Abstract
It is generally believed that rice landraces with long culms are susceptible to lodging, and have not been utilized for breeding to improve lodging resistance. However, little is known about the structural culm strength of landraces and their beneficial genetic loci. Therefore, in this study, genome-wide association studies (GWAS) were performed using a rice population panel including Japanese rice landraces to identify beneficial loci associated with strong culms. As a result, the landraces were found to have higher structural culm strength and greater diversity than the breeding varieties. Genetic loci associated with strong culms were identified, and it was demonstrated that haplotypes with positive effects of those loci were present in a high proportion of these landraces. These results indicated that the utilization of the strong culm-associated loci present in Japanese rice landraces may further improve the lodging resistance of modern breeding varieties that have relied on semi-dwarfism.
Similar content being viewed by others
Introduction
Increasing grain yield is important to meet the global demand for major crops, which is expected to increase at least until 20501,2,3. In the 1960s, as a result of semidwarf 1 (sd1) being introduced into rice, plant height was reduced, and since then, lodging-resistant varieties have been developed and are widely used. This event averted a food security crisis and along with similar interventions in wheat and other cereals has been recognized as the “Green Revolution”4,5,6. However, the short stature promoted by sd1 in improved rice varieties, is not enough to avoid lodging especially under strong typhoons that will further increase intensity in East and Southeast Asia7, 8. It has also been suggested that semi-dwarf varieties have reduced culm strength and plant biomass compared to the original variety7, 9. Therefore, it is important to aim for high yields by increasing lodging resistance provided by increased culm strength instead of relying solely on culm dwarfism in the future.
There are two types of lodging resistance in transplanting rice cultivation: bending-type and breaking-type lodging10. Breaking-type lodging can cause the panicles to fall to the surface, making harvesting difficult11 and causing viviparous germination of grains soaked in water that will reduce the grain eating quality12. Therefore, enhancing breaking-type lodging resistance is an important issue for crop production. The bending moment at breaking (BM) of basal culm is an index of breaking-type lodging resistance13. It is composed of section modulus, which in turn is affected by the diameter and thickness of a culm, and bending stress, which is affected by the chemical composition such as cellulose and lignin and their densities in culm cell wall14, 15.
Identification and pyramiding of quantitative trait loci (QTLs) associated with strong culms in addition to sd1 constitute an important approach to breeding rice varieties with high breaking strength to withstand typhoons of increasing intensity. In our previous studies, superior alleles of STRONG CULM 1 and 2 (SCM1 and SCM2) QTLs, which increase culm strength, were detected in an indica variety Habataki16, similarly STRONG CULM 3 and 4 (SCM3 and SCM4) QTLs were detected in a tropical japonica variety Chugoku 11717, and the genes responsible for SCM2 and SCM3 were identified as APO1 ABERRANT PANICLE ORGANIZATION 1 (APO1)16, 18 and FINE CULM 1 (FC1)17, 19, 20, respectively. In addition, many other QTLs associated with strong culms have been identified21,22,23,24,25,26,27,28.
It has been reported that, japonica landrace varieties in Japan have more agriculturally useful traits when compared with japonica breeding varieties. For instance, under phosphorus-limiting conditions, the japonica landrace Akamai developed an extensive root system compared with that of the japonica breeding variety Koshihikari to obtain more phosphorus and efficiently redistributed it within the plant29. However, few studies have comprehensively compared various varieties for culm-associated traits among Japanese genetic resources13, 28.
Therefore, we hypothesize that there are novel QTLs that could enhance culm strength in japonica varieties cultivated in Japan, including landrace varieties that have not been used for breeding yet. To date, the progeny of crosses between two parents have often been used as materials for analysing and identifying QTLs30,31,32. However, this method of analysis has the disadvantages of requiring cross-breeding to grow the materials, limiting the number of varieties that can be analysed at one time. On the other hand, the development of next-generation sequencing has recently enabled the use of GWAS, which have been shown to be powerful tools for solving the abovementioned problems, as these studies allow comprehensive analysis of varieties without the need for crossing33,34,35,36. Therefore, to test our hypothesis, we used GWAS to search for strong culm-associated QTLs in Japanese japonica varieties, including landraces.
Thus, in the current study, we examined the phenotypic characteristics of lodging resistance associated traits in Japanese rice varieties and novel QTLs associated with strong culms from japonica varieties using GWAS. Furthermore, we investigated the proportions of the superior alleles between landraces and breeding varieties.
Results
Phenotypic analysis of traits associated with lodging resistance in Japanese rice
In GWAS, it is necessary that phenotypes in rice panels are highly diverse to detect QTLs efficiently35. Therefore, traits associated with lodging resistance were measured to evaluate the phenotypic diversity. Figure 1 shows the frequency distribution of phenotypes of Japanese rice, including both landraces (in yellow) and breeding varieties (in blue). For the outer diameter of the minor axis (ODMI), the landraces ranged from 3.78 to 7.68 mm in 2018 and 3.56–7.50 mm in 2019, while the breeding varieties ranged from 3.69 to 5.74 mm in 2018 and 3.59–6.09 mm in 2019. For the BM, the landraces ranged from 740 to 3,997 gf cm in 2018 and 611–4,288 gf cm in 2019, while the breeding varieties ranged from 800 to 2,439 gf cm in 2018 and 643–2,169 gf cm in 2019. With respect to culm length, the landraces ranged from 84.3 to 143.0 cm in 2018 and 67.9–155.7 cm in 2019, while the breeding varieties ranged from 64.8–116.6 cm in 2018 and 68.7–118.7 cm in 2019. The minimum value was nearly the same between the landraces and the breeding varieties except for the value of culm length in 2018. However, the maximum value of ODMI was 34% higher in 2018 and 23% higher in 2019. Moreover, the maximum value of BM was 64% higher in 2018 and 98% higher in 2019, and the maximum culm length was 23% higher in 2018 and 31% higher in 2019. A comparison of the variance of the mean phenotypic values of landraces and breeding varieties showed that the variance of each trait was higher in landraces than in breeding varieties in both years (Supplementary Data 1). Broad-sense heritability was in the order of culm length, ODMI and BM, from highest to lowest, in both years (Supplementary Data 2). Furthermore, the characteristics of lodging resistance-associated traits were investigated for each landrace and breeding variety. Figure 2 shows the mean values of the landraces and the breeding varieties. The mean value of landraces for ODMI, BM and culm length were higher than that of breeding varieties for both years (P < 0.001). These results showed that the phenotypes for the lodging resistance-associated traits were continuously distributed and varied widely, and that the landraces were more diverse than the breeding varieties and have thicker and stronger culms as well as culm length.
Frequency distribution of lodging resistance-associated traits. Histograms of phenotypic values: (a) ODMI in 2018; (b) BM in 2018; (c) culm length in 2018; (d) ODMI in 2019; (e) BM in 2019; (f) culm length in 2019. The yellow, blue and black bars indicate landraces, breeding varieties and unidentified varieties, respectively.
Trait values of landraces vs. breeding varieties. Bar graphs of mean (SD) phenotypic values: (a) ODMI in 2018; (b) BM in 2018; (c) culm length in 2018; (d) ODMI in 2019; (e) BM in 2019; (f) culm length in 2019. The yellow and blue bars indicate landraces and breeding varieties, respectively. *** indicates significant differences at P < 0.001 (two-tailed Welch’s t-test). Two-tailed Welch’s t-test was performed by R software (https://www.R-project.org/)65.
To investigate the relationship between each trait, correlation coefficients were calculated for all varieties and for landraces and breeding varieties separately (Supplementary Fig. 1). For all varieties, BM and culm length were highly positively correlated with ODMI (r ≥ 0.7). Comparing landraces and breeding varieties, BM and culm length were more positively correlated with ODMI for landraces in each year. In all cases, the correlation coefficient between BM and culm length was lower than that between the other traits.
The genetic change of rice varieties in Japan in terms of lodging resistance was identified based on their phenotypic values. Figure 3 and Supplementary Fig. 2 shows the phenotypic changes of the varieties arranged in chronological order from the left for each of the lodging resistance-associated traits (data from 2019 were used, those whose year of establishment was unclear were removed). The ODMI and the BM tended to decrease with time. Among the breeding varieties, only Zengokuwase, which was established in 1917, showed thick culms that were greater than 6.00 mm (ODMI), and no breeding varieties with thick culms (greater than 5.00 mm) after the year 2000 were found. For the BM, no varieties with values larger than 2,000 gf cm existed after Nadahikari, which was established in 1977. The culm length tended to decrease with time, and among the varieties released since 1960, only three varieties had long culms (more than 100 cm).
Phenotypic values of the studied varieties arranged in chronological order. Bar graphs of trait values of the varieties arranged by year of establishment, from left to right: (a) ODMI; (b) BM; (c) culm length. The 2019 data of phenotypic values were used, those whose year of establishment was unclear were removed. The yellow and blue bars indicate landraces and breeding varieties, respectively.
Identification of QTLs associated with strong culms via GWAS
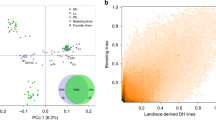
Using GWAS, we tried to identify QTLs containing superior alleles associated with a strong culm present among landraces. It has been reported that false positives are likely to occur if the population structure of the panel used for GWAS is segregated35. Therefore, before conducting our GWAS, the genetic structure of the Japanese rice varieties (excluding varieties in Hokkaido) was determined. Figure 4 shows first and second principal component (PC1 and PC2, respectively) from a principal component analysis based on the whole-genome data of the 168 and 326 varieties. These results showed that although there were no clear and strong clusters in the temperate japonica tested in this study, there were loose population structures between landraces and breeding varieties in PC1 and PC2. No distinct population structures between landraces and cultivars were found in PC3 and PC4 (Supplementary Fig. 3). Therefore, in addition to the GWAS without principal components (PCs), GWAS with PC1 and GWAS with PC1 and PC2 were also performed.
Genetic structure of temperate japonica rice varieties in Japan. Principal component analysis of temperate japonica rice in Japan based on whole-genome sequence data, with the x-axis representing the PC1 and the y-axis representing the PC2: (a) 2018; (b) 2019. Values in parentheses indicate the percentage contribution of each principal component. The yellow, blue and black markers indicate landraces, breeding varieties and unidentified varieties, respectively. Principal component analysis was performed using the package ‘pcaMethods’67 for R software (https://www.R-project.org/)65.
Next, GWAS was performed to identify QTLs associated with a strong culm. Figure 5 and Table 1 show Manhattan plots for each trait and the QTLs identified in both years without PCs as fixed factors. Q–Q plots, Local Manhattan plots and linkage disequilibrium (LD) can be found as Supplementary Figs. 4–10. For the ODMI, the same peaks above the threshold (− log10 (P) > 5) were detected on chromosomes (chrs.) 2S, 2L, 6L, 8L and 10S in both years. With respect to the BM, the same peaks above the threshold (− log10 (P) > 5) were detected on chr. 3S in both years. Peaks above the threshold (− log10 (P) > 5) were detected for culm length in each year, but they were not overlapping in both years. The peaks at 1S for ODMI and BM were slightly different from year to year. Peaks at 8L were detected for the ODMI. On the other hand, the peak positions were slightly different for the BM between 2018 and 2019. Among these abovementioned peaks, the peak at 2L in 2019 showed the highest value (− log10 (P) = 9.89) for the ODMI, and the peak at 3S in 2019 showed the highest value (− log10 (P) = 9.21) for the BM. A peak at 3S was also detected for ODMI in 2019 and culm length in 2018. When PC1 was considered as a fixed factor, the peaks showed less variation compared to when PCs were not considered, but the peak at 2L was higher for ODMI in 2019 (Fig. 5, Supplementary Fig. 11). When PC1 and PC2 were considered as fixed factors, only the peaks at 2L were above the threshold for ODMI in both years (Fig. 5, Supplementary Fig. 12). The peaks at 3S, which were above the threshold (− log10 (P) > 5) for ODMI in 2019, for BM in 2018 and 2019 and for culm length in 2018 when PCs were not considered, remained above the threshold (− log10 (P) > 5), although the peak was reduced in all cases when PC1 and PC2 were considered as fixed factors (Fig. 5, Supplementary Fig. 12). Q–Q plots with PC1 considered and those with PC1 and PC2 considered can be found as Supplementary Figs. 13, 14.
Manhattan plots of lodging resistance-associated traits. (a) ODMI in 2018; (b) BM in 2018; (c) culm length in 2018; (d) ODMI in 2019; (e) BM in 2019; (f) culm length in 2019. The x-axis indicates the SNPs or indels that physically mapped on each chromosome. The red and blue dashed lines indicate the threshold lines (− log10 (P) = 5) set in this study and the Bonferroni correction, respectively. Manhattan plots were created using the package ‘rrBLUP’68 for R software (https://www.R-project.org/)65.
Pleiotropic effects and combined effects of the QTLs
From the results of the GWAS, QTLs with particularly low P-values for ODMI or BM were detected on chrs. 2L and 3S, respectively. Therefore, targeting these QTLs, the pleiotropic effects and combined effects associated with lodging resistance were investigated. Figure 6 shows box plots of the trait values by the genotypes classified at the peak marker position in 2019. The genotypes for each variety can be found as Supplementary Data 3. The varieties in which the reference genotype (based on the Nipponbare genome) of the QTLs on chr. 2L was replaced by the alternative genotype had significantly higher ODMI and culm length values than those that was not replaced. In the comparison of breeding varieties only, the mean values of all traits were increased in varieties where the QTL of chr. 2L was replaced by the alternative genotype, but the differences were not significant (Supplementary Fig. 15). The varieties in which the reference genotype of the QTL on chr. 3S was replaced by an alternative genotype were significantly higher in ODMI, BM and culm length than those in which this QTL was not replaced. These results indicated that the substitution of the reference genotype for the alternative genotype of the QTL on chr. 2L had a pleiotropic effect on increasing the ODMI and the culm length, and the substitution of the reference genotype for the alternative genotype of the QTL on chr. 3S had a pleiotropic effect on increasing the ODMI, BM and the culm length. The varieties in which alternative genotypes of the QTLs were substituted on both chrs. 2L and 3S presented ODMI and culm length values that were significantly higher than those of any other variety group, and an increasing combined effect of QTLs on these traits was observed.
Combined effects of QTLs for lodging resistance-associated traits. Box plots of the trait values by the genotypes classified at the peak marker position in 2019: (a) ODMI; (b) BM; (c) culm length. Test results: (d) ODMI; (e) BM; f culm length. “2L” and “3S” mean the peak positions on chrs. 2L and 3S, respectively. “(−)” and “(+)” indicate the reference and alternative genotypes, respectively (based on the Nipponbare genome). The white, grey and black boxplots indicate the genotype combinations where both are references, one is a reference and the other is an alternative, and both alternatives, respectively. In the boxplots, different letters indicate significant differences at P < 0.05 (Steel–Dwass’s test). In the table, n.s. indicates no significant difference. *, **, *** indicate significant differences at P < 0.05, 0.01 and 0.001, respectively (Steel–Dwass’s test). Steel–Dwass test was performed using the package ‘NSM3’ for R software (https://www.R-project.org/)65.
To investigate the genetic origin of the two QTLs, the genotypes were classified according to peak SNPs for each landrace and breeding variety. Figure 7 shows the ratio of genotypes per QTL for the landraces and breeding varieties. For the QTL on chr. 2L, 29% of the landraces and 5% of the breeding varieties had the alternative genotype with positive effects. For the QTL on chr. 3S, 20% of the landraces and 2% of the breeding varieties had the alternative genotype with positive effects.
Ratio of genotypes per QTL for landraces and breeding varieties. Stacked bar graphs showing the ratio of genotypes at the peak: (a) on chr. 2L; (b) on chr. 3S. The grey and red bars indicate the reference and alternative genotypes, respectively. The numbers on the bars indicate the ratio, and the numbers in parentheses indicate the number of varieties.
Candidate genes for QTLs associated with strong culms
In LD for the QTL on chr. 2L, there were total of 66 genes according to the MSU Rice Genome Annotation Project (MSU) database and 49 genes according to the RAP-DB (RAP) database. In LD for the QTL on chr. 3S, there were total of 34 genes according to the MSU database and 30 genes according to the RAP database. From these, the candidate genes for each QTL were estimated. Table 2 shows the candidate genes that have mutations in coding DNA sequences (CDSs) and amino acid replacements or deletions in LD for each QTL. For the QTL on chr. 2L, eight candidate genes with amino acid replacements or deletions were evaluated. Among them, LOC_Os02g47410 was related to protein kinases, LOC_Os02g47570 was related to phosphatases, and LOC_Os02g47770 was related to zinc-finger homeodomain proteins. In contrast, GS2 was included in candidate genes with mutations in the putative promoter region (see Supplementary Table 1). For the QTL on chr. 3S, nine candidate genes with amino acid replacements were evaluated. Among them, LOC_Os03g02320 and LOC_Os03g02410 were kinase-associated genes.
Discussion
Phenotypic characteristics of lodging resistance in Japanese temperate japonica rice varieties
To date, few studies have evaluated the traits associated with strong culms of landraces13, 28. However, in this study, by comprehensively analysing the lodging resistance of a large number of varieties, we found that some landraces showed very high culm strength (Fig. 1). According to previous reports, the BM of Takanari, a semi-dwarf strong-culm indica and temperate japonica cross variety, was approximately 1,276–2,266 gf cm7, 24, 27, while that of its sister variety Habataki was approximately 1,650 gf cm17. In the present study, among the landrace varieties in Japan, there were varieties with a BM of approximately 4,000 gf cm (Fig. 1). Therefore, it was thought that some semi-dwarf varieties had high lodging resistance and that landraces in Japan had low lodging resistance because they are generally too tall and the effect of strong culms is not enough to counteract the bending moment given by a high centre of gravity, but it was found that some landraces have phenotypic characteristics associated with extremely strong culms. Although stronger culms are required to prevent the long culm plants from lodging, the effect on increasing lodging resistance given by improved strength from the landraces have a potential to be superior than the effect of plant height on decreasing lodging resistance.
Transition of breeding for lodging resistance in Japanese rice varieties
It has been reported that rice breeding for lodging resistance in Asia has been performed by shortening culms4, 37. In this study, it was shown that, compared with the landraces, the breeding varieties had shorter culms (Figs. 1, 2). Furthermore, it was confirmed that culm length tended to decrease over time (Fig. 3, Supplementary Fig. 2). In contrast, the traits associated with strong culms also tended to decrease over time (Fig. 3, Supplementary Fig. 2), and compared with the landraces, the breeding varieties had thinner and weaker culms (Figs. 1, 2). These results suggest that the improvement of lodging resistance has been dependent on breeding for relatively short culms and that breeding for relatively strong culms has not been carried out.
Both beneficial QTLs for strong culm-associated traits on chr. 2L and 3S were found to be present in a relatively high proportion of landraces (Fig. 7). Previous research showed that haplotypes exhibiting a panicle weight-type phenotype of SPINDLY (OsSPY; involved in gibberellin signalling)38 were present at a higher proportion in landraces than in breeding varieties39. Moreover, it has been shown that there is a trade-off between panicle weight and panicle number17. From these findings, it was suggested that, during the process of breeding, the emphasis on panicle weight changed to panicle number, and at the same time, the culm strength decreased.
Genetic population structure and strong culm-related loci in Japanese rice varieties detected via GWAS
The risk of false positives has been pointed out in GWAS when the genetic population structure to be analysed is separated35. In the panel of Japanese temperate japonica varietiess analyzed in this study, no clear separation of population structures as reported for rice subpopulations40 was found, while loose population structures were found between landraces and breeding varieties in PC1 and PC2 (Fig. 4). Therefore, in this study, in addition to GWAS without considering PCs, GWAS was also performed considering up to PC2 to increase the reliability of the detected QTLs.
Since the detected QTLs may change due to changes in expressed genes caused by environmental factors such as meteorological conditions41, 42, it is necessary to perform GWAS in multiple years to isolate more general loci. In this study, common peaks were detected at chrs. 2S, 2L, 6L, 8L and 10S for ODMI and at chr. 3S for BM in both 2018 and 2019, among which a particularly high peak was detected at chr. 2L for ODMI and at chr. 3S for BM (Fig. 5, Supplementary Fig. 5–10, Table 1). The peaks at chr. 2L for ODMI and 3S for BM remained above the threshold even when PC1 and PC2 were taken into account (Fig. 5, Supplementary Fig. 12). A QTL for culm diameter on chr. 5L was identified in a previous study on 135 temperate japonica varieties from different panels28, but was not detected in this study. Conversely, the QTLs on chr. 2L and 3S identified in this study were not detected in that previous study28. Unlike that previous study28, the panel used in the present study included temperate japonica varieties from the rice core collection of Japanese landraces composed by National Agriculture and Food Research Organization (NARO) and landraces from Okinawa, which may have unique genetic variations due to geographical factors. Therefore, it is possible that a novel QTL that was not previously detected was identified in this study. On the other hand, the APO1 and FC1 identified in previous studies16, 17 were not detected in this study. This suggests that the genes that enhance culm strength are different between temperate japonica and other rice subpopulations, such as indica and tropical japonica, and that the novel QTLs detected in this study may be unique to temperate japonica.
The QTL on the long arm of chromosome 2
As a result of investigating candidate genes of this QTL, eight candidate genes with amino acid replacements or deletions were evaluated (Table 2). Among them, LOC_Os02g47770 is a gene associated with a zinc-finger homeodomain (ZF-HD) protein43. Members of the rice ZF-HD gene family OsZHD1 and OsZHD2 have been reported to affect leaf morphogenesis and root meristem activity, respectively44, 45. These results suggested that, among the candidate genes, OsZHD7 might be the gene responsible for modulating culm diameter. On the other hand, GRAIN SIZE 2 (GS2) was included in the genes with mutations in the region presumed to be the promoter (Supplemental Table 1). Since it has been reported that GS2 affects grain size and weight46, 47, it might be possible that GS2 has pleiotropic effects not only on grains but also on culm morphogenesis.
The QTL on the short arm of chromosome 3
The interval of QTL on chr. 3S was identified as 0.5–0.9 Mb (Table 1). To date, QTLs for grain eating quality have been reported to be at around the 0.5 Mb mark on the short arm of chromosome 3, where the simple sequence repeat marker RM4108 (synonymous with RM14281) is located48,49,50. These previous studies have reported that Koshihikari, which is a good-tasting variety in Japan, has a beneficial allele related to grain eating quality near this marker, but this allele was not present in varieties such as Moritawase and Nipponbare. However, the haplotypes of the QTL located within 0.5–0.9 Mb associated with BM in this study were similar among Koshihikari, Moritawase, and Nipponbare (Table 1, Supplementary Data 4). This suggests that QTLs for BM and grain eating quality are likely to exist outside of LD. This means that there is a strong possibility that some of the varieties already have both strong culm-related and good-grain eating quality-related alleles at the position of chr. 3S. Therefore, these varieties can be used for breeding to improve lodging resistance without decreasing grain eating quality.
It has been shown that genes such as APO1 and FC1 have pleiotropic effects on the traits associated with strong culm and panicle morphogenesis16, 17. In this study, each trait was positively correlated, and peaks were detected in this QTL not only for BM but also for ODMI and culm length (Supplementary Fig. 1, 7, Fig. 5). From these results, it is necessary to clarify the pleiotropic and physiological functions of this QTL, including panicle morphogenesis, and to evaluate the usefulness of this QTL other than its effect on culm strength.
As a result of investigating candidate genes of this QTL, nine candidate genes with amino acid replacements were evaluated (Table 2). Among them, LOC_Os03g02320 is related to serine/threonine protein kinase, and LOC_Os03g02410 is related to GHMP kinase (Table 2). Serine/threonine protein kinases have been reported to affect plant morphogenesis51,52,53. In addition, it has been suggested that GHMP kinases may be involved in multiple responses stimulated by various plant hormones in Arabidopsis thaliana54. These results suggest that these candidate genes might be responsible for strong culms. In addition, two variants of SNPs replacing amino acid sequences in the CDS of LOC_Os03g02320 (Supplementary Data 5) were found by the database to be reference types in the indica varieties IR8 and Teqing (https://ricegenome.dna.affrc.go.jp/). Therefore, if this is the responsible gene, the alternative allele might be available for breeding these indica varieties to enhance culm strength. However, it has been reported that DNA sequence variation in the promoter region and epigenetic mutations such as DNA methylation can modulate plant traits55, 56, thus it is necessary to carefully identify the responsible genes by focusing not only on their coding DNA sequence but also on gene expression levels.
Combined effects of QTLs on culm thickness
Pyramiding QTLs is an important breeding strategy57, and it has been reported that additive effects of pyramiding and combining have been achieved for culm thickness17, 27. In the present study, as a result of examining the combined effects of QTLs, an additive combined effect was confirmed for the two identified QTLs (Fig. 6). From these results, it was suggested that culm thickness is a trait that tends to have an additive effect due to the accumulation of QTLs, and this trait may have not yet reached its maximum limit. In other words, it was suggested that additional strong-culm lines can be bred by pyramiding these QTLs and the genes associated with strong culms, such as APO116 and FC117, both of which have been identified thus far.
The combination of QTLs with the highest value for ODMI comprised alternative (non-Nipponbare) types on both chrs. 2L and 3S, and the combination that showed the lowest value comprised the reference (Nipponbare) type (Fig. 6). The QTLs on chrs. 2L and 3S of Koshihikari, Hitomebore, Hinohikari and Akitakomachi, which are the most widely cultivated varieties in Japan58, were all of the reference type (Supplementary Data 3). These varieties have weak to medium lodging resistance (https://ineweb.narcc.affrc.go.jp/index.html), and rice breeding for lodging resistance in Japan has depended on reduced culm length so far (Fig. 3, Supplementary Fig. 2). Therefore, it was suggested that these major Japanese varieties can be improved for lodging resistance by pyramiding the QTLs of these alternative types.
Prospect of breeding strategy for the “Next Green Revolution”
In this study, the landrace Houmanshindenine had an ODMI of more than 6.25 mm and a BM of approximately 4,000 gf cm (Fig. 1 and Supplementary Fig. 16). In addition, the genotype of the peak at chr. 2 of Houmanshindenine was an alternative type (Supplementary Data 3), suggesting that it may have a superior allele for strong culm-associated traits. For these reasons, the presence of beneficial alleles that enhance culm strength in landraces has been unutilised for many years; thus, these landraces have not been used as breeding materials and have been rarely inherited in modern breeding varieties.
To achieve high rice grain yields that contribute to food security and constitute the “Next Green Revolution”, one key strategy is to breed varieties with culms strong enough to withstand typhoons of increasing intensity, in addition to the strategy of semi-dwarf. A further increase in lodging resistance by increasing culm strength would also make it possible to obtain next generation plant type with larger above-ground biomass and panicles. The use of superior alleles of these landraces as breeding materials to dramatically increase fertility in the future may help to achieve this strategy.
Conclusion
In this study, we conducted a comprehensive phenotypic analysis of strong culm-associated traits in temperate japonica varieties in Japan. As a result, it was suggested that the improvement of lodging resistance in temperate japonica varieties in Japan by breeding was dependent of shortening culms. Furthermore, novel QTLs for culm strength-associated traits were identified using GWAS, and it was suggested that these superior haplotypes mainly exist in landraces. In other words, we have shown that the use of superior strong culm-related genes present in landraces that were previously thought to have low lodging resistance due to their long culm could further improve the lodging resistance of modern breeding varieties.
Methods
Plant material and cultivation
Landrace and breeding varieties collected from various regions in Japan except Hokkaido were used for rice (Oryza sativa L.) panels. A total of 168 varieties were grown in 2018, and 326 were grown in 2019. The entire list of varieties can be found as Supplementary Data 6. Some of these varieties had the same variety name but were registered under different IDs, and these were tested as different varieties. The realized additive relation matrix was calculated to determine the kinship between the varieties, and the data can be found in Supplementary Data 7 (in 2019; variety numbers are the same as in Supplementary Data 6). All plant seeds used in the current study are owned by Nagoya University and these seeds were obtained under a collaboration agreement with Nagoya University. Experimental research and field studies on plants, including the collection of plant materials in the current study, was carried out in compliance with relevant institutional, national and international guidelines and legislation. Field experiments were carried out in a paddy field in the Field Museum Honmachi, Field Science Center, Faculty of Agriculture, Tokyo University of Agriculture and Technology. The seeds were sown in germination boxes on May 7, 2018, and May 7, 2019. The seedlings at the four-leaf stage were transplanted to a paddy field on May 24, 2018, and May 23, 2019, with one seedling per hill. The planting density was 22.2 hills m−2, and the spacing was 15 × 30 cm. The residue of rice from the previous year was ploughed as compost during the winter, and N, P2O5 and K2O were applied at 50 kg ha−1, 60 kg ha−1 and 60 kg ha−1, respectively. Weed and pest control were performed as needed, and the fields were cultivated under irrigated conditions.
Phenotyping
Approximately 15 days after heading, the main culm of eight and three moderately growing individuals (dough stage) were used for phenotypic measurements in 2018 and 2019, respectively. The arithmetic mean of the traits of this individual replication for each variety was determined as the phenotypic value of the variety for each year. The leaf sheaths wrapped at the basal elongated internodes (the elongated internodes were defined as internodes that extended 4 cm or more) were removed. The ODMI at the centre of the basal elongated nodes was measured using a digital calliper. The BM was measured by a three-point bending test using a Tensilon universal material testing machine (RTG-1210; A&D, Tokyo, Japan) according an established method previously reported13. The basal elongated internodes were placed on fulcrums 4 cm apart, and a load was applied to the centre of the internodes and was measured at the time of breaking. BM was calculated by the following Eq. (1):
where M is the BM, P is the load at breaking and L is the distance between fulcrums. The culm length was defined as the length of the main culm from the base to the neck of panicle in the stretched state, measured using a ruler. Cross sections of basal elongated internodes were stained with toluidine blue and photographed using a stereoscopic microscope (SZX12; Olympus, Tokyo, Japan).
Genotyping
DNA extraction and sequencing were performed as described in a previous study35. Total DNA was extracted from leaves per variety using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany). Total DNA was sheared into −500 bp fragments by Covaris S2 (Covaris, Massachusetts, USA) and a DNA library was constructed using the NEBNext DNA Library Prep Reagent Set for Illumina (New England Biolabs, Massachusetts, USA). The DNA library was sequenced by the Illumina HiSeq 2000 (Illumina, California, USA). All reads were mapped to the Os-Nipponbare-Reference-IRGSP-1.0 pseudomolecules by BWA-MEM with the -M option in BWA software59. The mapped reads were realigned using RealignerTargetCreator and indelRealigner in GATK software60. The SNPs and indels were labelled using GATK’s UnifiedGenotyper with the -glm BOTH option. Nucleotide variants with a missing rate of > 0.2 and a minor allele frequency of < 0.05 were removed, resulting in the identification of 435,768 SNPs and indels in the 168-variety panel, and 477,640 SNPs and indels in the 326-variety panel. For the classification of polymorphisms, the promoter region was defined as the 2 kb region upstream of the translation start of each gene. The effects of genomic variants on gene function were predicted using SnpEff software (version 4.3 T)61. The information on gene location and coding sequence was obtained from Generic Feature Format Version 3 (GFF3) of the MSU Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/)62. Annotation information for genes was based on the content within the MSU Rice Genome Annotation Project and the RAP-DB (http://rapdb.dna.affrc.go.jp/)62, 63. LD heatmaps were constructed using the package ‘LDheatmap’ (version 0.99–7)64 for R software65. Genes with CDS variations and amino acid replacements or deletions and genes with promoter variations in the LD of each QTL were estimated as candidate genes (Genes of unclear function and genes related to transposon were removed). Sequence information for indica varieties for comparison of a candidate gene was obtained from TASUKE + (https://ricegenome.dna.affrc.go.jp/)66.
GWAS
The genetic structure of the analysed population was estimated using the “nipals” parameter of the package ‘pcaMethods’ (version 1.78.0)67 for R software65. The GWAS was performed using a linear mixed model (LMM) represented by the following Eq. (2):
where y is the vector of phenotypes, X is the matrix of fixed effects containing the nucleotide polymorphism, the grand mean and PCs calculated by genetic relationship analysis, β is the vector of effects, Z is the incidence matrix relating y and u. The variable u is a model of the genetic background of each variety as a random effect, with u ∼ N(0, Kσ2g), where K is the kinship matrix calculated from the nucleotide polymorphisms and σ2g is the genetic variance. ɛ is the matrix of residual effects, with ɛ ∼ N(0, Iσ2e), where I is the identity matrix and σ2e is the residual variance. The realized additive relation matrix was computed using a function “A.mat” and GWAS was performed without considering the principal component, with a fixed factor (PC1) and with fixed factors (PC1 and PC2) for the landraces and the breeding varieties using a function “GWAS” in the package ‘rrBLUP’ (version 4.6.1)68 for R software65. In the present GWAS, the threshold was set at − log10 (P) > 5, which was higher than that of several previous studies using rice panels69,70,71.
Statistical analysis
Two-tailed Welch’s t-test was performed by R software (version 4.0.5)65 to compare whether there was a difference in the means of phenotypic trait values between the groups of landraces and breeding varieties and between breeding varieties with and without a QTL. Steel–Dwass test was performed using the “pSDCFlig” function with “Asymptotic” method of the package ‘NSM3’ (version 1.16) for R software65 to compare the multiple groups classified by genotypes. The correlation coefficients between phenotypic trait values were calculated using R software65. The coefficients of determination (R2) and the adjusted R2 for the year of variety establishment and phenotypic values were calculated using R software65. Broad-sense heritability was calculated by the following Eq. (3)–(6)72, 73:
where H2 is broad-sense heritability, Mg and Me are the mean sums of squares for genotype and residual error, obtained from analysis of variance (ANOVA), r is the number of replications per genotype.
References
Godfray, H. C. J. et al. Food security: the challenge of feeding 9 billion people. Science 327, 812–818 (2010).
Tilman, D., Balzer, C., Hill, J. & Befort, B. L. Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. USA 108, 20260–20264 (2011).
Ray, D. K., Ramankutty, N., Mueller, N. D., West, P. C. & Foley, J. A. Recent patterns of crop yield growth and stagnation. Nat. Commun. 3, 1293. https://doi.org/10.1038/ncomms2296 (2012).
Hargrove, T. R. & Cabanilla, V. L. The impact of semidwarf varieties on Asian rice-breeding programs. Bioscience 29, 731–735 (1979).
Khush, G. S. Green revolution: preparing for the 21st century. Genome 42, 646–655 (1999).
Hedden, P. The genes of Green Revolution. Trends Genet. 19, 5–9 (2003).
Ookawa, T. et al. Precise estimation of genomic regions controlling lodging resistance using a set of reciprocal chromosome segment substitution lines in rice. Sci. Rep. 6, 30572. https://doi.org/10.1038/srep30572 (2016).
Mei, W. & Xie, S. P. Intensification of landfalling typhoons over the northwest Pacific since the late 1970s. Nat. Geosci. 9, 753–757 (2016).
Okuno, A. et al. New approach to increasing rice lodging resistance and biomass yield through the use of high gibberellin producing varieties. PLoS ONE 9, e86870. https://doi.org/10.1371/journal.pone.0086870 (2014).
Hirano, K., Ordonio, R. L. & Matsuoka, M. Engineering the lodging resistance mechanism of post-green revolution rice to meet future demands. Proc. Jan. Acad. Ser. B Phys. Biol. Sci. 93, 220–233 (2017).
Xangsayasane, P., Phongchanmisai, S., Bounphanousai, C. & Fukai, S. Combine harvesting efficiency as affected by rice field size and other factors and its implication for adoption of combine contracting service. Plant Prod. Sci. 22, 68–76 (2019).
Islam, M. S. et al. Lodging-related morphological traits of hybrid rice in a tropical irrigated ecosystem. Field Crops Res. 101, 240–248 (2007).
Ookawa, T. & Ishihara, K. Varietal difference of physical characteristics of the culm related to lodging resistance in paddy rice. Jpn. J. Crop Sci. 61, 419–425 (1992).
Ookawa, T. & Ishihara, K. Varietal difference of the cell wall components affecting the bending stress of the culm in relation to the lodging resistance in paddy rice. Jpn. J. Crop Sci. 62, 378–384 (1993).
Zhang, W. et al. Nitrogen fertilizer application affects lodging resistance by altering secondary cell wall synthesis in japonica rice (Oryza sativa). J. Plant Res. 130, 859–871 (2017).
Ookawa, T. et al. New approach for rice improvement using a pleiotropic QTL gene for lodging resistance and yield. Nat. Commun. 1, 132. https://doi.org/10.1038/ncomms1132 (2010).
Yano, K. et al. Isolation of a novel lodging resistance QTL gene involved in strigolactone signaling and its pyramiding with a QTL gene involved in another mechanism. Mol. Plant 8, 303–314 (2015).
Ikeda, K., Nagasawa, N. & Nagato, Y. ABERRANT PANICLE ORGANIZATION 1 temporally regulates meristem identity in rice. Dev. Biol. 282, 349–360 (2005).
Takeda, T. et al. The OsTB1 gene negatively regulates lateral branching in rice. Plant J. 33, 513–520 (2003).
Minakuchi, K. et al. FINE CULM1 (FC1) works downstream of strigolactones to inhibit the outgrowth of axillary buds in rice. Plant Cell Physiol. 51, 1127–1135 (2010).
Kashiwagi, T. & Ishimaru, K. Identification and functional analysis of a locus for improvement of lodging resistance in rice. Plant Physiol. 134, 676–683 (2004).
Ishimaru, K. et al. New target for rice lodging resistance and its effect in a typhoon. Planta 227, 601–609 (2008).
Kashiwagi, T. Identification of quantitative trait loci for resistance to bending-type lodging in rice (Oryza sativa L.). Euphytica 198, 353–367 (2014).
Mulsanti, I. W. et al. Finding the superior allele of japonica-type for increasing stem lodging resistance in indica rice varieties using chromosome segment substitution lines. Rice 11, 25. https://doi.org/10.1186/s12284-018-0216-3 (2018).
Jiang, M. et al. Mapping of QTLs associated with lodging resistance in rice (Oryza sativa L.) using the recombinant inbred lines derived from two high yielding cultivars, Tachisugata and Hokuriku 193. Plant Growth Regul. 87, 267–276 (2019).
Samadi, A. F. et al. Identification of quantitative trait loci for breaking and bending types lodging resistance in rice, using recombinant inbred lines derived from Koshihikari and a strong culm variety Leaf Star. Plant Growth Regul. 89, 83–98 (2019).
Nomura, T. et al. Next generation long-culm rice with superior lodging resistance and high grain yield, Monster Rice 1. PLoS ONE 14, e0221424. https://doi.org/10.1371/journal.pone.0221424 (2019).
Chigira, K. et al. Landraces of temperate japonica rice have superior alleles for improving culm strength associated with lodging resistance. Sci. Rep. 10, 19855. https://doi.org/10.1038/s41598-020-76949-8 (2020).
Dissanayaka, D. M. S. B., Maruyama, H., Nishida, S., Tawaraya, K. & Wasaki, J. Landrace of japonica rice, Akamai exhibits enhanced root growth and efficient leaf phosphorus remobilization in response to limited phosphorus availability. Plant Soil 414, 327–338 (2017).
Xiao, J., Li, J., Yuan, L. & Tanksley, S. D. Identification of QTLs affecting traits of agronomic importance in a recombinant inbred population derived from a subspecific rice cross. Theor. Appl. Genet. 92, 230–244 (1996).
Mei, H. W. et al. Gene actions of QTLs affecting several agronomic traits resolved in a recombinant inbred rice population and two backcross populations. Theor. Appl. Genet. 110, 649–659 (2005).
Kwon, S. J. et al. QTL mapping of agronomic traits using an RIL population derived from a cross between temperate japonica cultivars in rice (Oryza sativa L.). Breed. Sci. 58, 271–279 (2008).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Zhao, K. et al. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2, 467. https://doi.org/10.1038/ncomms1467 (2011).
Yano, K. et al. Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice. Nat. Genet. 48, 927–934 (2016).
Wang, Q., Tang, J., Han, B. & Huang, X. Advances in genome-wide association studies of complex traits in rice. Theor. Appl. Genet. 133, 1415–1425 (2020).
Asano, K. et al. Artificial selection for a green revolution gene during japonica rice domestication. Proc. Natl. Acad. Sci. USA 108, 11034–11039 (2011).
Shimada, A. et al. The rice SPINDLY gene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1, and modulating brassinosteroid synthesis. Plant J. 48, 390–402 (2006).
Yano, K. et al. GWAS with principal component analysis identifies a gene comprehensively controlling rice architecture. Proc. Natl. Acad. Sci. USA 116, 21262–21267 (2019).
McCouch, S. R. et al. Open access resources for genome-wide association mapping in rice. Nat. Commun. 7, 10532. https://doi.org/10.1038/ncomms10532 (2016).
Cao, G. et al. Impact of epistasis and QTL×environment interaction on the developmental behavior of plant height in rice (Oryza sativa L.). Theor. Appl. Genet. 103, 153–160 (2001).
Zhang, Z. H., Qu, X. S., Wan, S., Chen, L. H. & Zhu, Y. G. Comparison of QTL controlling seedling vigour under different temperature conditions using recombinant inbred lines in rice (Oryza sativa). Ann. Bot. 95, 423–429 (2005).
Hu, W., DePamphilis, C. W. & Ma, H. Phylogenetic analysis of the plant-specific zinc finger-homeobox and mini zinc finger gene families. J. Integr. Plant Biol. 50, 1031–1045 (2008).
Xu, Y. et al. Overexpression of OsZHD1, a zinc finger homeodomain class homeobox transcription factor, induces abaxially curled and drooping leaf in rice. Planta 239, 803–816 (2014).
Yoon, J. et al. Homeobox transcription factor OsZHD2 promotes root meristem activity in rice by inducing ethylene biosynthesis. J. Exp. Bot. 71, 5348–5364 (2020).
Hu, J. et al. A rare allele of GS2 enhances grain size and grain yield in rice. Mol. Plant 8, 1455–1465 (2015).
Duan, P. et al. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 15203. https://doi.org/10.1038/nplants.2015.203 (2016).
Wada, T., Ogata, T., Tsubone, M., Uchimura, Y. & Matsue, Y. Mapping of QTLs for eating quality and physicochemical properties of the japonica rice ‘Koshihikari’. Breed. Sci. 58, 427–435 (2008).
Takeuchi, Y. et al. Major QTLs for eating quality of an elite Japanese rice cultivar, Koshihikari, on the short arm of chromosome 3. Breed. Sci. 58, 437–445 (2008).
International Rice Genome Sequencing Project & Sasaki, T. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Li, J. et al. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signalling. Cell 110, 213–222 (2002).
Mao, X., Zhang, H., Tian, S., Chang, X. & Jing, R. TaSnRK2.4, an SNF1-type serine/threonine protein kinase of wheat (Triticum aestivum L.), confers enhanced multistress tolerance in Arabidopsis. J. Exp. Bot. 61, 683–696 (2010).
Song, M. et al. A leaf shape mutant provides insight into PINOID Serine/Threonine Kinase function in cucumber (Cucumis sativus L.). J. Integr. Plant Biol. 61, 1000–1014 (2019).
Xiao, W. et al. Genome-wide identification, classification and expression analysis of GHMP genes family in Arabidopsis thaliana. Plant Syst. Evol. 301, 2125–2140 (2015).
Duan, P. et al. Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol. Plant 10, 685–694 (2017).
Miura, K. et al. A metastable DWARF1 epigenetic mutant affecting plant stature in rice. Proc. Natl. Acad. Sci. USA 106, 11218–11223 (2009).
Ashikari, M. & Matsuoka, M. Identification, isolation and pyramiding of quantitative trait loci for rice breeding. Trends Plant Sci. 11, 344–350 (2006).
Kobayashi, A., Hori, K., Yamamoto, T. & Yano, M. Koshihikari: a premium short-grain rice cultivar—Its expansion and breeding in Japan. Rice 11, 15. https://doi.org/10.1186/s12284-018-0207-4 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 4. https://doi.org/10.1186/1939-8433-6-4 (2013).
Sakai, H. et al. Rice Annotation Project Database (RAP-DB): an integrative and interactive database for rice genomics. Plant Cell Physiol. 54, 6. https://doi.org/10.1093/pcp/pcs183 (2013).
Shin, J.-H., Blay, S., McNeney, B. & Graham, J. LDheatmap: an R function for graphical display of pairwise linkage disequilibria between single nucleotide polymorphisms. J. Stat. Softw. 16, 3 (2006).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing https://www.R-project.org/ (2021).
Kumagai, M. et al. TASUKE+: a web-based platform for exploring GWAS results and large-scale resequencing data. DNA Res. 26, 445–452 (2019).
Stacklies, W., Redestig, H., Scholz, M., Walther, D. & Selbig, J. pcaMethods—a bioconductor package providing PCA methods for incomplete data. Bioinformatics 23, 1164–1167 (2007).
Endelman, J. B. Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4, 250–255 (2011).
Wissuwa, M. et al. Unmasking novel loci for internal phosphorus utilization efficiency in rice germplasm through genome-wide association analysis. PLoS ONE 10, e0124215. https://doi.org/10.1371/journal.pone.0124215 (2015).
Zhu, Y. et al. Identification and fine mapping of a stably expressed QTL for cold tolerance at the booting stage using an interconnected breeding population in rice. PLoS ONE 10, e0145704. https://doi.org/10.1371/journal.pone.0145704 (2015).
Li, C. et al. Genome-wide association mapping of resistance against rice blast strains in South China and identification of a new Pik allele. Rice 12, 47. https://doi.org/10.1186/s12284-019-0309-7 (2019).
Singh, M., Ceccarelli, S. & Hamblin, J. Estimation of heritability from varietal trials data. Theor. Appl. Genet. 86, 437–441 (1993).
Kruijer, W. et al. Marker-based estimation of heritability in immortal populations. Genetics 199, 379–398 (2015).
Acknowledgements
This work was supported by JSPS Grant-in-Aid for JSPS Fellows Grant Number 20J13277.
Author information
Authors and Affiliations
Contributions
T.N. and T.O. conceived and designed the study, wrote the manuscript and secure funding. T.N., Y.S. and K.Y. designed the data analysis. T.N., Y.S., K.C. S.A. and O.S. performed the experiments and analysed the data. M.M. and K.Y. provided the plant materials and genome data. F.P. and M.R. reviewed and edited the manuscript. All authors commented the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nomura, T., Seki, Y., Matsuoka, M. et al. Potential of rice landraces with strong culms as genetic resources for improving lodging resistance against super typhoons. Sci Rep 11, 15780 (2021). https://doi.org/10.1038/s41598-021-95268-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-95268-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.