRC008474, sloppy paired: RPRC000987), vitellogenic ovary (pipe: RPRC005278), or in both, vitellogenic ovary and unfertilized eggs (windbeutel: RPRC007848 and proboscipedia: RPRC002128). We also identified a subset of transcripts derived from the Hox cluster: abdominal-A (RPRC000598), Abdominal-B (RPRC012974), Deformed (RPRC000437), proboscipedia and Ultrabithorax (RPRC000565) (Table 3A, Table 4), suggesting maternal/zygotic transcription of the HOX genes in R. prolixus.
Taken together, our data support the notion that maternal expression of developmental genes is widespread in R. prolixus and maintained (either by mRNA stability or zygotic transcription) during early developmental stages. The results agree with other gene specific studies that have reported maternal expression in the triatomine, during oogenesis and embryo development46,48,50,69. This compelling feature deserves further investigation, however it is currently limited by the inability to discriminate maternal and zygotic effects in R. prolixus by parental RNAi.
Maternal gene expression validation
To corroborate the gene expression patterns revealed by the transcriptomic analyses, 12 genes out of 81 genes involved in oogenesis and early embryo development were chosen for qRT-PCR. The analysis was performed with three independent biological replicates different from those used for RNA-seq. The relative transcript levels over time are shown in Fig. 4. All of the selected genes showed transcript variation consistent with the results derived from the transcriptome analysis, indicating that our approach, although it was not intended to be quantitative, was valid for the identification of expressed genes. All the genes analyzed showed presence of the mRNA in unfertilized eggs, which confirms that they were maternally provided. The different expression level between the vitellogenic ovary and the unfertilized egg could be explained by the nature of the samples, which implied ovarioles depleted of choriogenic egg chambers. Rp-sqd showed higher relative transcript levels than the other genes of interest at the different analyzed times.
Real-time quantitative PCR of expression of candidate genes in the different stages. X axis: developmental times analyzed. Y axis: expression relative to the reference gene. Values are expressed as mean ± SEM of 3 independent experiments. Rp-arm: Armadillo; Rp-BicD: Bicaudal D; Rp-cact: Cactus; Rp-capu: Cappuccino; Rp-dl: Dorsal; Rp-egh: Egghead; Rp-egl: Egalitarian; Rp-exu: Exuperantia; Rp-pb: Proboscipedia; Rp-pum: Pumilio; Rp-sqd: Squid; Rp-stau: Staufen. Graphs were performed using GraphPad Prism 7. * < 0.1; ** < 0.05.
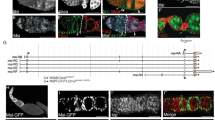
To extend the validation, we further studied the ortholog gene Rp-BicD (Additional file 8) by in situ hybridization and parental RNAi. Due to the limitation of the parental RNAi technique to distinguish maternal from zygotic phenotypes, we selected a gene that displayed a maternal, stage-specific expression, during oogenesis and early development of R. prolixus and that has not been studied in other insect beyond D. melanogaster. The structure of an ovariole is shown in Fig. 5A-C. The expression of Rp-BicD is cytoplasmic and the mRNA is detected by the antisense probe in both, the germarium and follicular epithelium of previtellogenic and vitellogenic oocytes, while the sense probe does not show any sign of hybridization (Fig. 5D-F). Rp-BicD mRNA is also detected in unfertilized eggs confined in the central region of the egg, with a diffusion of the signal towards the embryo surface (Fig. 5G). In embryos up to the onset of blastoderm stage, Rp-BicD mRNA could not be detected (data not shown), in agreement with our qRT-PCR data, although we cannot rule out transcripts below the detection limit of the in situ hybridization technique. Similar to D. melanogaster, the only insect species in which BicD has been studied, the expression decays just prior to the formation of the cellular blastoderm102. The dynamic of Rp-BicD expression suggests that the maternal transcript occurs during the initial nuclear cleavages but is degraded before cellularization of the blastoderm9,103.

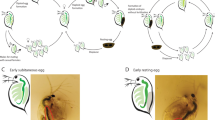
adapted from McLaughlin and Bratu116. (B) Structure of the ovariole showing the different stages that characterized oogenesis: previtellogenic, vitellogenic and choriogenic stage. (C) Ovariole of a control female showing the nuclei distribution by DAPI staining and the actin filaments by phalloidin staining50,117. (D) Detection of the sense probe by in situ hybridization assays in early stages of oogenesis. Scale Bar: 20 µm. (E) Detection of Rp-BicD transcript in the germarium and follicular cells of the ovariole with the antisense probe. Note the detection in Z3 of the germarium indicates Rp-BicD specific expression. Scale Bar: 20 µm. (F) Higher magnification of the developing oocytes. Notice the expression of Rp-BicD in the follicular cells. Scale Bar: 50 µm. (G) Detection of Rp-BicD transcript in unfertilized eggs by in situ hybridization. The arrowhead indicated the region of transcript accumulation. (H) Blastoderm embryo derived from control females. Note the nuclei distribution in the surface as revealed by DAPI staining. Scale bar: 100 µm. P: Posterior pole of the egg. (I) Egg from silenced (RNAiBicD) females. Note the lack of nuclei in the surface of the embryo. Scale bar: 100 µm. P: Posterior pole of the egg. The developmental stages corresponding to both, control (G) and silenced (H), represent the cellular blastoderm (24 hPL). (J) Ovariole of a control females showing the nuclei distribution by DAPI staining. Scale bar: 100 µm. (K) Follicular epithelium of vitellogenic oocytes from control females showing the nuclei distribution of the follicular epithelium by DAPI staining. Scale bar: 100 µm. (L) Ovariole of silenced (RNAiBicD) females showing the nuclei distribution by DAPI staining. Scale bar: 100 µm. (M) Follicular epithelium in vitellogenic oocytes from silenced (RNAiBicD) females showing the nuclei distribution of the follicular epithelium by DAPI staining. Scale bar: 100 µm.
Silencing of Rp-BicD produces anembryonic eggs. (A) Schematic of the ovariole showing the germarium host mitotically active cells (i.e. nurse cells) in zone 1 (Z1), zone 2 (Z2) and zone 3 (Z3), and previtellogenic (Pv), vitellogenic (V) and choriogenic (Ch) oocytes. Each oocyte becomes encapsulated by follicle cells (Fc), and remains connected to the germarium through the trophic cords (Tch). Germinal vesicle (Gv). Schematic representation
In order to determine the function of Rp-BicD we performed parental RNAi. We injected non-fed virgin females (n = 20) with different concentrations (0.5 to 2.5 µg/female) of dsRNABicD. As control, we used dsRNA dsRNAβ-lac (see methods). After feeding and mating, dsRNABicD and dsRNAβ-lac injected females were evaluated for survival, egg deposition, embryo lethality, and ovary and/or embryonic phenotype (Additional file 9). In our hands, 45% females injected with dsRNABicD did not survived to reach mating and egg deposition. Fertility was assessed in the surviving females (n = 9) by the number of eggs laid, while embryo lethality was studied by incubation of the eggs for the time of embryogenesis to finish (> 14 days). Compared to the control, the silenced females laid less eggs (n = 43). From these, only four corresponding to the group injected with the lowest concentration of dsRNABicD hatched to first-instar larvae (Additional file 9). The freshly laid eggs from both control and silenced females have the characteristic pink color due to the presence of the Rhodnius heme-binding protein (RHBP) in the yolk and do not show any visible chorion abnormality (Supplementary Fig. 10). As development proceeds, we monitor the embryogenesis by the coloration of the embryo, which can be observed through the chorion, white and transparent101. The control eggs showed the development of embryo pigmentation through the chorion, eventually resulting in hatchlings. The eggs derived from silenced females did not show evidence of pigmentation, suggesting that the development was arrested. The dissection of fixed silenced eggs confirmed the absence of any distinguishable embryonic structure, suggesting that BicD might act at very early stages of embryogenesis. DAPI staining of fixed early embryos showed that, different than the control, the nuclei are not observed in the surface, thus, the blastoderm was not formed. This suggests that the lack of Rp-BicD might affect a process as early as the embryonic nuclear cleavage. (Fig. 5H-I). The morphology of the ovary analyzed under the dissection microscope and the cellular pattern, oocyte and follicle cells, as judged by nuclear staining, did not show conspicuous differences between the control and silenced females. Therefore, silencing of Rp-BicD did not alter the normal morphology of the ovaries (Fig. 5J-M). The unaffected pattern of the follicle cells agrees with the normal pattern observed in the chorion in eggs derived from silenced females. The observed phenotype shows some differences with the BicD phenotype in D. melanogaster. In heterozygous BicD females the progression of oogenesis is not affected, but it causes sterility or inviable embryos that consist of a mirror-image duplication of 2–4 posterior segments104,105. Homozygous BicD females show ovaries with no diploid germ cell nuclei visible in older egg chambers and no oocyte development. The lack of BicD affects the zygotic viability, in which null flies die as pupae or young adults106. Our data indicate that in R. prolixus the function might be affecting the earliest steps of embryogenesis, but not the formation of the egg. Interestingly, in D. melanogaster, BicD and Egalitarian (Egl) are part of a complex that transports and localizes mRNAs in the oocyte, crucial for specifying the embryo axes104,107,108,109. Egl and BicD homologues have been identified in Caenorhabditis elegans and mammals, and proposed as a part of an evolutionarily conserved cytoskeletal system for mRNA transport110,111. Our qRT-PCR results show similar expression dynamics for Rp-BicD and Rp-egl, suggesting that this contemporary expression might reflect the conservation of the BicD/Egl localization machinery in R. prolixus, an aspect yet unexplored in this insect.
Conclusions
Our data provide a framework for functional studies of R. prolixus oogenesis and embryogenesis. We identified at least three temporal patterns of gene expression: 1. genes that are maternally expressed and rapidly decreased (Rp-capu, Rp-egh, Rp-exu); 2. genes with transient expression (Rp-arm, Rp-BicD, Rp-egl, Rp-sqd, Rp-pb, Rp-stau); 3. genes with invariant expression (Rp-cact, Rp-dl, Rp-pum). These dynamic suggest temporal roles based on known phenotypes: 1. Early oogenesis (genes that affect germ cell survival or results in atresic follicles): Rp-ATG-8112; Rp-cactus46; Rp-Piwi-2, Rp-Piwi-3, Rp-Argonauta-348; 2. Late oogenesis (genes that results in eggs with incomplete yolk load or impaired choriogenesis): Rp-Bicaudal C50; ULK/Rp-ATG-1113; Rp-ATG-3114; Rp-ATG-6115; 3. Embryonic patterning genes: Rp-dorsal46; Rp-Bicaudal D (this work).
Genome analysis opened an exciting path to study the molecular mechanism involved in R. prolixus oogenesis, a model established by the pioneer work of Erwin Huebner. These studies will enrich our knowledge on the evolution of development and, in the case of R. prolixus, might contribute to envisage new strategies to control the reproduction of Chagas disease vectors.
Change history
09 November 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41598-022-23779-5
References
Moussian, B. & Roth, S. Dorsoventral axis formation in the Drosophila embryo–shaping and transducing a morphogen gradient. Curr. Biol. 15, R887-899. https://doi.org/10.1016/j.cub.2005.10.026 (2005).
Dzamba, B. J. & DeSimone, D. W. Extracellular matrix (ECM) and the sculpting of embryonic tissues. Curr. Top. Dev. Biol. 130, 245–274. https://doi.org/10.1016/bs.ctdb.2018.03.006 (2018).
Salazar-Ciudad, I. Morphological evolution and embryonic developmental diversity in metazoa. Development 137, 531–539. https://doi.org/10.1242/dev.045229 (2010).
Davis, G. K. & Patel, N. H. Short, long, and beyond: Molecular and embryological approaches to insect segmentation. Annu. Rev. Entomol. 47, 669–699. https://doi.org/10.1146/annurev.ento.47.091201.145251 (2002).
Liu, P. Z. & Kaufman, T. C. Short and long germ segmentation: unanswered questions in the evolution of a developmental mode. Evol. Dev. 7, 629–646. https://doi.org/10.1111/j.1525-142X.2005.05066.x (2005).
El-Sherif, E., Lynch, J. A. & Brown, S. J. Comparisons of the embryonic development of Drosophila, Nasonia, and Tribolium. Wiley Interdiscip. Rev. Dev. Biol. 1, 16–39. https://doi.org/10.1002/wdev.3 (2012).
Mellanby, H. Memoirs: The early embryonic development of Rhodnius prolixus (Hemiptera, Heteroptera). Q. J. Microscop. Sci. s2-78, 71–90 (1935).
Lynch, J. A. & Roth, S. The evolution of dorsal-ventral patterning mechanisms in insects. Genes Dev. 25, 107–118. https://doi.org/10.1101/gad.2010711 (2011).
Tadros, W. & Lipshitz, H. D. The maternal-to-zygotic transition: a play in two acts. Development 136, 3033–3042. https://doi.org/10.1242/dev.033183 (2009).
Behura, S. K. et al. Comparative genomic analysis of Drosophila melanogaster and vector mosquito developmental genes. PLoS ONE 6, e21504. https://doi.org/10.1371/journal.pone.0021504 (2011).
Harker, B. W. et al. Stage-specific transcription during development of Aedes aegypti. BMC Dev. Biol. 13, 29. https://doi.org/10.1186/1471-213X-13-29 (2013).
Ewen-Campen, B. et al. The maternal and early embryonic transcriptome of the milkweed bug Oncopeltus fasciatus. BMC Genomics 12, 61. https://doi.org/10.1186/1471-2164-12-61 (2011).
Fan, X. B. et al. An overview of embryogenesis: External morphology and transcriptome profiling in the Hemipteran Insect Nilaparvata lugens. Front. Physiol. 11, 106. https://doi.org/10.3389/fphys.2020.00106 (2020).
Lavore, A. et al. Comparative analysis of zygotic developmental genes in Rhodnius prolixus genome shows conserved features on the tracheal developmental pathway. Insect Biochem. Mol. Biol. 64, 32–43. https://doi.org/10.1016/j.ibmb.2015.06.012 (2015).
Preuss, K. M., Lopez, J. A., Colbourne, J. K. & Wade, M. J. Identification of maternally-loaded RNA transcripts in unfertilized eggs of Tribolium castaneum. BMC Genomics 13, 671. https://doi.org/10.1186/1471-2164-13-671 (2012).
Hu, X., Ke, L., Wang, Z. & Zeng, Z. Dynamic transcriptome landscape of Asian domestic honeybee (Apis cerana) embryonic development revealed by high-quality RNA sequencing. BMC Dev. Biol. 18, 11. https://doi.org/10.1186/s12861-018-0169-1 (2018).
Simon, S. et al. Comparative transcriptomics reveal developmental turning points during embryogenesis of a hemimetabolous insect, the damselfly Ischnura elegans. Sci. Rep. 7, 13547. https://doi.org/10.1038/s41598-017-13176-8 (2017).
Clark, E. Dynamic patterning by the Drosophila pair-rule network reconciles long-germ and short-germ segmentation. PLoS Biol. 15, e2002439. https://doi.org/10.1371/journal.pbio.2002439 (2017).
Clark, E. & Peel, A. D. Evidence for the temporal regulation of insect segmentation by a conserved sequence of transcription factors. Development https://doi.org/10.1242/dev.155580 (2018).
Damen, W. G., Janssen, R. & Prpic, N. M. Pair rule gene orthologs in spider segmentation. Evol. Dev. 7, 618–628. https://doi.org/10.1111/j.1525-142X.2005.05065.x (2005).
Farzana, L. & Brown, S. J. Hedgehog signaling pathway function conserved in Tribolium segmentation. Dev. Genes Evol. 218, 181–192. https://doi.org/10.1007/s00427-008-0207-2 (2008).
Janssen, R. & Budd, G. E. Deciphering the onychophoran “segmentation gene cascade”: Gene expression reveals limited involvement of pair rule gene orthologs in segmentation, but a highly conserved segment polarity gene network. Dev. Biol. 382, 224–234. https://doi.org/10.1016/j.ydbio.2013.07.010 (2013).
Benton, M. A. A revised understanding of Tribolium morphogenesis further reconciles short and long germ development. PLoS Biol. 16, e2005093. https://doi.org/10.1371/journal.pbio.2005093 (2018).
Oppenheim, S. J., Baker, R. H., Simon, S. & DeSalle, R. We can’t all be supermodels: The value of comparative transcriptomics to the study of non-model insects. Insect Mol. Biol. 24, 139–154. https://doi.org/10.1111/imb.12154 (2015).
Arbeitman, M. N. et al. Gene expression during the life cycle of Drosophila melanogaster. Science 297, 2270–2275. https://doi.org/10.1126/science.1072152 (2002).
De Renzis, S., Elemento, O., Tavazoie, S. & Wieschaus, E. Unmasking activation of the zygotic genome using chromosomal deletions in the Drosophila embryo. PLoS ONE 5, 1036–1051 (2007).
Lecuyer, E. et al. Global analysis of mRNA localization reveals a prominent role in organizing cellular architecture and function. Cell 131, 174–187. https://doi.org/10.1016/j.cell.2007.08.003 (2007).
Tomancak, P. et al. Global analysis of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 8, R145. https://doi.org/10.1186/gb-2007-8-7-r145 (2007).
Tomancak, P. et al. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol. https://doi.org/10.1186/gb-2002-3-12-research0088 (2002).
Pridohl, F. et al. Transcriptome sequencing reveals maelstrom as a novel target gene of the terminal system in the red flour beetle Tribolium castaneum. Development 144, 1339–1349. https://doi.org/10.1242/dev.136853 (2017).
Amsel, D., Vilcinskas, A. & Billion, A. Evaluation of high-throughput isomiR identification tools: illuminating the early isomiRome of Tribolium castaneum. BMC Bioinformatics 18, 359. https://doi.org/10.1186/s12859-017-1772-z (2017).
Coura, J. R. & Borges-Pereira, J. Chagas disease. What is known and what should be improved: A systemic review. Rev. Soc. Bras. Med. Trop. 45, 286–296. https://doi.org/10.1590/s0037-86822012000300002 (2012).
Chagas, C. R. J. Nova tripanosomíase humana. Estudos sobre a morphologia e o ciclo evolutivo do Schizotrypanum cruzi n. gen. n. esp., agente da nova entidade mórbida do homem. Mem. Inst. Oswaldo Cruz. 1, 159–218 (1909).
Wigglesworth, V. B. The hormonal regularion of growth and reproduction in insects. Adv. Insect. Physiol. 2, 247–336 (1964).
Wigglesworth, V. B. The origin of sensory neurons in an insect. Q. J. Microsc. Sci 93, 93–112 (1953).
Wigglesworth, V. B. The physiology of ecdysis in Rhodnius prolixus (Hemiptera). II. Factors controlling moulting and metamorphosis. Q. J. Microsc. Sci. 77, 191–222 (1934).
Wigglesworth, V. B. The Principles of Insect Physiology (Methuen, 1939).
Huebner, E. & Anderson, E. A cytological study of the ovary of Rhodnius prolixus. Cytoarchitecture and development of the trophic chamber. J. Morphol. 138, 1–40. https://doi.org/10.1002/jmor.1051380102 (1972).
Huebner, E. & Anderson, E. A cytological study of the ovary of Rhodnius prolixus. I. The ontogeny of the follicular epithelium. J. Morphol. 136, 459–493. https://doi.org/10.1002/jmor.1051360405 (1972).
Huebner, E. & Anderson, E. A cytological study of the ovary of Rhodnius prolixus. II. Oocyte differentiation. J. Morphol. 137, 385–415. https://doi.org/10.1002/jmor.1051370402 (1972).
Lutz, D. A. & Huebner, E. Development and cellular differentiation of an insect telotrophic ovary (Rhodnius prolixus). Tissue Cell 12, 773–794. https://doi.org/10.1016/0040-8166(80)90029-4 (1980).
Lutz, D. A. & Huebner, E. Development of nurse cell-oocyte interactions in the insect telotrophic ovary (Rhodnius prolixus). Tissue Cell 13, 321–335. https://doi.org/10.1016/0040-8166(81)90008-2 (1981).
Huebner, E. Nurse cell-oocyte interaction in the telotrophic ovarioles of an insect, Rhodnius prolixus. Tissue Cell 13, 105–125. https://doi.org/10.1016/0040-8166(81)90042-2 (1981).
Huebner, E. Oocyte-follicle cell interaction during normal oogenesis and atresia in an insect. J. Ultrastruct. Res. 74, 95–104. https://doi.org/10.1016/s0022-5320(81)80112-8 (1981).
Mesquita, R. D. et al. Genome of Rhodnius prolixus, an insect vector of Chagas disease, reveals unique adaptations to hematophagy and parasite infection. Proc. Natl. Acad. Sci. U.S.A. 112, 14936–14941. https://doi.org/10.1073/pnas.1506226112 (2015).
Berni, M. et al. Toll signals regulate dorsal-ventral patterning and anterior-posterior placement of the embryo in the hemipteran Rhodnius prolixus. EvoDevo 5, 38. https://doi.org/10.1186/2041-9139-5-38 (2014).
Lavore, A., Esponda-Behrens, N., Pagola, L. & Rivera-Pomar, R. The gap gene Kruppel of Rhodnius prolixus is required for segmentation and for repression of the homeotic gene sex comb-reduced. Dev. Biol. 387, 121–129. https://doi.org/10.1016/j.ydbio.2013.12.030 (2014).
Brito, T. et al. Transcriptomic and functional analyses of the piRNA pathway in the Chagas disease vector Rhodnius prolixus. PLoS Negl. Trop. Dis. 12, e0006760. https://doi.org/10.1371/journal.pntd.0006760 (2018).
Nunes-da-Fonseca, R., Berni, M., Tobias-Santos, V., Pane, A. & Araujo, H. M. Rhodnius prolixus: From classical physiology to modern developmental biology. Genesis 55, e22995. https://doi.org/10.1002/dvg.22995 (2017).
Pascual, A., Vilardo, E. S., Taibo, C., Sabio y García, J. & Rivera-Pomar, R. Bicaudal C is required for the function of the follicular epithelium during oogenesis in Rhodnius prolixus. Dev. Genes Evol. 231, 33. https://doi.org/10.1007/s00427-021-00673-0 (2021).
Leyria, J., Orchard, I. & Lange, A. B. Transcriptomic analysis of regulatory pathways involved in female reproductive physiology of Rhodnius prolixus under different nutritional states. Sci. Rep. 10, 11431. https://doi.org/10.1038/s41598-020-67932-4 (2020).
Leyria, J., Orchard, I. & Lange, A. B. What happens after a blood meal? A transcriptome analysis of the main tissues involved in egg production in Rhodnius prolixus, an insect vector of Chagas disease. PLoS Negl. Trop. Dis. 14, e0008516. https://doi.org/10.1371/journal.pntd.0008516 (2020).
Medeiros, M. N. et al. Transcriptome and gene expression profile of ovarian follicle tissue of the triatomine bug Rhodnius prolixus. Insect. Biochem. Mol. Biol. 41, 823–831. https://doi.org/10.1016/j.ibmb.2011.06.004 (2011).
Coelho, V. L. et al. Analysis of ovarian transcriptomes reveals thousands of novel genes in the insect vector Rhodnius prolixus. Sci. Rep. 11, 1918. https://doi.org/10.1038/s41598-021-81387-1 (2021).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590-596. https://doi.org/10.1093/nar/gks1219 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36. https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Giraldo-Calderon, G. I. et al. VectorBase: An updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res. 43, D707-713. https://doi.org/10.1093/nar/gku1117 (2015).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185. https://doi.org/10.1093/bioinformatics/bts356 (2012).
Garcia-Alcalde, F. et al. Qualimap: Evaluating next-generation sequencing alignment data. Bioinformatics 28, 2678–2679. https://doi.org/10.1093/bioinformatics/bts503 (2012).
Garber, M., Grabherr, M. G., Guttman, M. & Trapnell, C. Computational methods for transcriptome annotation and quantification using RNA-seq. Nat. Methods 8, 469–477. https://doi.org/10.1038/nmeth.1613 (2011).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578. https://doi.org/10.1038/nprot.2012.016 (2012).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212. https://doi.org/10.1093/bioinformatics/btv351 (2015).
Seemann, T. & Gladman, S. Fasta Statistics: Display summary statistics for a fasta file, https://github.com/galaxyproject/tools-iuc (2012).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512. https://doi.org/10.1038/nprot.2013.084 (2013).
Zdobnov, E. M. & Apweiler, R. InterProScan–an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848. https://doi.org/10.1093/bioinformatics/17.9.847 (2001).
topGO: Enrichment Analysis for Gene Ontology v. 2.40.0 (Bioconductor, R package, 2020).
Ginzburg, N., Cohen, M. & Chipman, A. D. Factors involved in early polarization of the anterior-posterior axis in the milkweed bug Oncopeltus fasciatus. Genesis 55, e23027. https://doi.org/10.1002/dvg.23027 (2017).
Lavore, A., Pagola, L., Esponda-Behrens, N. & Rivera-Pomar, R. The gap gene giant of Rhodnius prolixus is maternally expressed and required for proper head and abdomen formation. Dev. Biol. 361, 147–155. https://doi.org/10.1016/j.ydbio.2011.06.038 (2012).
Folmes, C. D. & Terzic, A. Metabolic determinants of embryonic development and stem cell fate. Reprod. Fertil. Dev. 27, 82–88. https://doi.org/10.1071/RD14383 (2014).
Miyazawa, H. & Aulehla, A. Revisiting the role of metabolism during development. Development 145, 131110. https://doi.org/10.1242/dev.131110 (2018).
Atella, G. C. et al. Oogenesis and egg development in triatomines: A biochemical approach. An. Acad. Bras. Cienc. 77, 405–430. https://doi.org/10.1590/S0001-37652005000300005 (2005).
Fraga, A. et al. Glycogen and glucose metabolism are essential for early embryonic development of the red flour beetle Tribolium castaneum. PLoS ONE 8, e65125. https://doi.org/10.1371/journal.pone.0065125 (2013).
Vital, W. et al. Germ band retraction as a landmark in glucose metabolism during Aedes aegypti embryogenesis. BMC Dev. Biol. 10, 25. https://doi.org/10.1186/1471-213X-10-25 (2010).
Moraes, J. et al. Glucose metabolism during embryogenesis of the hard tick Boophilus microplus. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 146, 528–533. https://doi.org/10.1016/j.cbpa.2006.05.009 (2007).
Santos, R. et al. Carbohydrate accumulation and utilization by oocytes of Rhodnius prolixus. Arch. Insect Biochem. Physiol. 67, 55–62. https://doi.org/10.1002/arch.20217 (2008).
Santos, R., Rosas-Oliveira, R., Saraiva, F. B., Majerowicz, D. & Gondim, K. C. Lipid accumulation and utilization by oocytes and eggs of Rhodnius prolixus. Arch. Insect Biochem. Physiol. 77, 1–16. https://doi.org/10.1002/arch.20414 (2011).
Pontes, E. G., Leite, P., Majerowicz, D., Atella, G. C. & Gondim, K. C. Dynamics of lipid accumulation by the fat body of Rhodnius prolixus: the involvement of lipophorin binding sites. J. Insect Physiol. 54, 790–797. https://doi.org/10.1016/j.jinsphys.2008.02.003 (2008).
Fruttero, L. L., Frede, S., Rubiolo, E. R. & Canavoso, L. E. The storage of nutritional resources during vitellogenesis of Panstrongylus megistus (Hemiptera: Reduviidae): The pathways of lipophorin in lipid delivery to developing oocytes. J. Insect Physiol. 57, 475–486. https://doi.org/10.1016/j.jinsphys.2011.01.009 (2011).
Baugh, L. R., Hill, A. A., Slonim, D. K., Brown, E. L. & Hunter, C. P. Composition and dynamics of the Caenorhabditis elegans early embryonic transcriptome. Development 130, 889–900. https://doi.org/10.1242/dev.00302 (2003).
Wei, Z., Angerer, R. C. & Angerer, L. M. A database of mRNA expression patterns for the sea urchin embryo. Dev. Biol. 300, 476–484. https://doi.org/10.1016/j.ydbio.2006.08.034 (2006).
Wang, Q. T. et al. A genome-wide study of gene activity reveals developmental signaling pathways in the preimplantation mouse embryo. Dev. Cell 6, 133–144. https://doi.org/10.1016/s1534-5807(03)00404-0 (2004).
Lefebvre, F. A. & Lecuyer, E. Flying the RNA Nest: Drosophila reveals novel insights into the transcriptome dynamics of early development. J. Dev. Biol. 6, 5. https://doi.org/10.3390/jdb6010005 (2018).
Driever, W. & Nusslein-Volhard, C. The bicoid protein determines position in the Drosophila embryo in a concentration-dependent manner. Cell 54, 95–104. https://doi.org/10.1016/0092-8674(88)90183-3 (1988).
Ephrussi, A., Dickinson, L. K. & Lehmann, R. Oskar organizes the germ plasm and directs localization of the posterior determinant nanos. Cell 66, 37–50. https://doi.org/10.1016/0092-8674(91)90137-n (1991).
Gavis, E. R. & Lehmann, R. Localization of nanos RNA controls embryonic polarity. Cell 71, 301–313. https://doi.org/10.1016/0092-8674(92)90358-j (1992).
Broadus, J., Fuerstenberg, S. & Doe, C. Q. Staufen-dependent localization of prospero mRNA contributes to neuroblast daughter-cell fate. Nature 391, 792–795. https://doi.org/10.1038/35861 (1998).
Gore, A. V. et al. The zebrafish dorsal axis is apparent at the four-cell stage. Nature 438, 1030–1035. https://doi.org/10.1038/nature04184 (2005).
Takizawa, P. A., Sil, A., Swedlow, J. R., Herskowitz, I. & Vale, R. D. Actin-dependent localization of an RNA encoding a cell-fate determinant in yeast. Nature 389, 90–93. https://doi.org/10.1038/38015 (1997).
Hughes, J. R., Bullock, S. L. & Ish-Horowicz, D. Inscuteable mRNA localization is dynein-dependent and regulates apicobasal polarity and spindle length in Drosophila neuroblasts. Curr. Biol. 14, 1950–1956. https://doi.org/10.1016/j.cub.2004.10.022 (2004).
Long, R. M. et al. Mating type switching in yeast controlled by asymmetric localization of ASH1 mRNA. Science 277, 383–387. https://doi.org/10.1126/science.277.5324.383 (1997).
Neuman-Silberberg, F. S. & Schupbach, T. The Drosophila dorsoventral patterning gene gurken produces a dorsally localized RNA and encodes a TGF alpha-like protein. Cell 75, 165–174 (1993).
Simmonds, A. J., dosSantos, G., Livne-Bar, I. & Krause, H. M. Apical localization of wingless transcripts is required for wingless signaling. Cell 105, 197–207. https://doi.org/10.1016/s0092-8674(01)00311-7 (2001).
Zhang, J. et al. The role of maternal VegT in establishing the primary germ layers in Xenopus embryos. Cell 94, 515–524. https://doi.org/10.1016/s0092-8674(00)81592-5 (1998).
Melton, D. A. Translocation of a localized maternal mRNA to the vegetal pole of Xenopus oocytes. Nature 328, 80–82. https://doi.org/10.1038/328080a0 (1987).
Adereth, Y., Dammai, V., Kose, N., Li, R. & Hsu, T. RNA-dependent integrin alpha3 protein localization regulated by the Muscleblind-like protein MLP1. Nat. Cell Biol. 7, 1240–1247. https://doi.org/10.1038/ncb1335 (2005).
Lambert, J. D. & Nagy, L. M. Asymmetric inheritance of centrosomally localized mRNAs during embryonic cleavages. Nature 420, 682–686. https://doi.org/10.1038/nature01241 (2002).
Lawrence, J. B. & Singer, R. H. Intracellular localization of messenger RNAs for cytoskeletal proteins. Cell 45, 407–415. https://doi.org/10.1016/0092-8674(86)90326-0 (1986).
Mingle, L. A. et al. Localization of all seven messenger RNAs for the actin-polymerization nucleator Arp2/3 complex in the protrusions of fibroblasts. J. Cell Sci. 118, 2425–2433. https://doi.org/10.1242/jcs.02371 (2005).
Zhang, H. L. et al. Neurotrophin-induced transport of a beta-actin mRNP complex increases beta-actin levels and stimulates growth cone motility. Neuron 31, 261–275. https://doi.org/10.1016/s0896-6273(01)00357-9 (2001).
Esponda-Behrens, N. Estudios funcionales comparados de la evolución de la segmentación en insectos. Facultad de Ciencias Exactas. https://doi.org/10.35537/10915/43085 (2014).
Suter, B., Romberg, L. M. & Steward, R. Bicaudal-D, a Drosophila gene involved in developmental asymmetry: localized transcript accumulation in ovaries and sequence similarity to myosin heavy chain tail domains. Genes Dev. 3, 1957–1968 (1989).
Salz, H. K. et al. The Drosophila female-specific sexdetermination gene. Sex-lethal, has stage-, tissue-, and sex-specific RNAs suggesting multiple modes of regulation. Genes Dev. 3, 708–719 (1989).
Suter, B. & Steward, R. Requirement for phosphorylation and localization of the Bicaudal-D protein in Drosophila oocyte differentiation. Cell 67, 917–926. https://doi.org/10.1016/0092-8674(91)90365-6 (1991).
Mohler, J. & Wieschaus, E. F. Dominant maternal-effect mutations of Drosophila melanogaster causing the production of double-abdomen embryos. Genetics 112, 803–822 (1986).
Ran, B., Bopp, R. & Suter, B. Null alleles reveal novel requirements for Bic-D during Drosophila oogenesis and zygotic development. Development 120, 1233–1242 (1994).
Bullock, S. L. & Ish-Horowicz, D. Conserved signals and machinery for RNA transport in Drosophila oogenesis and embryogenesis. Nature 414, 611–616. https://doi.org/10.1038/414611a (2001).
Mach, J. M. & Lehmann, R. An Egalitarian-BicaudalD complex is essential for oocyte specification and axis determination in Drosophila. Genes Dev. 11, 423–435. https://doi.org/10.1101/gad.11.4.423 (1997).
Vazquez-Pianzola, P. et al. The mRNA transportome of the BicD/Egl transport machinery. RNA Biol. 14, 73–89. https://doi.org/10.1080/15476286.2016.1251542 (2017).
Baens, M. & Marynen, P. A human homologue (BICD1) of the Drosophila bicaudal-D gene. Genomics 45, 601–606. https://doi.org/10.1006/geno.1997.4971 (1997).
Aguirre-Chen, C., Bulow, H. E. & Kaprielian, Z. C. elegans bicd-1, homolog of the Drosophila dynein accessory factor Bicaudal D, regulates the branching of PVD sensory neuron dendrites. Development 138, 507–518. https://doi.org/10.1242/dev.060939 (2011).
Pereira, J. et al. Silencing of RpATG8 impairs the biogenesis of maternal autophagosomes in vitellogenic oocytes, but does not interrupt follicular atresia in the insect vector Rhodnius prolixus. PLoS Negl. Trop. Dis. 14, e0008012. https://doi.org/10.1371/journal.pntd.0008012 (2020).
Bomfim, L. & Ramos, I. Deficiency of ULK1/ATG1 in the follicle cells disturbs ER homeostasis and causes defective chorion deposition in the vector Rhodnius prolixus. FASEB J. 34, 13561–13572. https://doi.org/10.1096/fj.202001396R (2020).
Santos, A. & Ramos, I. ATG3 is important for the Chorion ultrastructure during oogenesis in the insect vector Rhodnius prolixus. Front. Physiol. 12, 638026. https://doi.org/10.3389/fphys.2021.638026 (2021).
Vieira, P. H., Bomfim, L., Atella, G. C., Masuda, H. & Ramos, I. Silencing of RpATG6 impaired the yolk accumulation and the biogenesis of the yolk organelles in the insect vector R. prolixus. PLoS Negl. Trop. Dis. 12, 6507. https://doi.org/10.1371/journal.pntd.0006507 (2018).
McLaughlin, J. M. & Bratu, D. P. Drosophila melanogaster oogenesis: An overview. Methods Mol Biol. 1328, 1–20. https://doi.org/10.1007/978-1-4939-2851-4_1 (2015).
Pascual, A. Genómica del desarrollo embrionario de Rhodnius prolixus. Facultad de Ciencias Exactas, Área Ciencias Biológicas. https://doi.org/10.35537/10915/90493 (2019).
Acknowledgements
The authors thank all members of Rivera-Pomar lab, in particular to Elías Gazza and Agustín Rolandelli, Nipam Patel and members of his lab for fruitful discussions, and Herbert Jäckle for critical comments on the manuscript. We thank Viviana Decker (Instituto Nacional de Tecnología Agropecuaria, Pergamino) for kindly sharing the qRT-PCR facility. R.R-P is professor at UNNOBA and investigator of CONICET. A.P is postdoctoral fellow of CONICET and UNNOBA. This work was funded by grants of The Company of Biologists to A.P. and ANPCyT (PICT-2013-1554), UNNOBA (SIB 0626/2019) and Alexander von Humboldt Research Prize to R.R.-P.
Author information
Authors and Affiliations
Contributions
A.P.: Sample preparation; Bioinformatic analysis; Data visualization; Gene expression validation; RNAi experiments; drafting the manuscript and figures; preparation of the supplementary information. R.R.-P.: Conceptualization and experimental design; Sample preparation; Bioinformatic analysis; Writing of the manuscript; figure conceptualization; Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained errors in Figure 4, where the bar graph showing the expression level of the Rp-pb gene, was omitted.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pascual, A., Rivera-Pomar, R. Dynamics of maternal gene expression in Rhodnius prolixus. Sci Rep 12, 6538 (2022). https://doi.org/10.1038/s41598-022-09874-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09874-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.